Abstract
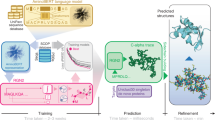
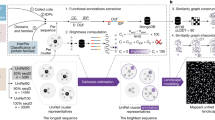
The analysis of the relationship between sequence and structure similarities during the evolution of a protein family has revealed a limit of sequence divergence for which structural conservation can be confidently assumed and homology modeling is reliable. Below this limit, the twilight zone corresponds to sequence divergence for which homology modeling becomes increasingly difficult and requires specific methods. Either with conventional threading methods or with recent deep learning methods, such as AlphaFold, the challenge relies on the identification of a template that shares not only a common ancestor (homology) but also a conserved structure with the query. As both homology and structural conservation are transitive properties, mining of sequence databases followed by multidimensional scaling (MDS) of the query sequence space can reveal intermediary sequences to infer homology and structural conservation between the query and the template. Here, as a case study, we studied the plethodontid receptivity factor isoform 1 (PRF1) from Plethodon jordani, a member of a pheromone protein family present only in lungless salamanders and weakly related to cytokines of the IL6 family. A variety of conventional threading methods led to the cytokine CNTF as a template. Sequence mining, followed by phylogenetic and MDS analysis, provided missing links between PRF1 and CNTF and allowed reliable homology modeling. In addition, we compared automated models obtained from web servers to a customized model to show how modeling can be improved by expert information.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Kendrew JC, Bodo G, Dintzis HM, Parrish RG, Wyckoff H, Phillips DC (1958) A three-dimensional model of the myoglobin molecule obtained by x-ray analysis. Nature 181:662–666. https://doi.org/10.1038/181662a0
Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H, Shindyalov IN, Bourne PE (2000) The Protein Data Bank. Nucleic Acids Res 28:235–242. https://doi.org/10.1093/nar/28.1.235
Perdigao N, Heinrich J, Stolte C, Sabir KS, Buckley MJ, Tabor B, Signal B, Gloss BS, Hammang CJ, Rost B, Schafferhans A, O’Donoghue SI (2015) Unexpected features of the dark proteome. Proc Natl Acad Sci U S A 112(52):15898–15903. https://doi.org/10.1073/pnas.1508380112
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, Tunyasuvunakool K, Bates R, Zidek A, Potapenko A, Bridgland A, Meyer C, Kohl SAA, Ballard AJ, Cowie A, Romera-Paredes B, Nikolov S, Jain R, Adler J, Back T, Petersen S, Reiman D, Clancy E, Zielinski M, Steinegger M, Pacholska M, Berghammer T, Bodenstein S, Silver D, Vinyals O, Senior AW, Kavukcuoglu K, Kohli P, Hassabis D (2021) Highly accurate protein structure prediction with AlphaFold. Nature 596:583–589. https://doi.org/10.1038/s41586-021-03819-2
Tunyasuvunakool K, Adler J, Wu Z, Green T, Zielinski M, Zidek A, Bridgland A, Cowie A, Meyer C, Laydon A, Velankar S, Kleywegt GJ, Bateman A, Evans R, Pritzel A, Figurnov M, Ronneberger O, Bates R, Kohl SAA, Potapenko A, Ballard AJ, Romera-Paredes B, Nikolov S, Jain R, Clancy E, Reiman D, Petersen S, Senior AW, Kavukcuoglu K, Birney E, Kohli P, Jumper J, Hassabis D (2021) Highly accurate protein structure prediction for the human proteome. Nature 596:590–596. https://doi.org/10.1038/s41586-021-03828-1
Sander C, Schneider R (1991) Database of homology-derived protein structures and the structural meaning of sequence alignment. Proteins 9:56–68. https://doi.org/10.1002/prot.340090107
Rost B (1999) Twilight zone of protein sequence alignments. Protein Eng 12:85–94. https://doi.org/10.1093/protein/12.2.85
Devos D, Valencia A (2000) Practical limits of function prediction. Proteins 41:98–107
Sali A, Blundell TL (1993) Comparative protein modelling by satisfaction of spatial restraints. J Mol Biol 234:779–815. https://doi.org/10.1006/jmbi.1993.1626
