Abstract
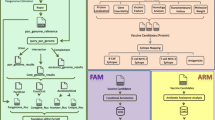
There is still a lack of vaccines for many bacterial infections for which the best treatment option would be a prophylactic one. On the other hand, effectiveness has been questioned for some existing vaccines, prompting new developments. Therapeutic vaccines are also becoming a treatment option in specific cases where antibiotics tend to fail. In this scenario, refinement and extension of the classical reverse vaccinology approach is allowing scientists to find new and more effective antigens. In this chapter, we describe an in silico methodology that integrates pangenomic, immunoinformatic, structural, and evolutionary approaches for the screening of potential antigens in a given bacterial species. The strategy focuses on targeting relatively conserved epitopes in core proteins to design broadly cross-protective vaccines and avoid allele-specific immunity. The proposed methodological steps and computational tools can be easily implemented in a reverse vaccinology approach not only to identify new leads with strong immune response but also to develop diagnostic assays.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Levine MM, Dougan G, Good MF, Liu MA, Nabel GJ, Nataro JP, Rappuoli R (2017) New generation vaccines, 4th edn. Taylor & Francis Group, Boca Raton, FL
Bidmos FA, Siris S, Gladstone CA, Langford PR (2018) Bacterial vaccine antigen discovery in the reverse vaccinology 2.0 era: progress and challenges. Front Immunol 9:2315. https://doi.org/10.3389/fimmu.2018.02315
Tettelin H, Masignani V, Cieslewicz MJ et al (2005) Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: implications for the microbial “pan-genome.”. PNAS 102:13950–13955
Vernikos G, Medini D, Riley DR, Tettelin H (2015) Ten years of pan-genome analyses. Curr Opin Microbiol 23:148–154
Rappuoli R (2000) Reverse vaccinology. Curr Opin Microbiol 3:445–450
Pizza M, Scarlato V, Masignani V et al (2000) Identification of vaccine candidates against serogroup B meningococcus by whole-genome sequencing. Science 287:1816–1820
Moriel DG, Scarselli M, Serino L, Mora M, Rappuoli R, Masignani V (2008) Genome-based vaccine development: a short cut for the future. Hum Vaccin 4:184–188
Moriel DG, Tan L, Goh KGK, Phan M-D, Ipe DS, Lo AW, Peters KM, Ulett GC, Beatson SA, Schembri MA (2016) A novel protective vaccine antigen from the core Escherichia coli genome. mSphere 1(6). https://doi.org/10.1128/mSphere.00326-16
Moriel DG, Bertoldi I, Spagnuolo A et al (2010) Identification of protective and broadly conserved vaccine antigens from the genome of extraintestinal pathogenic Escherichia coli. PNAS 107:9072–9077
Ariel N, Zvi A, Makarova KS, Chitlaru T, Elhanany E, Velan B, Cohen S, Friedlander AM, Shafferman A (2003) Genome-based bioinformatic selection of chromosomal Bacillus anthracis putative vaccine candidates coupled with proteomic identification of surface-associated antigens. Infect Immun 71:4563–4579
Amela I, Cedano J, Querol E (2007) Pathogen proteins eliciting antibodies do not share epitopes with host proteins: a bioinformatics approach. PLoS One 2:e512
Barh D, Tiwari S, Jain N, Ali A, Santos AR, Misra AN, Azevedo V, Kumar A (2011) In silico subtractive genomics for target identification in human bacterial pathogens. Drug Dev Res 72:162–177
Rappuoli R, Pizza M, Masignani V, Vadivelu K (2018) Meningococcal B vaccine (4CMenB): the journey from research to real world experience. Exp Rev Vaccin 17:1111–1121
Giuliani MM, Adu-Bobie J, Comanducci M et al (2006) A universal vaccine for serogroup B meningococcus. Proc Natl Acad Sci U S A 103:10834–10839
Tomar N, De RK (2014) Immunoinformatics: a brief review. Methods Mol Biol 1184:23–55
Capelli R, Peri C, Villa R et al (2018) BPSL1626: reverse and structural vaccinology reveal a novel candidate for vaccine design against Burkholderia pseudomallei. Antibodies 7:26
