Abstract
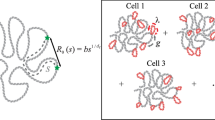
In the absence of a clear molecular understanding of the mechanism that stabilizes specific contacts in interphasic chromatin, we resort to the principle of maximum entropy to build a polymeric model based on the Hi-C data of the specific system one wants to study. The interactions are set by an iterative Monte Carlo algorithm to reproduce the average contacts summarized by the Hi-C map. The study of the ensemble of conformations generated by the algorithm can report a much richer set of information than the experimental map alone, including colocalization of multiple sites, fluctuations of the contacts, and kinetical properties.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Dekker J, Rippe K, Dekker M, Kleckner N (2002) Capturing chromosome conformation. Science 295:1306–1311
Lieberman-Aiden E, van Berkum NL, Williams L et al (2009) Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326:289–293
Giorgetti L, Galupa R, Nora EP et al (2014) Predictive polymer modeling reveals coupled fluctuations in chromosome conformation and transcription. Cell 157:950–963
Nagano T, Lubling Y, Stevens TJ et al (2013) Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 502:59–64
Tiana G, Amitai A, Pollex T et al (2016) Structural fluctuations of the chromatin fiber within topologically associating domains. Biophys J 110:1234–1245
Jost D, Carrivain P, Cavalli G, Vaillant C (2014) Modeling epigenome folding: formation and dynamics of topologically associated chromatin domains. Nucleic Acids Res 42:9553–9561
Barbieri M, Chotalia M, Fraser J et al (2012) Complexity of chromatin folding is captured by the strings and binders switch model. Proc Natl Acad Sci U S A 109:16173–16178
Benedetti F, Dorier J, Burnier Y, Stasiak A (2013) Models that include supercoiling of topological domains reproduce several known features of interphase chromosomes. Nucleic Acids Res 42:2848–2855
Brackley CA, Taylor S, Papantonis A et al (2013) Nonspecific bridging-induced attraction drives clustering of DNA-binding proteins and genome organization. Proc Natl Acad Sci U S A 110:E3605–E3611
Brackley CA, Johnson J, Michieletto D et al (2017) Nonequilibrium chromosome looping via molecular slip links. Phys Rev Lett 119:138101
Fudenberg G, Imakaev M, Lu C et al (2016) Formation of chromosomal domains by loop extrusion. Cell Rep 15:2038–2049
Davidson IF, Bauer B, Goetz D et al (2019) DNA loop extrusion by human cohesin. Science 366:1338–1345. https://doi.org/10.1126/science.aaz3418
Kim Y, Shi Z, Zhang H et al (2019) Human cohesin compacts DNA by loop extrusion. Science 366(6471):1345–1349. https://doi.org/10.1126/science.aaz4475
Jaynes ET (1957) Information theory and statistical mechanics. Phys Rev 106:620–630
Redolfi J, Zhan Y, Valdes-Quezada C et al (2019) DamC reveals principles of chromatin folding in vivo without crosslinking and ligation. Nat Struct Mol Biol 26:471–480
Zhan Y, Giorgetti L, Tiana G (2017) Modelling genome-wide topological associating domains in mouse embryonic stem cells. Chromosom Res 25:5–14
Tiana G, Giorgetti L (2018) Integrating experiment, theory and simulation to determine the structure and dynamics of mammalian chromosomes. Curr Opin Struct Biol 49:11–17
Tiana G, Villa F, Zhan Y et al (2014) MonteGrappa: an iterative Monte Carlo program to optimize biomolecular potentials in simplified models. Comput Phys Commun 186:93–104
Norgaard AB, Ferkinghoff-Borg J, Lindorff-Larsen K (2008) Experimental parameterization of an energy function for the simulation of unfolded proteins. Biophys J 94:182–192
Olivares-Chauvet P, Mukamel Z, Lifshitz A et al (2016) Capturing pairwise and multi-way chromosomal conformations using chromosomal walks. Nature 540:296–300
Ferrenberg A, Swendsen R (1989) Optimized Monte Carlo data analysis. Phys Rev Lett 63:1195–1198
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Zhan, Y., Giorgetti, L., Tiana, G. (2022). Polymer Folding Simulations from Hi-C Data. In: Bicciato, S., Ferrari, F. (eds) Hi-C Data Analysis. Methods in Molecular Biology, vol 2301. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1390-0_13
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1390-0_13
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1389-4
Online ISBN: 978-1-0716-1390-0
eBook Packages: Springer Protocols