Abstract
Over the last century research in Drosophila has resulted in many fundamental contributions to our understanding of the biology of multicellular organisms. Many of these breakthroughs have been based on the identification of novel gene functions in large-scale genetic screens. However, conventional forward-genetic screens have been limited by the random nature of mutagenesis and difficulties in mapping causal mutations, while reverse-genetic RNAi screens suffer from incomplete knockdown of gene expression. Recently developed large-scale CRISPR-Cas9 libraries promise to address these limitations by allowing the induction of targeted mutations in genes with spatial and temporal control. Here, we provide a guide for tissue-specific CRISPR screening in Drosophila, including the characterization of Gal4 UAS-Cas9 lines, selection of sgRNA libraries, and various quality control measures. We also discuss confounding factors that can give rise to false-positive and false-negative results in such experiments and suggest strategies on how to detect and avoid them. Conditional CRISPR screening represents an exciting new approach for functional genomics in vivo and is set to further expand our knowledge of the molecular underpinning of development, homeostasis, and disease.
You have full access to this open access chapter, Download protocol PDF
Similar content being viewed by others
Key words
1 Introduction
Genetic screens aim to identify novel gene functions by testing many genes in parallel using unbiased approaches. In Drosophila such screens have been particularly successful and have identified many of the genes that control development, behavior, and disease in multicellular animals [1,2,3]. Central to genetic screening is a scalable method to reliably perturb gene expression, ideally including the capacity to abrogate gene function. Furthermore, large-scale screening can be facilitated by reverse-genetic approaches, which use reagents that are rationally designed to target specific genes, thereby allowing to test a large number of genes with limited resources.
Traditional screening methods in Drosophila have a number of limitations. Mutagenesis screens using chemicals, radiation, or transposable elements have the capacity to knockout gene function, but only do so relatively infrequently, thereby making comprehensive screens labor and resource intensive. RNAi screens use rationally designed reagents that minimize the number of experiments needed to test a large number of genes, but only knockdown gene expression, thereby missing phenotypes due to residual gene expression.
CRISPR-Cas9 gene editing is a highly scalable method for functional genomics [4]. It makes use of the RNA-guided endonuclease Cas9, which is directed to the genomic target site by a single guide RNA (sgRNA) [5] . The Cas9/sgRNA complex initially binds DNA through a protospacer adjacent motif (PAM), which in the case of Streptococcus pyogenes Cas9 is NGG, and represents the only restriction on genomic target space. Otherwise, target specificity is exclusively encoded in the 5′ 20 nucleotides of the sgRNA. Since it is relatively straightforward to generate new sgRNAs, this method lends itself to the generation of large resources for functional genomic screening. Binding of the Cas9 ribonucleoprotein (RNP) particle and base pairing of the sgRNA with the target sequence activates the endonuclease activity of Cas9, resulting in a cut of both DNA strands 3 base pairs upstream of the PAM. The DNA double-strand break is recognized by the endogenous DNA repair machinery and repaired by either homology-directed repair (HDR) or non-homologous end joining (NHEJ). While the former pathway results in the repair of the lesion with high fidelity, NHEJ is an error-prone repair pathway and frequently leads to small insertions and deletions (indels) at the target site (Fig. 1).
CRISPR-Cas9 for gene targeting in Drosophila. (a) In CRISPR-Cas9 gene editing a small sgRNA guides the endonuclease Cas9 to the genomic target site. Binding activates the enzymatic activity, which creates a DNA double-strand break. This is recognized by the endogenous DNA repair machinery. DNA repair can result in the restoration of the original sequence, which can be cut again by Cas9, or the induction of mutations, often small insertions and deletions (indels), which can be in- or out-of-frame. (b) Conditional CRISPR mutagenesis in Drosophila involves crossing Gal4 UAS-Cas9 transgenic flies to transgenic sgRNA lines. Offspring expresses CRISPR components in Gal4-expressing cells, leading to mutations at the target locus and subsequent phenotypes
CRISPR screening typically makes use of Cas9-induced indels, which in the coding sequence of protein-coding genes can abrogate gene function. However, in many cases only indels that represent out-of-frame mutations have a strong functional impact, while in-frame indels are often silent. Moreover, in some species it has been shown that in some instances even the effect of out-of-frame indels can be attenuated by alternative splicing or genetic compensation [6,7,8]. As a result the efficiency of CRISPR mutagenesis is not only limited by Cas9/sgRNA activity but also by the spectrum and position of the induced mutations. Several strategies have been developed to increase the fraction of bi-alleleic knockout cells, including algorithms to predict sgRNA activity [9], bioinformatic prediction of target sites where in-frame indels are likely to disrupt gene function [10, 11], and sgRNA multiplexing to induce several mutations or larger deletions in a gene [12, 13].
CRISPR genome engineering has been quickly adopted and combined with the sophisticated genetic tools available in the Drosophila model system. Early work has mostly focused on establishing methods to edit the genome in Drosophila germ cells, allowing the generation of heritable genome edits [14,15,16,17]. More recently, several labs have also developed techniques to acutely induce knockouts in genes in somatic tissues with spatial and temporal control [18,19,20,21]. This is achieved by expressing CRISPR components under the control of regulatory elements from genes with restricted expression patterns, either by cloning the Cas9 coding sequence downstream of such regulatory elements or by expressing Cas9 components under control of the Gal4/UAS system. These are then combined with transgenic sgRNAs, either expressed ubiquitously or via Gal4/UAS, through a genetic cross.
Several labs are currently in the process of generating large-scale sgRNA libraries to facilitate unbiased screening for novel gene functions in Drosophila [19, 20, 22]. These use in part different designs, including the promoters that drive sgRNA expression and the number of sgRNAs per gene. Some libraries have already been used to mutagenize many genes either ubiquitously or in selected tissues and have been found to be highly effective, often performing significantly better than previous technologies [19, 20]. Here we provide a practical guide for performing conditional CRISPR screening in Drosophila and discuss strategies for quality control.