Pajon R, Yero D, Niebla O et al (2009) Identification of new meningococcal serogroup B surface antigens through a systematic analysis of neisserial genomes. Vaccine 28:532–541
He Y, Xiang Z, Mobley HLT (2010) Vaxign: the first web-based vaccine design program for reverse vaccinology and applications for vaccine development. J Biomed Biotechnol 2010:297505. https://doi.org/10.1155/2010/297505
Naz K, Naz A, Ashraf ST, Rizwan M, Ahmad J, Baumbach J, Ali A (2019) PanRV: pangenome-reverse vaccinology approach for identifications of potential vaccine candidates in microbial pangenome. BMC Bioinformatics 20:123. https://doi.org/10.1186/s12859-019-2713-9
Solanki V, Tiwari V (2018) Subtractive proteomics to identify novel drug targets and reverse vaccinology for the development of chimeric vaccine against Acinetobacter baumannii. Sci Rep 8:9044
Jaiswal V, Chanumolu SK, Gupta A, Chauhan RS, Rout C (2013) Jenner-predict server: prediction of protein vaccine candidates (PVCs) in bacteria based on host-pathogen interactions. BMC Bioinformatics 14:211
Zvi A, Rotem S, Bar-Haim E, Cohen O, Shafferman A (2011) Whole-genome immunoinformatic analysis of F. tularensis: predicted CTL epitopes clustered in hotspots are prone to elicit a T-cell response. PLoS One 6:e20050
Solanki V, Tiwari M, Tiwari V (2019) Prioritization of potential vaccine targets using comparative proteomics and designing of the chimeric multi-epitope vaccine against Pseudomonas aeruginosa. Sci Rep 9:1. https://doi.org/10.1038/s41598-019-41496-4
Dhanda SK, Usmani SS, Agrawal P, Nagpal G, Gautam A, Raghava GPS (2017) Novel in silico tools for designing peptide-based subunit vaccines and immunotherapeutics. Brief Bioinform 18:467–478
Sanchez-Trincado JL, Gomez-Perosanz M, Reche PA (2017) Fundamentals and methods for T- and B-cell epitope prediction. J Immunol Res 2017:2680160. https://doi.org/10.1155/2017/2680160
Fleri W, Paul S, Dhanda SK, Mahajan S, Xu X, Peters B, Sette A (2017) The immune epitope database and analysis resource in epitope discovery and synthetic vaccine design. Front Immunol 8:278. https://doi.org/10.3389/fimmu.2017.00278
Vita R, Mahajan S, Overton JA, Dhanda SK, Martini S, Cantrell JR, Wheeler DK, Sette A, Peters B (2019) The immune epitope database (IEDB): 2018 update. Nucleic Acids Res 47:D339–D343
Cozzi R, Scarselli M, Ferlenghi I (2013) Structural vaccinology: a three-dimensional view for vaccine development. Curr Top Med Chem 13:2629–2637
Anasir MI, Poh CL (2019) Structural vaccinology for viral vaccine design. Front Microbiol 10:738. https://doi.org/10.3389/fmicb.2019.00738
Nuccitelli A, Rinaudo CD, Brogioni B et al (2013) Understanding the molecular determinants driving the immunological specificity of the protective pilus 2a backbone protein of group B streptococcus. PLoS Comput Biol 9:e1003115
Capelli R, Marchetti F, Tiana G, Colombo G (2017) SAGE: a fast computational tool for linear epitope grafting onto a foreign protein scaffold. J Chem Inf Model 57:6–10
Raeven RHM, van Riet E, Meiring HD, Metz B, Kersten GFA (2019) Systems vaccinology and big data in the vaccine development chain. Immunology 156:33–46
Oberg AL, Kennedy RB, Li P, Ovsyannikova IG, Poland GA (2011) Systems biology approaches to new vaccine development. Curr Opin Immunol 23:436–443
De Groot AS, Ardito M, Moise L, Gustafson EA, Spero D, Tejada G, Martin W (2011) Immunogenic consensus sequence T helper epitopes for a Pan-Burkholderia biodefense vaccine. Immunome Res 7(2). https://doi.org/10.4172/1745-7580.1000043
Goodswen SJ, Kennedy PJ, Ellis JT (2014) Enhancing in silico protein-based vaccine discovery for eukaryotic pathogens using predicted peptide-MHC binding and peptide conservation scores. PLoS One 9:e115745. https://doi.org/10.1371/journal.pone.0115745
Seib KL, Zhao X, Rappuoli R (2012) Developing vaccines in the era of genomics: a decade of reverse vaccinology. Clin Microbiol Infect 18(Suppl 5):109–116
Goodswen SJ, Kennedy PJ, Ellis JT (2018) A gene-based positive selection detection approach to identify vaccine candidates using Toxoplasma gondii as a test case protozoan pathogen. Front Genet 9:332. https://doi.org/10.3389/fgene.2018.00332