Wallace IM, Blackshields G, Higgins DG (2005) Multiple sequence alignments. Curr Opin Struct Biol 15:261–266. https://doi.org/10.1016/j.sbi.2005.04.002
Kemena C, Notredame C (2009) Upcoming challenges for multiple sequence alignment methods in the high-throughput era. Bioinformatics 25:2455–2465. https://doi.org/10.1093/bioinformatics/btp452
Rollmann SM, Houck LD, Feldhoff RC (1999) Proteinaceous pheromone affecting female receptivity in a terrestrial salamander. Science 285:1907–1909. https://doi.org/10.1126/science.285.5435.1907
UniProt C (2019) UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res 47:D506–D515. https://doi.org/10.1093/nar/gky1049
El-Gebali S, Mistry J, Bateman A, Eddy SR, Luciani A, Potter SC, Qureshi M, Richardson LJ, Salazar GA, Smart A, Sonnhammer ELL, Hirsh L, Paladin L, Piovesan D, Tosatto SCE, Finn RD (2019) The Pfam protein families database in 2019. Nucleic Acids Res 47:D427–D432. https://doi.org/10.1093/nar/gky995
Mitchell AL, Attwood TK, Babbitt PC, Blum M, Bork P, Bridge A, Brown SD, Chang HY, El-Gebali S, Fraser MI, Gough J, Haft DR, Huang H, Letunic I, Lopez R, Luciani A, Madeira F, Marchler-Bauer A, Mi H, Natale DA, Necci M, Nuka G, Orengo C, Pandurangan AP, Paysan-Lafosse T, Pesseat S, Potter SC, Qureshi MA, Rawlings ND, Redaschi N, Richardson LJ, Rivoire C, Salazar GA, Sangrador-Vegas A, Sigrist CJA, Sillitoe I, Sutton GG, Thanki N, Thomas PD, Tosatto SCE, Yong SY, Finn RD (2019) InterPro in 2019: improving coverage, classification and access to protein sequence annotations. Nucleic Acids Res 47:D351–D360. https://doi.org/10.1093/nar/gky1100
Burley SK, Berman HM, Bhikadiya C, Bi C, Chen L, Di Costanzo L, Christie C, Dalenberg K, Duarte JM, Dutta S, Feng Z, Ghosh S, Goodsell DS, Green RK, Guranovic V, Guzenko D, Hudson BP, Kalro T, Liang Y, Lowe R, Namkoong H, Peisach E, Periskova I, Prlic A, Randle C, Rose A, Rose P, Sala R, Sekharan M, Shao C, Tan L, Tao YP, Valasatava Y, Voigt M, Westbrook J, Woo J, Yang H, Young J, Zhuravleva M, Zardecki C (2019) RCSB Protein Data Bank: biological macromolecular structures enabling research and education in fundamental biology, biomedicine, biotechnology and energy. Nucleic Acids Res 47:D464–D474. https://doi.org/10.1093/nar/gky1004
Andreeva A, Kulesha E, Gough J, Murzin AG (2020) The SCOP database in 2020: expanded classification of representative family and superfamily domains of known protein structures. Nucleic Acids Res 48:D376–D382. https://doi.org/10.1093/nar/gkz1064
Holm L, Sander C (1998) Removing near-neighbour redundancy from large protein sequence collections. Bioinformatics 14:423–429. https://doi.org/10.1093/bioinformatics/14.5.423
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948. https://doi.org/10.1093/bioinformatics/btm404
Madeira F, Park YM, Lee J, Buso N, Gur T, Madhusoodanan N, Basutkar P, Tivey ARN, Potter SC, Finn RD, Lopez R (2019) The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res 47:W636–W641. https://doi.org/10.1093/nar/gkz268
Edgar RC (2004) MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5:113. https://doi.org/10.1186/1471-2105-5-113
Notredame C, Higgins DG, Heringa J (2000) T-Coffee: a novel method for fast and accurate multiple sequence alignment. J Mol Biol 302:205–217. https://doi.org/10.1006/jmbi.2000.4042
Armougom F, Moretti S, Poirot O, Audic S, Dumas P, Schaeli B, Keduas V, Notredame C (2006) Expresso: automatic incorporation of structural information in multiple sequence alignments using 3D-Coffee. Nucleic Acids Res 34:W604–W608. https://doi.org/10.1093/nar/gkl092
Nicholas KB, Nicholas HB Jr, Deerfield DWI (1999) GeneDoc: analysis and visualization of genetic variation. EMBNEW.NEWS 4:14
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739. https://doi.org/10.1093/molbev/msr121
Pele J, Becu JM, Abdi H, Chabbert M (2012) Bios2mds: an R package for comparing orthologous protein families by metric multidimensional scaling. BMC Bioinformatics 13:133. https://doi.org/10.1186/1471-2105-13-133
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402. https://doi.org/10.1093/nar/25.17.3389