2 Materials
2.1 Transgenic Drosophila Strains Expressing Cas9
Lines that allow optimizing Cas9 expression levels independent from the strength of the Gal4 driver are available from the Vienna Drosophila Stock Center at https://stockcenter.vdrc.at [19]. UAS-uCas9 lines have the VDRC IDs 340000-340007. The HD_CFD Tools collection at VDRC also contains a number of stocks in which UAS-uCas9 transgenes are already combined with popular Gal4 drivers. In addition, a line (VDRC ID 340008) is available that allows induction of Cas9 in negatively marked clones through FLP/FRT recombination [19]. A number of different UAS-Cas9 lines (UAS-Cas9.C, UAS-Cas9.P, UAS-Cas9.P2), as well as lines already combined with Gal4 drivers, are also available from the Bloomington Drosophila Stock Center (BDSC) at https://bdsc.indiana.edu/ [22]. Users should characterize Gal4 UAS-Cas9 lines for their intended purpose as described in Subheading 3.1.
2.2 Transgenic sgRNA Lines
Several large-scale sgRNA collections, as well as smaller collections from individual laboratories, are available from public stock centers. The Vienna Drosophila Stock Center distributes the Heidelberg CRISPR Fly Design (HD_CFD) library [19]. The Bloomington Drosophila Stock Center distributes the TRiP-KO (as well as TRiP-OE for CRISPR activation) library and the WKO library, as well as a number of smaller collections and individual lines [20,21,22,23]. Fly Stocks at the National Institute of Genetics (https://shigen.nig.ac.jp/fly/nigfly/) distributes a collection of sgRNA lines originally intended for germline mutagenesis [16]. These resources differ in their design and performance in various applications and users should consult Subheading 3.2 and Fig. 2 to choose the lines best suited for their experiments.
Cas9 and sgRNA expression constructs for efficient CRISPR mutagenesis in Drosophila. (a) The UAS-uCas9 series comprises a number of UAS-Cas9 constructs with upstream open reading frames (uORF) in between the Cas9 coding sequence and the UAS-hsp70 promoter. Translation of the downstream Cas9 ORF is inversely correlated with the length of the uORF. Choosing the appropriate vector allows titration of Cas9 expression to optimal levels. (b) sgRNA expression vectors frequently used in Drosophila CRISPR experiments. Plasmids differ in their use of promoters for sgRNA expression and whether they allow sgRNA multiplexing or not. Transgenes with different U6 promoters have been shown to mediate gene editing with different efficiency [18]
2.3 Fly Strains to Mark Cells with Active CRISPR-Cas9
A number of fly lines that allow to mark cells that underwent CRISPR mutagenesis have been described and are available upon request from the respective laboratories [24,25,26]. Note that these tools typically do not mark all cells that have been edited by CRISPR.
2.4 Plasmids
There are a number of sgRNA expression plasmids available from Addgene (https://www.addgene.org/) that allow generation of transgenic sgRNA lines. We recommend use of pCFD5 (Addgene 112645) for ubiquitous expression of sgRNAs with high efficiency and pCFD6 (Addgene 73915) for expression of sgRNAs under control of Gal4, which leads to tighter restriction of mutagenesis to Gal4-expressing cells [12].
In case users wish to generate UAS-uCas9 lines with insertion at different genomic loci the plasmids are also available from Addgene (Plasmids 127382 – 127387).
2.5 Antibody Staining
To profile cells and tissues expressing CRISPR-Cas9 components we recommend the following antibodies: Rabbit anti-phospho-Histone 3 (Millipore, Cat# 06-570) to stain mitotic cells, rabbit anti-cleaved Drosophila Dcp-1 (Millipore, Cat# AB3623) to stain cells undergoing apoptosis, and mouse anti-Fasciclin III (DSHB, Clone 7G10) to visualize cell morphology.
3 Methods
3.1 Generation and Characterization of Gal4 UAS-Cas9 Fly Lines
To induce mutations only in selected target tissues and at specific stages during development the expression of Cas9 is typically controlled via the Gal4/UAS system (see Note 1). A number of Gal4 UAS-Cas9 stocks are publicly available from the Vienna Drosophila Resource Center (VDRC) and the Bloomington Drosophila Stock Center (BDSC) (see Subheading 2.1 and Note 2). However, not all of these stocks contain the latest Cas9 constructs and often users will be required to generate their own lines. To this end UAS-Cas9 transgenes are combined with a Gal4 driver of interest by conventional genetic crosses. A number of different UAS-Cas9 constructs have been described [12, 18, 19]. We recommend use of the UAS-uCas9 series, as it allows to fine tune Cas9 expression levels (as detailed in Subheading 3.1.1). It can be desirable to recombine the Gal4 driver and UAS-Cas9 transgene on the same chromosome to simplify subsequent crosses (see Note 3). Eventually stable stocks are generated that harbor the Gal4 driver and the Cas9 transgene, which can then be crossed to transgenic sgRNA flies.
-
1.
Check if suitable Gal4 UAS-Cas9 lines are already available from the VDRC or BDSC. Order lines that are available and proceed to step 3.1.1.
-
2.
If no suitable lines are readily available, order individual UAS-Cas9 transgenic lines. We recommend use of the UAS-uCas9 series developed by our lab, which is available from the VDRC (see Note 2). We typically use UAS-uSCas9 for weak Gal4 drivers, UAS-uMCas9 for medium Gal4 drivers and UAS-uLCas9 for very strong Gal4 lines. However, we recommend to establish the optimal line empirically by testing several UAS-uCas9 transgenes in parallel.
-
3.
Cross UAS-uCas9 lines to the Gal4 driver.
-
4.
Select Gal4 UAS-uCas9 transgenic flies (use virgin females in case both transgenes are recombined on the same chromosome) and cross to a suitable balancer line.
-
5.
Select balanced Gal4 UAS-uCas9 transgenic flies and set up a stable transgenic stock.