Gilchuk P, Hill TM, Wilson JT, Joyce S (2015) Discovering protective CD8 T cell epitopes—no single immunologic property predicts it! Curr Opin Immunol 34:43–51
Akram A, Inman RD (2012) Immunodominance: a pivotal principle in host response to viral infections. Clin Immunol 143:99–115
Rodrigues MM, Ersching J (2015) Neglected tropical diseases, bioinformatics, and vaccines. J Infect Dis 211:175–177
Haft DH, DiCuccio M, Badretdin A et al (2018) RefSeq: an update on prokaryotic genome annotation and curation. Nucleic Acids Res 46:D851–D860
Harrison PW, Alako B, Amid C et al (2019) The european nucleotide archive in 2018. Nucleic Acids Res 47:D84–D88
Wattam AR, Davis JJ, Assaf R et al (2017) Improvements to PATRIC, the all-bacterial Bioinformatics Database and Analysis Resource Center. Nucleic Acids Res 45:D535–D542
Fu L, Niu B, Zhu Z, Wu S, Li W (2012) CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28:3150–3152
Fischer S, Brunk BP, Chen F, Gao X, Harb OS, Iodice JB, Shanmugam D, Roos DS, Stoeckert CJ (2011) Using OrthoMCL to assign proteins to OrthoMCL-DB groups or to cluster proteomes into new ortholog groups. Curr Protoc Bioinformatics Chapter 6:Unit 6.12.1-19
Train C-M, Glover NM, Gonnet GH, Altenhoff AM, Dessimoz C (2017) Orthologous Matrix (OMA) algorithm 2.0: more robust to asymmetric evolutionary rates and more scalable hierarchical orthologous group inference. Bioinformatics 33:i75–i82
Callister SJ, McCue LA, Turse JE, Monroe ME, Auberry KJ, Smith RD, Adkins JN, Lipton MS (2008) Comparative bacterial proteomics: analysis of the core genome concept. PLoS One 3:e1542. https://doi.org/10.1371/journal.pone.0001542
The UniProt Consortium (2017) UniProt: the universal protein knowledgebase. Nucleic Acids Res 45:D158–D169
GenBank Sample Record. https://www.ncbi.nlm.nih.gov/Sitemap/samplerecord.html. Accessed 27 Jun 2019
Yu NY, Wagner JR, Laird MR et al (2010) PSORTb 3.0: improved protein subcellular localization prediction with refined localization subcategories and predictive capabilities for all prokaryotes. Bioinformatics 26:1608–1615
Krogh A, Larsson B, von Heijne G, Sonnhammer EL (2001) Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J Mol Biol 305:567–580
Prilusky J, Felder CE, Zeev-Ben-Mordehai T, Rydberg EH, Man O, Beckmann JS, Silman I, Sussman JL (2005) FoldIndex: a simple tool to predict whether a given protein sequence is intrinsically unfolded. Bioinformatics 21:3435–3438
Conchillo-Solé O, de Groot NS, Avilés FX, Vendrell J, Daura X, Ventura S (2007) AGGRESCAN: a server for the prediction and evaluation of “hot spots” of aggregation in polypeptides. BMC Bioinformatics 8:65
Magnan CN, Baldi P (2014) SSpro/ACCpro 5: almost perfect prediction of protein secondary structure and relative solvent accessibility using profiles, machine learning and structural similarity. Bioinformatics 30:2592–2597
Dhanda SK, Mahajan S, Paul S et al (2019) IEDB-AR: immune epitope database-analysis resource in 2019. Nucleic Acids Res 47:W502. https://doi.org/10.1093/nar/gkz452
Jurtz V, Paul S, Andreatta M, Marcatili P, Peters B, Nielsen M (2017) NetMHCpan-4.0: improved peptide-MHC class i interaction predictions integrating eluted ligand and peptide binding affinity data. J Immunol 199:3360–3368
Jensen KK, Andreatta M, Marcatili P, Buus S, Greenbaum JA, Yan Z, Sette A, Peters B, Nielsen M (2018) Improved methods for predicting peptide binding affinity to MHC class II molecules. Immunology 154:394–406
Karosiene E, Lundegaard C, Lund O, Nielsen M (2012) NetMHCcons: a consensus method for the major histocompatibility complex class I predictions. Immunogenetics 64:177–186
Peri C, Solé OC, Corrada D, Gori A, Daura X, Colombo G (2015) Prediction of antigenic B and T cell epitopes via energy decomposition analysis: description of the web-based prediction tool BEPPE. Methods Mol Biol 1348:13–22
