Abstract
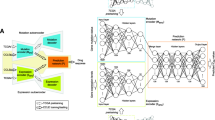
The prediction of the cancer cell lines sensitivity to a specific treatment is one of the current challenges in precision medicine. With omics and pharmacogenomics data being available for over 1000 cancer cell lines, several machine learning and deep learning algorithms have been proposed for drug sensitivity prediction. However, deciding which omics data to use and which computational methods can efficiently incorporate data from different sources is the challenge which several research groups are working on. In this review, we summarize recent advances in the representative computational methods that have been developed in the last 2 years on three public datasets: COSMIC, CCLE, NCI-60. These methods aim to improve the prediction of the cancer cell lines sensitivity to a given treatment by incorporating drug’s chemical information in the input or using a priori feature selection. Finally, we discuss the latest published method which aims to improve the prediction of clinical drug response of real patients starting from cancer cell line molecular profiles.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Reuter JA, Spacek DV, Snyder MP (2015) High-throughput sequencing technologies. Mol Cell 58:586–597
Iorio F, Knijnenburg TA, Vis DJ et al (2016) A landscape of pharmacogenomic interactions in cancer. Cell 166:740–754
Garnett MJ, Edelman EJ, Heidorn SJ et al (2012) Systematic identification of genomic markers of drug sensitivity in cancer cells. Nature 483:570–575
Azuaje F (2017) Computational models for predicting drug responses in cancer research. Brief Bioinform 18:820–829
Menden MP, AstraZeneca-Sanger Drug Combination DREAM Consortium, Wang D et al (2019) Community assessment to advance computational prediction of cancer drug combinations in a pharmacogenomic screen. Nat Commun 10
Huang C, Mezencev R, McDonald JF, Vannberg F (2017) Open source machine-learning algorithms for the prediction of optimal cancer drug therapies. PLoS One 12:e0186906
Weinstein JN (2012) Drug discovery: cell lines battle cancer. Nature 483:544–545
Wilding JL, Bodmer WF (2014) Cancer cell lines for drug discovery and development. Cancer Res 74:2377–2384
Yamori T (2003) Panel of human cancer cell lines provides valuable database for drug discovery and bioinformatics. Cancer Chemother Pharmacol 52:74–79
Barretina J, Caponigro G, Stransky N et al (2012) The cancer cell line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483:603–607
Forbes SA, Beare D, Boutselakis H et al (2017) COSMIC: somatic cancer genetics at high-resolution. Nucleic Acids Res 45:D777–D783
Alley MC, Scudiero DA, Monks A et al (1988) Feasibility of drug screening with panels of human tumor cell lines using a microculture tetrazolium assay. Cancer Res 48:589–601
Shoemaker RH (2006) The NCI60 human tumour cell line anticancer drug screen. Nat Rev Cancer 6:813–823
Nguyen T, Nguyen GTT, Nguyen T, Le D-H (2021) Graph convolutional networks for drug response prediction. bioRxiv. 2020.04.07.030908
Blessie EC, Chandra Blessie E, Karthikeyan E (2012) Sigmis: a feature selection algorithm using correlation based method. J Algorithms Computl Technol 6:385–394
Parca L, Pepe G, Pietrosanto M et al (2019) Modeling cancer drug response through drug-specific informative genes. Sci Rep 9:15222
Sánchez-Maroño N, Caamaño-Fernández M, Castillo E, Alonso-Betanzos A (2006) Functional networks and analysis of variance for feature selection. Intell Data Eng Autom Learn 2006:1031–1038
Chang Y, Park H, Yang H-J et al (2018) Cancer drug response profile scan (CDRscan): a deep learning model that predicts drug effectiveness from cancer genomic signature. Sci Rep 8:8857
Chiu Y-C, Chen H-IH, Zhang T et al (2019) Predicting drug response of tumors from integrated genomic profiles by deep neural networks. BMC Med Genet 12:18
Li M, Wang Y, Zheng R et al (2021) DeepDSC: a deep learning method to predict drug sensitivity of cancer cell lines. IEEE/ACM Trans Comput Biol Bioinform 18:575–582
Ali M, Aittokallio T (2019) Machine learning and feature selection for drug response prediction in precision oncology applications. Biophys Rev 11:31–39
Gillet J-P, Varma S, Gottesman MM (2013) The clinical relevance of cancer cell lines. J Natl Cancer Inst 105:452–458
Geeleher P, Cox NJ, Huang R (2014) Clinical drug response can be predicted using baseline gene expression levels and in vitro drug sensitivity in cell lines. Genome Biol 15:R47
Huang H-H, Dai J-G, Liang Y (2018) clinical drug response prediction by using a Lq penalized network-constrained logistic regression method. Cell Physiol Biochem 51:2073–2084
Liu P, Li H, Li S, Leung K-S (2019) Improving prediction of phenotypic drug response on cancer cell lines using deep convolutional network. BMC Bioinformatics 20:408
Moughari FA, Eslahchi C (2020) Author correction: ADRML: anticancer drug response prediction using manifold learning. Sci Rep 10:22360
Ma Y, Fu Y (2011) Manifold learning theory and applications. CRC Press
Wang JJ-Y, Huang JZ, Sun Y, Gao X (2015) Feature selection and multi-kernel learning for adaptive graph regularized nonnegative matrix factorization. Expert Syst Appl 42:1278–1286
Wei D, Liu C, Zheng X, Li Y (2019) Comprehensive anticancer drug response prediction based on a simple cell line-drug complex network model. BMC Bioinformatics 20:44
Wang L, Li X, Zhang L, Gao Q (2017) Improved anticancer drug response prediction in cell lines using matrix factorization with similarity regularization. BMC Cancer 17:513
Suphavilai C, Bertrand D, Nagarajan N (2018) Predicting cancer drug response using a recommender system. Bioinformatics 34:3907–3914
Garreta R, Moncecchi G (2013) Learning scikit-learn: machine Learning in Python. Packt Publishing Ltd.
