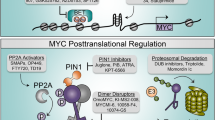
Abstract
Oncoproteins encoded by dominant oncogenes have long been considered as targets for chemotherapeutic intervention. However, oncogenic transcription factors have often been dismissed as “undruggable.” Members of the Myc family of transcription factors have been identified as promising targets for cancer chemotherapy in multiple publications reporting the requirement of Myc proteins for maintenance of almost every type of tumor. Here, we describe cell-based approaches to identify c-Myc small molecule inhibitors by screening complex libraries of diverse small molecules based on Myc functionality and specificity.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Dang CV, O’Donnell KA, Zeller KI, Nguyen T, Osthus RC, Li F (2006) The c-Myc target gene network. Semin Cancer Biol 16:253–264
Dang CV (2012) MYC on the path to cancer. Cell 149:22–35
Malynn BA, de Alboran IM, O’Hagan RC, Bronson R, Davidson L, DePinho RA, Alt FW (2000) N-myc can functionally replace c-myc in murine development, cellular growth, and differentiation. Genes Dev 14:1390–1399
Hermeking H (2003) The MYC oncogene as a cancer drug target. Curr Cancer Drug Targets 3:163–175
Arvanitis C, Felsher DW (2006) Conditional transgenic models define how MYC initiates and maintains tumorigenesis. Semin Cancer Biol 16:313–317
Schulte JH, Lindner S, Bohrer A, Maurer J, De Preter K, Lefever S, Heukamp L, Schulte S, Molenaar J, Versteeg R, Thor T, Künkele A, Vandesompele J, Speleman F, Schorle H, Eggert A, Schramm A (2012) MYCN and ALKF1174L are sufficient to drive neuroblastoma development from neural crest progenitor cells. Oncogene 10. [Epub ahead of print]
Soucek L, Whitfield J, Martins CP, Finch AJ, Murphy DJ, Sodir NM, Karnezis AN, Swigart LB, Nasi S, Evan GI (2008) Modelling Myc inhibition as a cancer therapy. Nature 455:679–683
Carabet LA, Rennie PS, Cherkasov A (2018) Therapeutic inhibition of Myc in cancer structural bases and computer-aided drug discovery approaches. Int J Mol Sci 20:120
Allen-Petersen BL, Sears RC (2019) Mission possible: advances in MYC therapeutic targeting in cancer. BioDrugs 33:539–553
Mateyak MK, Obaya AJ, Adachi S, Sedivy JM (1999) Phenotypes of c-Myc-deficient rat fibroblasts isolated by targeted homologous recombination. Cell Growth Differ 8:1039–1048
Wang H, Mannava S, Grachtchouk V, Zhuang D, Soengas MS, Gudkov AV, Prochownik EV, Nikiforov MA (2008) c-Myc depletion inhibits proliferation of human tumor cells at various stages of the cell cycle. Oncogene 27:1905–1915
Acknowledgments
This work was supported by NIH R01 CA120244 and ACS RSG-10-121-01 grants to M.A.N., and by CINSW and NHMRC grants to M.H. and M.D.N.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Burkhart, C.A., Haber, M., Norris, M.D., Gudkov, A.V., Nikiforov, M.A. (2021). Cell-Based Methods for the Identification of Myc-Inhibitory Small Molecules. In: Soucek, L., Whitfield, J. (eds) The Myc Gene. Methods in Molecular Biology, vol 2318. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1476-1_19
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1476-1_19
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1475-4
Online ISBN: 978-1-0716-1476-1
eBook Packages: Springer Protocols