3.1.1 Testing for Cas9-Mediated Toxicity
Cas9 protein at high expression levels is toxic to flies. It is unclear what the mechanism behind this toxicity is, but it does not require the ability to cut DNA, as high levels of nuclease-dead versions of the enzyme are also detrimental [18]. Toxicity can be so severe that it leads to lethality of the animal. At intermediate expression levels effects can be more subtle, such as an increase in the number of apoptotic cells, an influence on cell shape or the physiological response to external stimuli. Fortunately, relatively low levels of Cas9 expression are sufficient for highly efficient gene editing and are well tolerated in flies [19]. It is therefore important to optimize Cas9 expression levels to balance activity and toxicity.
Expression levels of Cas9 from UAS transgenes will depend on a number of factors: the strength of the Gal4 driver, the sequence of the UAS vector, and the insertion site of the transgene in the genome. We have recently developed a series of UAS-Cas9 transgenes that uses upstream open reading frames (uORFs) between the UAS regulatory region and the Cas9 coding sequence to predictably modulate the levels of Cas9 translation (UAS-uCas9 series [19]). With these transgenes it is possible to titrate levels of Cas9 independent of the strength of the Gal4 line and ensure an optimal balance between high gene editing efficiency and low toxicity.
-
1.
Verify that the chromosome harboring the uCas9 transgene in your Gal4 UAS-uCas9 line is homozygous viable (provided it is not recombined with a recessive lethal Gal4 driver). All UAS-uCas9 transgenes are inserted in attP landing sites that are homozygous viable (see Note 4). Homozygous strains facilitate downstream work and indicate that Cas9 is not expressed in amounts that impair viability.
-
2.
Compare the viability, proliferation, and morphology of Cas9-expressing cells in the Gal4 UAS-uCas9 strain with a line expressing only Gal4. Viability can be tested by performing anti-cleaved Dcp-1 staining, which marks cells undergoing apoptosis. Proliferating cells can be identified by an anti-phospho Histone 3 staining. Cell morphology can be monitored by staining cells with antibodies against cell surface markers (e.g., FasIII).
3.1.2 Testing for On-Target Efficiency of Gal4 UAS-uCas9 Lines
The aim of the previous step was to select a line with low enough Cas9 expression to not cause any adverse effects. However, for the success of conditional gene editing it is important that Cas9 is present in sufficient amounts to efficiently mediate DNA double-strand breaks at the target locus. It is therefore also important to test, if the selected fly strains can mediate high gene editing activity.
There are multiple ways to assay Cas9-mediated mutagenesis at the target locus. The most direct and quantitative way to do this is via PCR amplification of the target locus coupled with high-throughput sequencing of the amplicons. However, such assays can be time consuming and expensive to do and are difficult to interpret when the Gal4 driver is only active in a small fraction of cells. Instead, we propose to test Cas9 activity through the induction of relevant phenotypes. While less sensitive and not directly reflecting Cas9 activity (see Subheading 3.5 for a detailed discussion), this assay has the most practical relevance for downstream applications.
-
1.
Order several sgRNA lines targeting genes in which mutations are known to result in a clearly recognizable phenotype in the target tissue. If no such genes are known, sgRNAs targeting cell essential genes can be used and are expected to result in the death of the target cells.
-
2.
Cross transgenic sgRNA flies to Gal4 UAS-uCas9 flies and incubate crosses at 25 °C. Include a Gal4 UAS-Cas9 line with a Cas9 construct resulting in high expression levels (e.g., UAS-Cas9.P2).
-
3.
Record and compare the induced phenotype.
-
4.
The ideal Gal4 UAS-uCas9 line will induce phenotypes that are comparable to the ones induced by high-expression Cas9 constructs, but not result in non-specific phenotypes in the absence of sgRNAs.
3.1.3 Analysis of Spatial Mutagenesis Patterns
Expression of CRISPR components under the Gal4/UAS system aims to restrict mutagenesis in time and space, thereby allowing for a detailed analysis of gene function in the context of a multicellular organism. In many cases researchers will already have Gal4 lines in the lab, which they know to express at a given developmental stage in a desired pattern. However, using such lines to control expression of Cas9 will in some cases lead to mutations also in other cells and tissues. This is mainly due to two reasons: First, transgenes downstream of UAS promoters can be expressed at very low levels also in the absence of Gal4 (so-called leaky expression). Such low level expression can be sufficient for Cas9-mediated mutagenesis. Second, CRISPR gene editing leads to permanent mutations that are passed on during mitosis to both daughter cells. As a result the animal will acquire mutations in all cell lineages that expressed Gal4 at any stage during development. Of note, both of these factors play much less of a role when Gal4 lines are used to drive expression of a fluorescent protein, a method that is typically used to establish the expression pattern of a given Gal4 line. For this reason, mutagenesis patterns with CRISPR are sometimes substantially different from expectation. It is therefore advisable to experimentally test the spatial distribution of mutagenesis, in particular, if strict spatial control is an important factor for the success of the experiment.
The easiest way to visualize gene editing activity in whole animals is by using a fluorescent reporter that marks cells with Cas9-induced mutations. Several labs have developed such systems [24,25,26]. Upon successful gene targeting the induced mutations shift the reading frame or remove or inactivate an inhibitory sequence of the fluorophore coding sequence, thereby rendering the gene functional and the cell fluorescent. The advantage is that such systems allow visualization of gene edited cells with minimal work and without prior knowledge of the mutagenesis pattern. However, typically not all editing events result in reporter activity (see Note 5). Hence, the pattern of labeled cells is variable between different animals and an underestimate of the true gene editing pattern.
It is important to note that the spatial and temporal control of CRISPR activity does not only depend on the Gal4 line and UAS-Cas9 construct but is also significantly influenced by the sgRNA expression vector. While ubiquitous sgRNA expression from U6 promoters favors ubiquitous mutagenesis, sgRNA expression from UAS plasmids leads to CRISPR activity that is largely dependent on Gal4 (see below for a detailed discussion). It is therefore imperative that CRISPR activity reporters are induced with sgRNAs expressed from the same plasmid as the sgRNAs used in later experiments.
-
1.
Order reporter lines for CRISPR activity. The sgRNA targeting the reporter must be encoded in the same expression vector as will be used later on.
-
2.
Cross reporter flies to the Gal4 UAS-uCas9 line to be tested and incubate under the same conditions as to be used in later experiments.