Eddy SR (1998) Profile hidden Markov models. Bioinformatics 14:755–763. https://doi.org/10.1093/bioinformatics/14.9.755
Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJ (2015) The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 10:845–858. https://doi.org/10.1038/nprot.2015.053
Soding J, Remmert M (2011) Protein sequence comparison and fold recognition: progress and good-practice benchmarking. Curr Opin Struct Biol 21:404–411. https://doi.org/10.1016/j.sbi.2011.03.005
Soding J (2005) Protein homology detection by HMM-HMM comparison. Bioinformatics 21:951–960. https://doi.org/10.1093/bioinformatics/bti125
Zimmermann L, Stephens A, Nam SZ, Rau D, Kubler J, Lozajic M, Gabler F, Soding J, Lupas AN, Alva V (2018) A completely reimplemented MPI bioinformatics toolkit with a new HHpred server at its core. J Mol Biol 430:2237–2243. https://doi.org/10.1016/j.jmb.2017.12.007
Zheng W, Zhang C, Wuyun Q, Pearce R, Li Y, Zhang Y (2019) LOMETS2: improved meta-threading server for fold-recognition and structure-based function annotation for distant-homology proteins. Nucleic Acids Res 47:W429–W436. https://doi.org/10.1093/nar/gkz384
Pandurangan AP, Stahlhacke J, Oates ME, Smithers B, Gough J (2019) The SUPERFAMILY 2.0 database: a significant proteome update and a new webserver. Nucleic Acids Res 47:D490–D494. https://doi.org/10.1093/nar/gky1130
Gough J, Karplus K, Hughey R, Chothia C (2001) Assignment of homology to genome sequences using a library of hidden Markov models that represent all proteins of known structure. J Mol Biol 313:903–919. https://doi.org/10.1006/jmbi.2001.5080
Buchan DWA, Jones DT (2019) The PSIPRED protein analysis workbench: 20 years on. Nucleic Acids Res 47:W402–W407. https://doi.org/10.1093/nar/gkz297
Jones DT (1999) Protein secondary structure prediction based on position-specific scoring matrices. J Mol Biol 292:195–202. https://doi.org/10.1006/jmbi.1999.3091
Heffernan R, Yang Y, Paliwal K, Zhou Y (2017) Capturing non-local interactions by long short-term memory bidirectional recurrent neural networks for improving prediction of protein secondary structure, backbone angles, contact numbers and solvent accessibility. Bioinformatics 33:2842–2849. https://doi.org/10.1093/bioinformatics/btx218
Roy A, Kucukural A, Zhang Y (2010) I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc 5:725–738. https://doi.org/10.1038/nprot.2010.5
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y (2015) The I-TASSER suite: protein structure and function prediction. Nat Methods 12:7–8. https://doi.org/10.1038/nmeth.3213
Zhang Y (2008) I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9:40. https://doi.org/10.1186/1471-2105-9-40
Song Y, DiMaio F, Wang RY, Kim D, Miles C, Brunette T, Thompson J, Baker D (2013) High-resolution comparative modeling with RosettaCM. Structure 21:1735–1742. https://doi.org/10.1016/j.str.2013.08.005
Zhang Y, Skolnick J (2004) Scoring function for automated assessment of protein structure template quality. Proteins 57:702–710. https://doi.org/10.1002/prot.20264
Xu J, Zhang Y (2010) How significant is a protein structure similarity with TM-score = 0.5? Bioinformatics 26:889–895. https://doi.org/10.1093/bioinformatics/btq066
Zhang Y, Skolnick J (2005) TM-align: a protein structure alignment algorithm based on the TM-score. Nucleic Acids Res 33:2302–2309. https://doi.org/10.1093/nar/gki524
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera--a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612. https://doi.org/10.1002/jcc.20084
Derouet D, Rousseau F, Alfonsi F, Froger J, Hermann J, Barbier F, Perret D, Diveu C, Guillet C, Preisser L, Dumont A, Barbado M, Morel A, deLapeyriere O, Gascan H, Chevalier S (2004) Neuropoietin, a new IL-6-related cytokine signaling through the ciliary neurotrophic factor receptor. Proc Natl Acad Sci U S A 101:4827–4832. https://doi.org/10.1073/pnas.0306178101
