Abstract
By temporarily distorting the DNA double helix, the moving RNA polymerases can lead to the formation of non-B DNA structures. One of the most abundant and largest non-B DNA structures in the genome is the R-loop, a three-stranded structure forming when the nascent RNA hybridizes with its DNA template, thereby extruding the non-template DNA strand. Growing evidence suggests that at least a subset of R-loops could induce transcription stress and genome instability, although the direct, primary consequences of R-loop formation on the surrounding chromatin are still unclear.
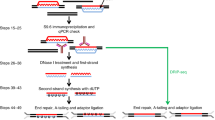
To understand the direct impact of R-loops on transcription and genome stability, accurate and quantitative mapping of R-loops is essential. R-loop mapping is commonly achieved using the antibody-based DNA:RNA Immunoprecipitation (DRIP) strategy. While it is reasonably straightforward to obtain robust DRIP enrichments from human cells, this has proved harder in yeast, where DRIP signals are often relatively weak, with a poor signal-to-noise ratio. Although it is unclear whether such weak signals stem from a technical or a biological reality, they make the accurate quantification of DRIP signals all the more important, especially when deep sequencing is used to monitor and quantify the distribution of R-loops genome-wide. Here we propose a DRIP protocol that has been optimized for the mapping and the quantification of R-loops in Schizosaccharomyces pombe but that can also be used in Saccharomyces cerevisiae. As a result, this protocol can be used to generate calibrated DRIP-seq data, where genomic DNA extracted from S. cerevisiae serves as spike-in reference.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Crossley MP, Bocek M, Cimprich KA (2019) R-loops as cellular regulators and genomic threats. Mol Cell 73:398–411
Crossley MP, Bocek MJ, Hamperl S, Swigut T, Cimprich KA (2020) qDRIP: a method to quantitatively assess RNA-DNA hybrid formation genome-wide. Nucleic Acids Res 48:e84
Malig M, Hartono SR, Giafaglione JM, Sanz LA, Chedin F (2020) Ultra-deep coverage single-molecule R-loop footprinting reveals principles of R-loop formation. J Mol Biol 432:2271–2288
Chédin F, Hartono SR, Sanz LA, Vanoosthuyse V (2021) Best practices for the visualization, mapping, and manipulation of R-loops. EMBO J 40:e106394
Castillo-Guzman D, Chédin F (2021) Defining R-loop classes and their contributions to genome instability. DNA Repair. https://doi.org/10.1016/j.dnarep.2021.103182
Sanz LA, Chédin F (2019) High-resolution, strand-specific R-loop mapping via S9.6-based DNA-RNA immunoprecipitation and high-throughput sequencing. Nat Protoc 14:1734–1755
Legros P, Malapert A, Niinuma S, Bernard P, Vanoosthuyse V (2014) RNA processing factors Swd2.2 and Sen1 antagonize RNA Pol III-dependent transcription and the localization of condensin at Pol III genes. PLoS Genet 10:e1004794
Hartono SR, Malapert A, Legros P, Bernard P, Chédin F, Vanoosthuyse V (2018) The affinity of the S9.6 antibody for double-stranded RNAs impacts the accurate mapping of R-loops in fission yeast. J Mol Biol 430:272–284
Flor-Parra I, Zhurinsky J, Bernal M, Gallardo P, Daga RR (2014) A Lallzyme MMX-based rapid method for fission yeast protoplast preparation. Yeast 31:61–66
Carrasco-Salas Y, Malapert A, Sulthana S, Molcrette B, Chazot-Franguiadakis L, Bernard P, Chédin F, Faivre-Moskalenko C, Vanoosthuyse V (2019) The extruded non-template strand determines the architecture of R-loops. Nucleic Acids Res 47:6783–6795
Wahba L, Costantino L, Tan FJ, Zimmer A, Koshland D (2016) S1-DRIP-seq identifies high expression and polyA tracts as major contributors to R-loop formation. Genes Dev 30:1327–1338
Crossley MP, Brickner JR, Song C, Zar SMT, Maw SS, Chédin F, Tsai M-S, Cimprich KA (2021) Catalytically inactive, purified RNase H1: a specific and sensitive probe for RNA-DNA hybrid imaging. J Cell Biol 220:e202101092
Nelson JD, Denisenko O, Bomsztyk K (2006) Protocol for the fast chromatin immunoprecipitation (ChIP) method. Nat Protoc 1:179–185
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Vachez, L., Teste, C., Vanoosthuyse, V. (2022). DNA:RNA Immunoprecipitation from S. pombe Cells for qPCR and Genome-Wide Sequencing. In: Aguilera, A., Ruzov, A. (eds) R-Loops . Methods in Molecular Biology, vol 2528. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2477-7_27
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2477-7_27
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2476-0
Online ISBN: 978-1-0716-2477-7
eBook Packages: Springer Protocols