-
3.
Visualize fluorescent cells at different levels (e.g., whole body and dissected tissues). Fluorescent cells indicate they have undergone CRISPR mutagenesis. Make sure to analyze several animals and aggregate results, as stochastically some edited cells will not activate the reporter (see Note 5).
3.2 Selection of Transgenic sgRNA Lines
sgRNAs determine at which genomic locus Cas9 will cut the DNA, leading to mutations after imprecise DNA repair. Tissue-specific CRISPR screens use collections of transgenic sgRNAs stably integrated into the Drosophila genome. Several labs have created such large-scale collections and several thousand of them are now available from public Drosophila stock centers [19, 20, 22]. Should these libraries not yet contain lines targeting genes of interest users can generate new sgRNA lines themselves, which is relatively straightforward and has been previously described [12] (see Note 6).
sgRNAs are generally designed to target unique sequences in the genome with few if any highly homologous sequences elsewhere, thereby reducing the likelihood of off-target mutagenesis. Furthermore, target sites are chosen such that sgRNAs direct Cas9 to the coding sequence of protein-coding genes (see Note 7). Mutations that change the reading frame have a high chance to disrupt gene function, but functional in-frame mutations can also occur and reduce the phenotypic penetrance of CRISPR mutagenesis (see below).
The different sgRNA resources differ mainly in the choice of promoter used for sgRNA expression and the number of sgRNAs targeting each gene (Fig. 2). Commonly used promoters include RNA polymerase III promoters of the U6 ribosomal RNAs and the UAS-hsp70 promoter present in most UAS vectors. There are three U6 genes in Drosophila and early work has shown that gene editing efficiency is affected by which promoter is used [18]. The U6:3 promoter mediates editing with high efficiency, while U6:2 is significantly less active and U6:1 has intermediate activity. All U6 promoters mediate ubiquitous expression of sgRNAs. In contrast the UAS-hsp70 promoter cassette is dependent on Gal4 and can therefore be expressed in a tissue-specific manner. However, UAS-hsp70 is a RNA polymerase II promoter and results in the transcription of mRNAs, which are capped, polyadenylated and exported into the cytoplasm, which is not compatible with sgRNA activity [12, 27]. To overcome this problem sgRNAs expressed from UAS vectors are flanked by tRNAs, which mediate excision of mature sgRNAs by the endogenous tRNA processing machinery [12, 13].
The different expression strategies have complementary strengths and weaknesses. U6 promoters, and in particular U6:3, reliably mediate high CRISPR-Cas9 activity, but in our experience frequently also mediate mutagenesis in cells that do not express Gal4, due to leaky Cas9 expression. In contrast, sgRNAs expressed from UAS-hsp70 are largely dependent on Gal4 (see Note 8) and in combination with UAS-Cas9 constructs result in much tighter restriction of mutagenesis to Gal4-expressing cells [12]. However, sgRNA levels can become limiting when these constructs are combined with weak Gal4 drivers [20].
The second aspect that differentiates publicly available sgRNA transgenes is the number of sgRNAs encoded in each line. Currently, almost all lines encode either one or two sgRNAs targeting the same gene. It has been shown that multiplexing sgRNAs leads to higher mutagenesis efficiency and more penetrant phenotypes [12]. This is because additional sgRNAs can compensate for sgRNAs with low efficiency and inducing independent mutations in the same gene increases the chances of abrogating gene function and can lead to the induction of larger deletions. However, the use of multiple sgRNAs also increases the risk of off-target mutagenesis and phenotypes caused by a general response to DNA damage.
3.2.1 The Heidelberg CRISPR Fly Design Library (Boutros Lab)
The lines of the Heidelberg CRISPR Fly Design (HD_CFD) library encode two sgRNAs in the pCFD6 expression vector and have been described in Port et al., 2020 [19]. pCFD6 is a UAS vector and mutagenesis with this library has been observed to be well restricted to Gal4-expressing cells. Each sgRNA pair targets sites preferentially located in the 5′ half of the coding sequence and usually spaced apart by approximately 500 bp. Characterization of a random sample of sgRNA lines indicated that the large majority of sgRNA lines is active and most lines generate indels at both target sites. Deletions between target sites also occur, but are less frequent. Phenotypic analysis of a large number of HD_CFD lines further supported the high efficiency of this library, with 80–90% of lines giving rise to the expected phenotype.
The HD_CFD library contains mostly sgRNAs targeting transcription factors, kinases, and phosphatases, as well as a number of genes with human homologs implicated in disease. There are currently about 2000 HD_CFD lines available from the VDRC.
3.2.2 The TRIP Knockout Lines (Perrimon Lab)
The Transgenic RNAi Project (TRiP) is producing sgRNA lines for gene activation (TRiP-OE) and gene knockouts (TRiP-KO) and these resources are described in Zirin et al., 2020 [22]. Since this chapter focuses on CRISPR loss-of-function screens, we will only describe the TRiP-KO library. This library currently consists of over 2000 transgenic lines, the majority of which encodes single sgRNAs in pCFD3. This vector expresses sgRNAs under the control of the strong, ubiquitous U6:3 promoter. A smaller proportion of lines express two sgRNAs per gene encoded in pCFD4, which uses two different U6 promoters to drive expression of each sgRNA. The TRiP has announced that production of TRiP-KO lines has been shifted to using the pCFD6 vector to improve the Gal4 dependency of mutagenesis [22]. The lines of the TRiP sgRNA libraries are available via the Bloomington Drosophila Stock Center (BDSC).
3.2.3 The Weizmann Knockout Project Lines (Schuldiner Lab)
sgRNA lines of the Weizmann knockout project encode two sgRNAs per gene and over 300 lines are currently available from the Bloomington Drosophila Stock Center. This resource is described in Meltzer et al. 2019 [20]. sgRNAs are expressed from either pCFD4 or pCFD5, which both mediate ubiquitous expression of sgRNAs. An evaluation of these tools in the nervous system suggested that sgRNAs expressed from pCFD5 have higher efficiency [20]. This might be due to the fact that in this vector both sgRNAs are expressed from the most efficient U6:3 promoter and are flanked by tRNAs. Targets covered by the WKO library are enriched for genes implicated in neuronal development, but also encompass genes involved in other biological processes. The efficiency of this library has been successfully demonstrated through a screen in the Drosophila mushroom body [20].