Senaldi G, Varnum BC, Sarmiento U, Starnes C, Lile J, Scully S, Guo J, Elliott G, McNinch J, Shaklee CL, Freeman D, Manu F, Simonet WS, Boone T, Chang MS (1999) Novel neurotrophin-1/B cell-stimulating factor-3: a cytokine of the IL-6 family. Proc Natl Acad Sci U S A 96:11458–11463. https://doi.org/10.1073/pnas.96.20.11458
Heinrich PC, Behrmann I, Haan S, Hermanns HM, Muller-Newen G, Schaper F (2003) Principles of interleukin (IL)-6-type cytokine signalling and its regulation. Biochem J 374:1–20. https://doi.org/10.1042/BJ20030407
Huising MO, Kruiswijk CP, Flik G (2006) Phylogeny and evolution of class-I helical cytokines. J Endocrinol 189:1–25. https://doi.org/10.1677/joe.1.06591
Rose-John S (2018) Interleukin-6 family cytokines. Cold Spring Harb Perspect Biol 10:a028415. https://doi.org/10.1101/cshperspect.a028415
Sims NA (2015) Cardiotrophin-like cytokine factor 1 (CLCF1) and neuropoietin (NP) signalling and their roles in development, adulthood, cancer and degenerative disorders. Cytokine Growth Factor Rev 26:517–522. https://doi.org/10.1016/j.cytogfr.2015.07.014
Somers W, Stahl M, Seehra JS (1997) 1.9 A crystal structure of interleukin 6: implications for a novel mode of receptor dimerization and signaling. EMBO J 16:989–997. https://doi.org/10.1093/emboj/16.5.989
Adams R, Burnley RJ, Valenzano CR, Qureshi O, Doyle C, Lumb S, Del Carmen LM, Griffin R, McMillan D, Taylor RD, Meier C, Mori P, Griffin LM, Wernery U, Kinne J, Rapecki S, Baker TS, Lawson AD, Wright M, Ettorre A (2017) Discovery of a junctional epitope antibody that stabilizes IL-6 and gp80 protein: protein interaction and modulates its downstream signaling. Sci Rep 7:37716. https://doi.org/10.1038/srep37716
Putoczki TL, Dobson RC, Griffin MD (2014) The structure of human interleukin-11 reveals receptor-binding site features and structural differences from interleukin-6. Acta Crystallogr D Biol Crystallogr 70:2277–2285. https://doi.org/10.1107/S1399004714012267
McDonald NQ, Panayotatos N, Hendrickson WA (1995) Crystal structure of dimeric human ciliary neurotrophic factor determined by MAD phasing. EMBO J 14:2689–2699
Robinson RC, Grey LM, Staunton D, Vankelecom H, Vernallis AB, Moreau JF, Stuart DI, Heath JK, Jones EY (1994) The crystal structure and biological function of leukemia inhibitory factor: implications for receptor binding. Cell 77:1101–1116. https://doi.org/10.1016/0092-8674(94)90449-9
Huyton T, Zhang JG, Luo CS, Lou MZ, Hilton DJ, Nicola NA, Garrett TP (2007) An unusual cytokine: Ig-domain interaction revealed in the crystal structure of leukemia inhibitory factor (LIF) in complex with the LIF receptor. Proc Natl Acad Sci U S A 104:12737–12742. https://doi.org/10.1073/pnas.0705577104
Deller MC, Hudson KR, Ikemizu S, Bravo J, Jones EY, Heath JK (2000) Crystal structure and functional dissection of the cytostatic cytokine oncostatin M. Structure 8:863–874. https://doi.org/10.1016/s0969-2126(00)00176-3
Schreuder HA, Rondeau JM, Tardif C, Soffientini A, Sarubbi E, Akeson A, Bowlin TL, Yanofsky S, Barrett RW (1995) Refined crystal structure of the interleukin-1 receptor antagonist. Presence of a disulfide link and a cis-proline. Eur J Biochem 227:838–847. https://doi.org/10.1111/j.1432-1033.1995.tb20209.x
Fiser A, Sali A (2003) Modeller: generation and refinement of homology-based protein structure models. Methods Enzymol 374:461–491. https://doi.org/10.1016/S0076-6879(03)74020-8
Panayotatos N, Radziejewska E, Acheson A, Somogyi R, Thadani A, Hendrickson WA, McDonald NQ (1995) Localization of functional receptor epitopes on the structure of ciliary neurotrophic factor indicates a conserved, function-related epitope topography among helical cytokines. J Biol Chem 270:14007–14014. https://doi.org/10.1074/jbc.270.23.14007
Perret D, Guillet C, Elson G, Froger J, Plun-Favreau H, Rousseau F, Chabbert M, Gauchat JF, Gascan H (2004) Two different contact sites are recruited by cardiotrophin-like cytokine (CLC) to generate the CLC/CLF and CLC/sCNTFRalpha composite cytokines. J Biol Chem 279:43961–43970. https://doi.org/10.1074/jbc.M407686200