Fiorucci S, Zacharias M (2010) Prediction of protein-protein interaction sites using electrostatic desolvation profiles. Biophys J 98:1921–1930
Rice P, Longden I, Bleasby A (2000) EMBOSS: the European Molecular Biology Open Software Suite. Trends Genet 16:276–277
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549
Garcia-Boronat M, Diez-Rivero CM, Reinherz EL, Reche PA (2008) PVS: a web server for protein sequence variability analysis tuned to facilitate conserved epitope discovery. Nucleic Acids Res 36:W35–W41
Rozas J, Ferrer-Mata A, Sánchez-DelBarrio JC, Guirao-Rico S, Librado P, Ramos-Onsins SE, Sánchez-Gracia A (2017) DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol Biol Evol 34:3299–3302
Xu B, Yang Z (2013) PAMLX: a graphical user interface for PAML. Mol Biol Evol 30:2723–2724
Pond SLK, Frost SDW, Muse SV (2005) HyPhy: hypothesis testing using phylogenies. Bioinformatics 21:676–679
Weaver S, Shank SD, Spielman SJ, Li M, Muse SV, Kosakovsky Pond SL (2018) Datamonkey 2.0: a modern web application for characterizing selective and other evolutionary processes. Mol Biol Evol 35:773. https://doi.org/10.1093/molbev/msx335
Baum BR (1989) PHYLIP: phylogeny inference package. Version 3.2. Joel Felsenstein. Q Rev Biol 64:539–541
Guindon S, Dufayard J-F, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321
Kane TL, Carothers KE, Lee SW (2018) Virulence factor targeting of the bacterial pathogen Staphylococcus aureus for vaccine and therapeutics. Curr Drug Targets 19:111–127
The Gene Ontology Consortium (2019) The gene ontology resource: 20 years and still GOing strong. Nucleic Acids Res 47:D330-D338
El-Gebali S, Mistry J, Bateman A et al (2019) The Pfam protein families database in 2019. Nucleic Acids Res 47:D427–D432
Mitchell AL, Attwood TK, Babbitt PC et al (2019) InterPro in 2019: improving coverage, classification and access to protein sequence annotations. Nucleic Acids Res 47:D351–D360
Carroll MC (2008) Complement and Humoral Immunity. Vaccine 26:I28–I33
Backert L, Kohlbacher O (2015) Immunoinformatics and epitope prediction in the age of genomic medicine. Genome Med 7:119
Gonzalez-Galarza FF, McCabe A, Melo Dos Santos EJ, Takeshita L, Ghattaoraya G, Jones AR, Middleton D (2018) Allele frequency net database. Methods Mol Biol 1802:49–62
HLA allele frequencies and reference sets with maximal population coverage. IEDB Solutions Center. http://help.iedb.org/hc/en-us/articles/114094151851-HLA-allele-frequencies-and-reference-sets-with-maximal-population-coverage. Accessed 26 Jun 2019
Greenbaum J, Sidney J, Chung J, Brander C, Peters B, Sette A (2011) Functional classification of class II human leukocyte antigen (HLA) molecules reveals seven different supertypes and a surprising degree of repertoire sharing across supertypes. Immunogenetics 63:325–335
Weiskopf D, Angelo MA, de Azeredo EL et al (2013) Comprehensive analysis of dengue virus-specific responses supports an HLA-linked protective role for CD8+ T cells. Proc Natl Acad Sci U S A 110:E2046–E2053
Lassaux P, Peri C, Ferrer-Navarro M et al (2013) A structure-based strategy for epitope discovery in Burkholderia pseudomallei OppA antigen. Structure 21:167–175
Gourlay LJ, Peri C, Ferrer-Navarro M et al (2013) Exploiting the Burkholderia pseudomallei acute phase antigen BPSL2765 for structure-based epitope discovery/design in structural vaccinology. Chem Biol 20:1147–1156
Gourlay LJ, Thomas RJ, Peri C et al (2015) From crystal structure to in silico epitope discovery in the Burkholderia pseudomallei flagellar hook-associated protein FlgK. FEBS J 282:1319–1333
Gaudesi D, Peri C, Quilici G et al (2015) Structure-based design of a B cell antigen from B. pseudomallei. ACS Chem Biol 10:803–812
van Gunsteren WF, Bakowies D, Baron R et al (2006) Biomolecular modeling: goals, problems, perspectives. Angew Chem Int Ed Eng 45:4064–4092
Telford JL (2008) Bacterial genome variability and its impact on vaccine design. Cell Host Microbe 3:408–416