3.2.4 The NIG sgRNA Lines (Kondo and Ueda Lab)
The fly facility at the National Institute of Genetics in Japan has created a large collection of transgenic sgRNA strains mainly targeting genes on the second chromosome. The experimental strategy has been previously described [16], but so far no report of a larger-scale screen with these lines has been published. The lines are mainly intended to create germline alleles. They can also be combined with a conditional Cas9 source for somatic mutagenesis, but genetic mosaicism is expected to be particularly pronounced, since they encode a single sgRNA expressed under the weakest U6 promoter.
3.2.5 Other Publicly Available sgRNA Collections
At the time of writing the Bloomington Drosophila Stock Center also distributed 99 sgRNA lines generated by the Rhagu lab targeting genes implicated in the regulation of phosphoinositides [23]. These encode two sgRNAs per gene expressed from a U6:2 promoter. The target sites are located at the beginning and end of the coding sequence of each gene. Deletions between the target sites are therefore deleting the entire gene, creating null mutations. However, in cases where such deletions do not occur, individual indels at the downstream target site have a relatively low chance of disrupting gene function.
In addition, the BDSC also distributes 54 sgRNA lines generated by the Han lab [21, 26]. These encode single sgRNAs expressed from the U6:3 promoter.
3.3 Controls and Experimental Conditions
3.3.1 Negative Controls
The goal of CRISPR screening is to reveal phenotypes induced by mutations in the target gene. However, a number of other factors have the potential to induce phenotypes independent of on-target mutagenesis. Such factors include toxicity associated with excessive amounts of Cas9 (as discussed above) and the cellular response to the DNA damage that is caused by Cas9. The goal of negative controls is to detect such effects:
-
1.
Order sgRNA lines targeting genes not expressed in the target tissue or known to not be involved in the process of interest. We do not recommend the use of “non-targeting” sgRNAs, as they do not induce DNA damage and, therefore, miss an often important contribution to non-specific phenotypes.
-
2.
Cross sgRNA flies to Gal4 UAS-uCas9 flies and incubate under the same conditions as envisioned for the screen. Raise in parallel non-transgenic flies and animals that only harbor sgRNA transgenes or Gal4 UAS-uCas9 transgenes.
-
3.
Compare the phenotype of animals expressing an active CRISPR system to animals harboring only individual components or are not transgenic.
3.3.2 Positive Controls
Positive controls will confirm or refute that a method, here conditional CRISPR-Cas9 mutagenesis, is suitable to uncover genes that play a role in the process of interest. To this end, mutagenesis is performed with sgRNAs targeting genes already known to be involved and where the phenotype upon conditional loss of function is established. Occasionally, when performing particularly innovative screens, no such prior knowledge might be available. In such cases, it is advisable to perform mutagenesis with sgRNAs targeting genes that are essential to cell survival or other recognizable phenotypes, to establish that efficient mutagenesis can be performed under the chosen conditions:
-
1.
Order sgRNA lines targeting genes known to be involved in the process of interest and where a phenotype of gene knockout is known or can be inferred.
-
2.
Cross sgRNA lines to Gal4 UAS-uCas9 lines and incubate under the same conditions as planned for the screen.
-
3.
Compare the phenotype of animals expressing the positive-control sgRNAs to animals expressing negative-control sgRNAs.
3.3.3 Experimental Conditions
A relatively low number of sgRNA lines that are typically used as positive and negative controls make it feasible to test different experimental conditions to find the optimal set-up for a screen. Parameters that should be tested include the number of flies in each vial, whether vials need to be flipped at a certain time point and the temperature at which the crosses are incubated. Furthermore, the exact time point at which the phenotype is recorded needs to be established, with a longer time between induction of CRISPR mutagenesis and analysis favoring stronger phenotypes. It is also advisable to test the variation between replicates of the same cross and to establish if differential phenotypes are observed in male and female animals.
3.3.4 Pilot Screen
Once the experimental conditions are established, a pilot screen should be performed to further optimize the screening workflow. While experiments with positive and negative controls will give information about the feasibility of a screen, only a pilot screen with a larger number of sgRNAs will reveal the breath and variability of phenotypes and highlight potential problems that might arise once experiments are done at large scale:
-
1.
Order a large selection of sgRNA lines. These will typically be several tens to a few hundred lines. Include lines targeting genes that you know or suspect might be involved in the process of interest, some that are highly unlikely to be involved and a number of genes that represent a random selection of the genes you plan to test in a future screen.
-
2.
Cross sgRNA lines to Gal4 UAS-uCas9 lines and incubate under the same conditions as planned for the screen.
-
3.
Establish a reliable system for setting up and tracking each cross during the screen, recording of the phenotype, and analysis of the data.
3.4 Confounding Factors
While conditional CRISPR mutagenesis is now well established and has been found to often yield better results than previous technologies, there remain some factors that can occasionally lead one astray. These can be broadly classified as false-negative and false-positive results.
3.4.1 Effects That Can Give Rise to False-Negative Results
3.4.1.1 Low Mutagenesis Efficiency
In some cases, animals expressing Cas9 and a gene-specific sgRNA will not or only infrequently acquire mutations at the target locus. The most likely cause is inactive sgRNAs. It is long established that sgRNAs vary in their potency to guide and activate Cas9. While there is a large literature on sgRNA activity prediction, the resulting algorithms have often been found to be poor predictors of sgRNA activity in Drosophila [28]. This is likely due to the fact that such models are typically based on data from transient assays performed in mammalian cells, which differ in key parameters from transgenic CRISPR systems in vivo. Furthermore, genomic variation, rather than sgRNA activity per se, can be the basis of low on-target mutagenesis, due to polymorphisms at the target site. As a result, it is currently not possible to reliably predict sgRNAs libraries consisting of only highly active sgRNAs.