Koras K, Juraeva D, Kreis J et al (2020) Feature selection strategies for drug sensitivity prediction. Sci Rep 10:9377
Kursa MB, Jankowski A, Rudnicki WR (2010) Boruta – a system for feature selection. Fundamenta Inform 101:271–285
Xu X, Gu H, Wang Y et al (2019) Autoencoder based feature selection method for classification of anticancer drug response. Front Genet 10:233
Dong Z, Zhang N, Li C et al (2015) Anticancer drug sensitivity prediction in cell lines from baseline gene expression through recursive feature selection. BMC Cancer 15:489
Emdadi A, Eslahchi C (2021) Auto-HMM-LMF: feature selection based method for prediction of drug response via autoencoder and hidden Markov model. BMC Bioinformatics 22:33
Neumann U, Genze N, Heider D (2017) EFS: an ensemble feature selection tool implemented as R-package and web-application. BioData Min 10:21
Ahmed KT, Park S, Jiang Q et al (2020) Network-based drug sensitivity prediction. BMC Med Genet 13:193
An B, Zhang Q, Fang Y et al (2020) Iterative sure independent ranking and screening for drug response prediction. BMC Med Inform Decis Mak 20:224
Zhu L, Li L, Li R, Zhu L (2011) Model-free feature screening for ultrahigh dimensional data. J Am Stat Assoc 106:1464–1475
Fang Y, Qin Y, Zhang N et al (2015) DISIS: prediction of drug response through an iterative sure independence screening. PLoS One 10:e0120408
Majumder B, Baraneedharan U, Thiyagarajan S et al (2015) Predicting clinical response to anticancer drugs using an ex vivo platform that captures tumour heterogeneity. Nat Commun 6(1):1–14
Ding Z, Zu S, Gu J (2016) Evaluating the molecule-based prediction of clinical drug responses in cancer. Bioinformatics 32:2891–2895
Turki T, Wei Z, Wang JTL (2018) A transfer learning approach via procrustes analysis and mean shift for cancer drug sensitivity prediction. J Bioinforma Comput Biol 16:1840014
Huang EW, Bhope A, Lim J et al (2020) Tissue-guided LASSO for prediction of clinical drug response using preclinical samples. PLoS Comput Biol 16:e1007607
Johnson WE, Evan Johnson W, Li C Adjusting batch effects in microarray experiments with small sample size using empirical Bayes methods. Batch Effects Noise Microarray Exp:113–129
Marangoni E, Poupon M-F (2014) Patient-derived tumour xenografts as models for breast cancer drug development. Curr Opin Oncol 26:556–561
Tentler JJ, Tan AC, Weekes CD et al (2012) Patient-derived tumour xenografts as models for oncology drug development. Nat Rev Clin Oncol 9:338–350
Weeber F, Ooft SN, Dijkstra KK, Voest EE (2017) Tumor organoids as a pre-clinical cancer model for drug discovery. Cell Chem Biol 24:1092–1100
Rae C, Amato F, Braconi C (2021) Patient-derived organoids as a model for cancer drug discovery. Int J Mol Sci 22:3483
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Pepe, G., Carrino, C., Parca, L., Helmer-Citterich, M. (2022). Dissecting the Genome for Drug Response Prediction. In: Carugo, O., Eisenhaber, F. (eds) Data Mining Techniques for the Life Sciences. Methods in Molecular Biology, vol 2449. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2095-3_7
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2095-3_7
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2094-6
Online ISBN: 978-1-0716-2095-3
eBook Packages: Springer Protocols