Khan AM, Hu Y, Miotto O, Thevasagayam NM, Sukumaran R, Abd Raman HS, Brusic V, Tan TW, Thomas August J (2017) Analysis of viral diversity for vaccine target discovery. BMC Med Genet 10:78
Graves CJ, Ros VID, Stevenson B, Sniegowski PD, Brisson D (2013) Natural selection promotes antigenic evolvability. PLoS Pathog 9:e1003766. https://doi.org/10.1371/journal.ppat.1003766
Xu Z, Chen H, Zhou R (2011) Genome-wide evidence for positive selection and recombination in Actinobacillus pleuropneumoniae. BMC Evol Biol 11:203
Cao P, Guo D, Liu J, Jiang Q, Xu Z, Qu L (2017) Genome-wide analyses reveal genes subject to positive selection in Pasteurella multocida. Front Microbiol 8:961. https://doi.org/10.3389/fmicb.2017.00961
Petersen L, Bollback JP, Dimmic M, Hubisz M, Nielsen R (2007) Genes under positive selection in Escherichia coli. Genome Res 17:1336–1343
Nandi T, Ong C, Singh AP et al (2010) A genomic survey of positive selection in Burkholderia pseudomallei provides insights into the evolution of accidental virulence. PLoS Pathog 6:e1000845
O’Connor DH, McDermott AB, Krebs KC et al (2004) A dominant role for CD8+-T-lymphocyte selection in simian immunodeficiency virus sequence variation. J Virol 78:14012–14022
de Oliveira T, Salemi M, Gordon M, Vandamme A-M, van Rensburg EJ, Engelbrecht S, Coovadia HM, Cassol S (2004) Mapping sites of positive selection and amino acid diversification in the HIV genome: an alternative approach to vaccine design? Genetics 167:1047–1058
Yang Z (1997) PAML: a program package for phylogenetic analysis by maximum likelihood. Comput Appl Biosci 13:555–556
Kosakovsky Pon S, Poon A, Frost S (2009) Estimating selection pressures on alignments of coding sequences. In: The phylogenetic handbook: a practical approach to phylogenetic analysis and hypothesis testing, 2nd edn. Cambridge University Press, New York, NY, pp 419–452
Anisimova M, Nielsen R, Yang Z (2003) Effect of recombination on the accuracy of the likelihood method for detecting positive selection at amino acid sites. Genetics 164:1229–1236
Kosakovsky Pond SL, Posada D, Gravenor MB, Woelk CH, Frost SDW (2006) GARD: a genetic algorithm for recombination detection. Bioinformatics 22:3096–3098
Dobson R, Stockdale C, Lapsley C, Wilkes J, McCulloch R (2011) Interactions among Trypanosoma brucei RAD51 paralogues in DNA repair and antigenic variation. Mol Microbiol 81:434–456
Holden NJ, Uhlin BE, Gally DL (2001) PapB paralogues and their effect on the phase variation of type 1 fimbriae in Escherichia coli. Mol Microbiol 42:319–330
Tsuru T, Kobayashi I (2008) Multiple genome comparison within a bacterial species reveals a unit of evolution spanning two adjacent genes in a tandem paralog cluster. Mol Biol Evol 25:2457–2473
Rappuoli R (2011) The challenge of developing universal vaccines. F1000 Med Rep 3:16. https://doi.org/10.3410/M3-16
Bambini S, Piet J, Muzzi A et al (2013) An analysis of the sequence variability of meningococcal fHbp, NadA and NHBA over a 50-year period in the Netherlands. PLoS One 8:e65043
Wachter J, Hill S (2016) Positive selection pressure drives variation on the surface-exposed variable proteins of the pathogenic neisseria. PLoS One 11:e0161348. https://doi.org/10.1371/journal.pone.0161348
Counoupas C, Pinto R, Nagalingam G, Hill-Cawthorne GA, Feng CG, Britton WJ, Triccas JA (2016) Mycobacterium tuberculosis components expressed during chronic infection of the lung contribute to long-term control of pulmonary tuberculosis in mice. NPJ Vaccines 1:16012
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Yero, D., Conchillo-Solé, O., Daura, X. (2021). Antigen Discovery in Bacterial Panproteomes. In: Pfeifer, B.A., Hill, A. (eds) Vaccine Delivery Technology. Methods in Molecular Biology, vol 2183. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-0795-4_5
Download citation
DOI: https://doi.org/10.1007/978-1-0716-0795-4_5
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-0794-7
Online ISBN: 978-1-0716-0795-4
eBook Packages: Springer Protocols