Another factor that has been shown to limit CRISPR mutagenesis activity is chromatin context. Target sites in heterochromatic regions of the genome are less accessible to Cas9, although some mutagenesis has been detected also at such sites. Since chromatin is dynamic and differs between cell types and states it is expected that activity of the same sgRNA will vary depending on where and when it is used.
3.4.1.2 mRNA and Protein Stability
After successful bi-allelic CRISPR mutagenesis only mutant copies of mRNA are transcribed. However, existing functional copies of mRNA will remain for some time and continue to be translated. The translated protein will also persist in the cell and perform its function until it is degraded. As a result at what point mutant phenotypes can be observed will depend on the level of preexisting mRNA and protein and their decay kinetics. This can be problematic in early stages of development or for experimental designs in which the CRISPR system is only activated shortly before phenotypes are recorded. Furthermore, actively dividing cells are likely to acquire phenotypes faster, as preexisting mRNA and protein is rapidly diluted in the growing cell population.
3.4.1.3 Silent Mutations and Genetic Compensation
Conditional CRISPR mutagenesis is based on error-prone DSB repair by non-homologous end joining, which typically leads to a variety of mutations at the target site, most prominently small indels. Often not all such mutations impact gene function. For example, in-frame mutations are often silent, as many proteins tolerate the deletion or insertion of a few amino acids at many positions. Furthermore, research in other organisms has established that even the impact of out-of-frame mutations can sometimes be suppressed, by mechanisms, such as alternative splicing, translational initiation at downstream start codons and upregulation of genes with overlapping functions [6, 7].
3.4.1.4 Genetic Mosaics
While sgRNA lines that completely fail to mediate on-target mutagenesis are rare, so are lines that lead to bi-alleleic knockout of the target gene in all cells. In the majority of cases CRISPR mutagenesis with one or two sgRNAs leads to genetic mosaics, in which cells that have been independently edited by Cas9 carry different mutations (Fig. 3). Some of these cells carry silent mutations or non-mutated alleles. This heterogeneity can attenuate phenotypes or lead to significant variation in observed phenotypes.
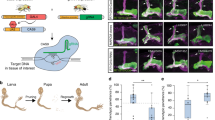
Potential limitations of conditional CRISPR experiments and strategies to avoid them. (a) CRISPR mutagenesis can in some cases be observed in broader pattern than the known Gal4 expression domain. This can be due to broad Gal4 expression in early development and Gal4 expression below the detection limit of other methods. To mitigate such effects Cas9 lines without excessive expression levels should be used. Gal4 expression can be further suppressed with temperature-dependent Gal80. (b) Mutations can be observed in cells that do not express Gal4. Ectopic mutations are the result of low level “leaky” expression of Cas9 in combination with sufficient amounts of sgRNA. To mitigate this effect tRNA-flanked sgRNAs can be expressed from the UAS vector pCFD6, which substantially improves Gal4 dependency of mutagenesis. (c) Conditional CRISPR mutagenesis frequently gives rise to genetic mosaics, which comprise cells with one, two, and no functional alleles of the target gene. These are typically caused by the induction of functional in-frame mutations and sgRNAs with low activity. To increase the frequency of bi-allelic knockout cells sgRNAs can be multiplexed to induce multiple mutations in the same gene
3.4.2 Effects That Can Give Rise to False-Positive Results
3.4.2.1 Off-Target Mutagenesis
The natural function of CRISPR-Cas9 as an adaptive immune system targeting quickly evolving pathogens means that it is unlikely to have evolved absolute target specificity. It is therefore not surprising that Cas9 can tolerate a certain degree of mismatches between the target and the sgRNA protospacer. In the context of genome editing this can lead to unwanted off-target effects, where Cas9 induces DSBs at loci elsewhere in the genome with incomplete homology to the sgRNA. It has been shown that the number of mismatches that can be tolerated are sgRNA specific and not easily predicted [29]. Unfortunately, a genome-wide assessment of off-target effects across many sgRNAs in Drosophila is currently lacking. However, results from large-scale experiments utilizing many sgRNAs suggest that off-target effects are not so common that they would lead to a high number of false-positive results [19, 20]. Nevertheless, it is imperative to exclude a causal role of off-target mutations in follow-up experiments (see below).
3.4.2.2 Large Deletions and Genomic Rearrangements
While small indels seem to be the dominant form of mutations induced by CRISPR-Cas9, several others have been documented. These include large deletions at the on-target locus, genomic rearrangements, including translocations and inversions, and chromotypsis [30,31,32,33]. Importantly, most assays used to profile CRISPR-induced mutations will not detect such events, as they are typically PCR-based and these genomic aberrations delete one or both primer binding sites. This is particularly problematic in samples that also contain alleles with small indels or which are not edited. Larger genomic alterations have the potential to give rise to phenotypes not related to the target gene, for example, by deleting neighboring genes or regulatory elements that can act on distantly located genes.
3.4.2.3 Loss of Heterozygosity
It has been shown in several organisms that DSBs caused by CRISPR-Cas9 can lead to mitotic recombination, when sister chromatids from homologous chromosomes are exchanged [24, 34,35,36]. This phenomenon is particularly common in Drosophila, because the homologous chromosomes pair during mitosis. The consequence of such recombination events is that the daughter cells become homozygous for the region of the chromosome that is distal to the cut site. If this region carries recessive mutations, these can give rise to phenotypes not observed in heterozygous tissue and not related to the CRISPR target gene.
3.5 Strategies to Confirm Causality of On-Target Mutations
A CRISPR screen will give a list of sgRNA lines that are associated with phenotypes of interest. However, as detailed in the previous section in some cases, such phenotypes could represent false-positive results and it is therefore crucial to test, if there exists a causal relationship between mutations in the target gene and the observed phenotype.