Plun-Favreau H, Elson G, Chabbert M, Froger J, deLapeyriere O, Lelievre E, Guillet C, Hermann J, Gauchat JF, Gascan H, Chevalier S (2001) The ciliary neurotrophic factor receptor alpha component induces the secretion of and is required for functional responses to cardiotrophin-like cytokine. EMBO J 20:1692–1703. https://doi.org/10.1093/emboj/20.7.1692
Skolnick J, Zhou H (2017) Why is there a glass ceiling for threading based protein structure prediction methods? J Phys Chem B 121:3546–3554. https://doi.org/10.1021/acs.jpcb.6b09517
Abdi H (2007) Metric multidimensional scaling. In: Salkind NJ (ed) Encyclopedia of measurement and statistics. Sage, Thousand Oaks, pp 598–605
Wu S, Zhang Y (2008) MUSTER: improving protein sequence profile-profile alignments by using multiple sources of structure information. Proteins 72:547–556. https://doi.org/10.1002/prot.21945
Wu S, Zhang Y (2007) LOMETS: a local meta-threading-server for protein structure prediction. Nucleic Acids Res 35:3375–3382. https://doi.org/10.1093/nar/gkm251
Xu D, Jaroszewski L, Li Z, Godzik A (2014) FFAS-3D: improving fold recognition by including optimized structural features and template re-ranking. Bioinformatics 30:660–667. https://doi.org/10.1093/bioinformatics/btt578
Xu Y, Xu D (2000) Protein threading using PROSPECT: design and evaluation. Proteins 40:343–354
Zhou H, Zhou Y (2005) Fold recognition by combining sequence profiles derived from evolution and from depth-dependent structural alignment of fragments. Proteins 58:321–328. https://doi.org/10.1002/prot.20308
Yang Y, Faraggi E, Zhao H, Zhou Y (2011) Improving protein fold recognition and template-based modeling by employing probabilistic-based matching between predicted one-dimensional structural properties of query and corresponding native properties of templates. Bioinformatics 27:2076–2082. https://doi.org/10.1093/bioinformatics/btr350
Madera M (2008) Profile comparer: a program for scoring and aligning profile hidden Markov models. Bioinformatics 24:2630–2631. https://doi.org/10.1093/bioinformatics/btn504
Kallberg M, Wang H, Wang S, Peng J, Wang Z, Lu H, Xu J (2012) Template-based protein structure modeling using the RaptorX web server. Nat Protoc 7:1511–1522. https://doi.org/10.1038/nprot.2012.085
Ovchinnikov S, Park H, Varghese N, Huang PS, Pavlopoulos GA, Kim DE, Kamisetty H, Kyrpides NC, Baker D (2017) Protein structure determination using metagenome sequence data. Science 355:294–298. https://doi.org/10.1126/science.aah4043
Palmer CA, Watts RA, Gregg RG, McCall MA, Houck LD, Highton R, Arnold SJ (2005) Lineage-specific differences in evolutionary mode in a salamander courtship pheromone. Mol Biol Evol 22:2243–2256. https://doi.org/10.1093/molbev/msi219
Acknowledgments
This study was supported by institutional grants from INSERM, CNRS, and University of Angers. MC is supported by CNRS. RB was supported by a fellowship from the University of Angers (France). AT was supported by a fellowship from the University of Carthage (Tunisia).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
1 Electronic Supplementary Material
Fig 1
Comparison of the customized and AlphaFold models. The customized MODELLER model (residues 10-187 of mature protién, white ribbon) is superimposed with the AlphaFold model deposited in UniProt (AF_Q9PUJ2-F1, slate) and the CNTF template (PDB access number: 1CNT, magenta). Phe-70 of PRF1 and Trp-64 of CNTF are shown as sticks. (ZIP 1.67MB)
Rights and permissions
Copyright information
© 2023 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Ben Boubaker, R., Tiss, A., Henrion, D., Chabbert, M. (2023). Homology Modeling in the Twilight Zone: Improved Accuracy by Sequence Space Analysis. In: Filipek, S. (eds) Homology Modeling. Methods in Molecular Biology, vol 2627. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2974-1_1
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2974-1_1
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2973-4
Online ISBN: 978-1-0716-2974-1
eBook Packages: Springer Protocols