Abstract
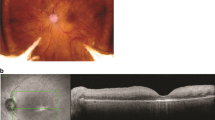
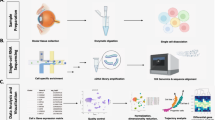
Transcriptome profiling at single-cell resolution allows us to identify and assess functional cell types and cellular states, including those within degenerating ocular tissues in retinitis pigmentosa. The technology is particularly valuable when studying tissues with high cellular heterogeneity, or when specific cell types are of interest. In this chapter, we introduce a detailed protocol of a medium-throughput single-nucleus RNA sequencing technique that utilizes frozen tissue as input sample. This protocol can be executed by any researcher with basic training in molecular biology techniques. With this protocol, a single experimenter can easily process two samples per day up to cDNA amplification, and library preparations can be done in batches of 8. Routinely we can obtain ~20 K nuclei per eye from 3 to 4 library preparations.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Huang S (2009) Non-genetic heterogeneity of cells in development: more than just noise. Development 136(23):3853–3862
Li L, Clevers H (2010) Coexistence of quiescent and active adult stem cells in mammals. Science 327(5965):542–545
Shalek AK et al (2014) Single-cell RNA-seq reveals dynamic paracrine control of cellular variation. Nature 510(7505):363–369
Hwang B, Lee JH, Bang D (2018) Single-cell RNA sequencing technologies and bioinformatics pipelines. Exp Mol Med 50(8):96
Whitmore SS et al (2014) Transcriptomic analysis across nasal, temporal, and macular regions of human neural retina and RPE/choroid by RNA-Seq. Exp Eye Res 129:93–106
Macosko EZ et al (2015) Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell 161(5):1202–1214
Zheng GX et al (2017) Massively parallel digital transcriptional profiling of single cells. Nat Commun 8:14049
Buenaventura DF, Corseri A, Emerson MM (2019) Identification of genes with enriched expression in early developing mouse cone photoreceptors. Invest Ophthalmol Vis Sci 60(8):2787–2799
Lukowski SW et al (2019) A single-cell transcriptome atlas of the adult human retina. EMBO J 38(18):e100811
Peng YR et al (2019) Molecular classification and comparative taxonomics of foveal and peripheral cells in primate retina. Cell 176(5):1222–1237. e22
Shekhar K et al (2016) Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics. Cell 166(5):1308–1323. e30
Voigt AP et al (2019) Molecular characterization of foveal versus peripheral human retina by single-cell RNA sequencing. Exp Eye Res 184:234–242
Huang HL et al (2010) Trypsin-induced proteome alteration during cell subculture in mammalian cells. J Biomed Sci 17:36
Grindberg RV et al (2013) RNA-sequencing from single nuclei. Proc Natl Acad Sci U S A 110(49):19802–19807
Habib N et al (2017) Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat Methods 14(10):955–958
Habib N et al (2016) Div-Seq: single-nucleus RNA-Seq reveals dynamics of rare adult newborn neurons. Science 353(6302):925–928
Rosenberg AB et al (2018) Single-cell profiling of the developing mouse brain and spinal cord with split-pool barcoding. Science 360(6385):176–182
Orozco LD et al (2020) Integration of eQTL and a single-cell atlas in the human eye identifies causal genes for age-related macular degeneration. Cell Rep 30(4):1246–1259 e6
Krishnaswami SR et al (2016) Using single nuclei for RNA-seq to capture the transcriptome of postmortem neurons. Nat Protoc 11(3):499–524
Butler A et al (2018) Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol 36(5):411–420
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Liu, CY., Chen, HH. (2023). Large-Scale Single-Nucleus RNA Sequencing Compatible with Complex Archived Samples. In: Tsang, S.H., Quinn, P.M. (eds) Retinitis Pigmentosa. Methods in Molecular Biology, vol 2560. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2651-1_30
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2651-1_30
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2650-4
Online ISBN: 978-1-0716-2651-1
eBook Packages: Springer Protocols