3.5.1 Perform Mutagenesis with Independent sgRNA Lines
In some cases, publicly available sgRNA libraries already contain independent lines targeting the same gene. If only a single line targeting the candidate gene exists, additional lines can be generated with relatively little effort. Lines targeting different positions within the same gene differ in their protospacer sequence and are therefore very unlikely to share off-target sequences with incomplete sequence homology. Therefore, phenotypes that are shared between independent lines are unlikely to arise from mutations caused by off-target cutting of Cas9. However, independent lines could still share other off-target effects, such as those caused by loss of heterozygosity or deletion of neighboring genes. If different phenotypes are obtained with lines targeting the same gene this might indicate differences in on-target knockout efficiency or the presence of off-target effects not shared between the lines.
3.5.2 Perturb Gene with Alternative Methods
To strengthen the link between the loss of function of a gene and an observed phenotype, it is often beneficial to perturb gene expression by orthogonal methods. For example, RNAi can be used to knockdown mRNA levels or nanobodies fused to degrons can mediate protein turnover of tagged proteins [37, 38]. While shared phenotypes are a strong indication for a causal relationship between gene function and the observed phenotype, differential outcomes are more difficult to interpret. These might arise due to off-target effects of one or the other method, but often reflect the different kinetics and strength of the different perturbation methods. For example, RNAi is often limited by a substantial amount of residual mRNA expression and a failure to reproduce a phenotype with this method could simply reflect this fact.
3.5.3 Create Sequence Verified Germline Alleles
The creation of a sequence verified germline allele is an essential step during the in-depth characterization of a gene. To this end sgRNA lines can be crossed to Cas9 lines combined with a germline driver (such as nanos-Gal4). Offspring will have an active CRISPR system in the germline and can give rise to mutant progeny. Mutant alleles can be sequenced using PCR amplicons generated with primers flanking the mutation sites. Furthermore, mutant alleles can be backcrossed into a desired genetic background, which will strongly reduce the likelihood of other genetic alterations being linked to the on-target mutation. Often multiple different alleles can be generated in a single experiment and crossed together to generate trans-heterozygous animals for analysis. In cases where knockout animals are not viable, mutations can be induced directly on FRT chromosomes to combine with conditional expression of FLP recombinase to induce homozygous mutant cells in heterozygous animals.
3.5.4 Rescue the Phenotype
Rescue experiments are a powerful way to confirm the link between gene function and phenotype. If an observed phenotype is caused by the absence of a certain gene, reintroduction of that gene should rescue the phenotype. This can be done by inserting a plasmid encoding the gene into the mutant fly strain. Using UAS plasmids will allow testing where in the organism the gene needs to be expressed in order to rescue the phenotype. However, transgene expression via the Gal4/UAS system typically leads to highly unphysiological expression levels. To express a gene at physiological levels and in its natural expression pattern, exogenous DNA sequences containing all regulatory elements are required. Bacterial artificial chromosomes are one way to insert such typically large stretches of DNA into the fly genome. An alternative approach is to correct the originally created mutation though CRISPR-assisted HDR.
4 Notes
-
1.
An alternative approach is to directly express Cas9 under the control of tissue-specific regulatory elements by encoding them in the same plasmid or by inserting the Cas9 sequence into an endogenous locus by homology-directed repair [21, 26]. The advantage of this approach is that the Gal4/UAS system can be used in parallel for other purposes and that Cas9 expression is induced faster than via Gal4. However, this method often leads to mutations in undesired locations, as it does not allow for dual conditional control of Cas9 and sgRNA expression, and Cas9 expression levels cannot be easily tuned for an optimal balance between activity and toxicity.
-
2.
Information about available Gal4 UAS-Cas9 stocks are available at https://stockcenter.vdrc.at/control/library_hdcfd and https://bdsc.indiana.edu/stocks/genome_editing/triptoolbox_grna.html. We recommend use of UAS-uCas9 transgenes.
-
3.
For some applications users will need to add other transgenes in their Gal4 UAS-Cas9 stocks, such as UAS-Fluorophore to label Cas9-expressing cells (note that this is an acute label and does not cover cells that might have expressed Cas9 at earlier time points during development) and tub-Gal80ts, which can be used to inhibit Cas9 expression by temperature.
-
4.
Sometimes chromosomes will not become homozygous not because of toxicity of the transgene but because of unrecognized recessive lethal mutations present in the original stock.
-
5.
CRISPR-Cas9 mutagenesis can lead with some frequency to larger deletions at the target locus, which can delete the promoter or fluorophore reporter in such constructs. The result are mutant cells without fluorophore expression. Since these events occur stochastically, the best remedy is to average the expression pattern of the reporter over many animals.
-
6.
We recommend the use of pCFD5 for ubiquitous expression of sgRNA and pCFD6 for Gal4-dependent sgRNA expression. Step-by-step protocols for cloning of one or several sgRNA into these vectors are available under http://www.crisprflydesign.org/ and as supplementary protocol in ref. 12.
-
7.
For other applications, such as CRISPR activation or CRISPR interference, sgRNA is designed to target other locations (e.g., the transcriptional start site).
-
8.
While expression of sgRNAs from the UAS vector pCFD6 is largely dependent on the presence of Gal4, some ubiquitous expression also exists. This is likely at least in part due to the presence of sequences encoding tRNAs, which have been shown to act as RNA polymerase III promoters. As a result, some mutagenesis in non-Gal4-expressing cells is also present when sgRNAs are expressed from pCFD6, although much reduced in comparison to strong ubiquitous sgRNA vectors.
References
Nüsslein-Volhard C, Wieschaus E (1980) Mutations affecting segment number and polarity in Drosophila. Nature 287:795–801
Boutros M, Ahringer J (2008) The art and design of genetic screens: RNA interference. Nat Rev Genet 9:554–566
St Johnston D (2002) The art and design of genetic screens: Drosophila melanogaster. Nat Rev Genet 3:176–188
Pickar-Oliver A, Gersbach CA (2019) The next generation of CRISPR–Cas technologies and applications. Nat Rev Mol Cell Biol 20:490–507
Jinek M et al (2012) A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science 337:816–821
El-Brolosy MA et al (2019) Genetic compensation triggered by mutant mRNA degradation. Nature 568:193–197
Smits AH et al (2019) Biological plasticity rescues target activity in CRISPR knock outs. Nat Methods 16:1087–1093
Mou H et al (2017) CRISPR/Cas9-mediated genome editing induces exon skipping by alternative splicing or exon deletion. Genome Biol 18:108
Cui Y, Xu J, Cheng M, Liao X, Peng S (2018) Review of CRISPR/Cas9 sgRNA design tools. Interdiscip Sci 10:455–465
Molla KA, Yang Y (2020) Predicting CRISPR/Cas9-induced mutations for precise genome editing. Trends Biotechnol 38:136–141
Michlits G et al (2020) Multilayered VBC score predicts sgRNAs that efficiently generate loss-of-function alleles. Nat Methods 17:708–716
Port F, Bullock SL (2016) Augmenting CRISPR applications in Drosophila with tRNA-flanked sgRNAs. Nat Methods 13:852–854
Xie K, Minkenberg B, Yang Y (2015) Boosting CRISPR/Cas9 multiplex editing capability with the endogenous tRNA-processing system. Proc Natl Acad Sci U S A 112:3570–3575
Gratz SJ et al (2013) Genome engineering of Drosophila with the CRISPR RNA-guided Cas9 nuclease. Genetics 194:1029–1035
Bassett AR, Tibbit C, Ponting CP, Liu J-L (2013) Highly efficient targeted mutagenesis of Drosophila with the CRISPR/Cas9 system. Cell Rep 4:220–228
Kondo S, Ueda R (2013) Highly improved gene targeting by germline-specific Cas9 expression in Drosophila. Genetics 195:715–721
Port F, Bullock SL (2016) Creating heritable mutations in drosophila with CRISPR-Cas9. Methods Mol Biol 1478:145–160
Port F, Chen H-M, Lee T, Bullock SL (2014) Optimized CRISPR/Cas tools for efficient germline and somatic genome engineering in Drosophila. Proc Natl Acad Sci U S A 111:E2967–E2976
Port F et al (2020) A large-scale resource for tissue-specific CRISPR mutagenesis in Drosophila. elife 9:e53865
Meltzer H et al (2019) Tissue-specific (ts)CRISPR as an efficient strategy for in vivo screening in Drosophila. Nat Commun 10:2113
Poe AR et al (2019) Robust CRISPR/Cas9-mediated tissue-specific mutagenesis reveals gene redundancy and perdurance in drosophila. Genetics 211:459–472
Zirin J et al (2020) Large-scale transgenic drosophila resource collections for loss- and gain-of-function studies. Genetics 214:755–767
Trivedi D et al (2020) A genome engineering resource to uncover principles of cellular organization and tissue architecture by lipid signaling. elife 9:e55793
Brunner E et al (2019) CRISPR-induced double-strand breaks trigger recombination between homologous chromosome arms. Life Sci Alliance 2(3):e201800267
Garcia-Marques J et al (2019) Unlimited genetic switches for cell-type-specific manipulation. Neuron 104:227–238, e7
Koreman GT et al (2021) Upgraded CRISPR/Cas9 tools for tissue-specific mutagenesis in Drosophila. Proc Natl Acad Sci U S A 118:e2014255118
Ren X et al (2013) Optimized gene editing technology for Drosophila melanogaster using germ line-specific Cas9. Proc Natl Acad Sci U S A 110:19012–19017
Port F, Muschalik N, Bullock SL (2015) Systematic evaluation of Drosophila CRISPR tools reveals safe and robust alternatives to autonomous gene drives in basic research. G3 (Bethesda) 5:1493–1502
Ren X et al (2014) Enhanced specificity and efficiency of the CRISPR/Cas9 system with optimized sgRNA parameters in Drosophila. Cell Rep 9:1151–1162
Leibowitz ML et al (2021) Chromothripsis as an on-target consequence of CRISPR–Cas9 genome editing. Nat Genet 53:895–905
Papathanasiou S et al (2021) Whole chromosome loss and genomic instability in mouse embryos after CRISPR-Cas9 genome editing. Nat Commun 12:5855
Kosicki M, Tomberg K, Bradley A (2018) Repair of double-strand breaks induced by CRISPR-Cas9 leads to large deletions and complex rearrangements. Nat Biotechnol 36:765–771
Shin HY et al (2017) CRISPR/Cas9 targeting events cause complex deletions and insertions at 17 sites in the mouse genome. Nat Commun 8:15464
Allen SE et al (2021) Versatile CRISPR/Cas9-mediated mosaic analysis by gRNA-induced crossing-over for unmodified genomes. PLoS Biol 19:e3001061
Sadhu MJ, Bloom JS, Day L, Kruglyak L (2016) CRISPR-directed mitotic recombination enables genetic mapping without crosses. Science 352:1113–1116
Alanis-Lobato G et al (2021) Frequent loss of heterozygosity in CRISPR-Cas9–edited early human embryos. Proc Natl Acad Sci U S A 118:e2004832117
Heigwer F, Port F, Boutros M (2018) RNA interference (RNAi) screening in drosophila. Genetics 208:853–874
Caussinus E, Kanca O, Affolter M (2012) Fluorescent fusion protein knockout mediated by anti-GFP nanobody. Nat Struct Mol Biol 19:117–121
Acknowledgments
The authors would like to thank Jun Zhou for helpful discussions and comments on the manuscript; Claudia Strein, Mona Stricker, and other members of the CRISPR Fly Design team in the Boutros lab for generating and characterizing CRISPR tools; Erich Brunner, Ben Ewen-Campden, Stephanie Mohr, Jonathan Zirin, and Norbert Perrimon for sharing unpublished information; and Lisa Meadows, Reinhard Klug, and the VDRC for distributing fly lines. Work in the laboratory of M.B. is in part supported by the European Research Council (ERC DECODE) and the DFG (SFB/TRR186, SFB1324).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Port, F., Boutros, M. (2022). Tissue-Specific CRISPR-Cas9 Screening in Drosophila. In: Dahmann, C. (eds) Drosophila. Methods in Molecular Biology, vol 2540. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2541-5_7
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2541-5_7
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2540-8
Online ISBN: 978-1-0716-2541-5
eBook Packages: Springer Protocols