Abstract
Background
Recurrence represents a well-known poor prognostic factor for colorectal cancer (CRC) patients. This study aimed to establish an effective prognostic prediction model based on noninvasive circulating tumor DNA methylation markers for CRC patients receiving radical surgery.
Results
Two methylation markers (cg11186405 and cg17296166) were identified by Cox regression and receiver operating characteristics, which could classify CRC patients into high recurrence risk and low recurrence risk group. The 3-year disease-free survival was significantly different between CRC patients with low and high recurrence risk [Training set: hazard ratio (HR) 28.776, 95% confidence interval (CI) 3.594–230.400; P = 0.002; Validation set: HR 7.796, 95% CI 1.425–42.660, P = 0.018]. The nomogram based on the above two methylation markers and TNM stage was established which demonstrated robust prognostic prediction potential, as evidenced by the decision curve analysis result.
Conclusions
A cell-free DNA methylation model consisting of two DNA methylation markers is a promising method for prognostic prediction in CRC patients.
Similar content being viewed by others
Background
Colorectal cancer (CRC) is the third leading malignancy and the second most common cause of global cancer-related death [1]. Radical surgery is the cornerstone of current therapeutic modalities for localized advanced CRC patients, which has improved their clinical outcomes dramatically. Unfortunately, about a quarter of CRC patients develop recurrence after radical surgery [2, 3]. Worse yet, CRC patients with recurrence have an inferior prognosis, with 60% of these patients suffering from death within 2 years [4].
The combination of imaging examination and serum tumor markers, such as carcinoembryonic antigen (CEA) and carbohydrate antigen19-9 (CA19-9), remains the routine monitoring approach for recurrence detection in CRC patients after radical surgery [5,6,7,8,9,10]. The sensitivity and specificity for postoperative recurrence and metastasis detection of CEA are 58–89% and 75–98% respectively, while those of CA19-9 range from 26 to 56% and 83 to 87% separately [11, 12]. These data indicated that CEA and CA19-9 have poor performance in prognosis monitoring. Therefore, there is an urgent clinical need to improve the diagnostic efficacy of CRC recurrence after radical surgery, which can remarkably improve the clinical outcomes of these patients.
As an epigenetic mechanism, DNA methylation plays a vital role in tumorigenesis and development [13]. Abnormal methylation in tumors often results in aberrant gene expression [14]. Tumor cells with necrosis and apoptosis can release DNA harboring abnormal methylation; these extracellular nucleic acid fragments are named circulating tumor DNA (ctDNA), which can truly reflect features of cancer [15,16,17]. Plenty of studies suggest a strong link between ctDNA methylation and tumorigenesis; ctDNA methylation has been utilized as a valuable marker for the early diagnosis of CRC, which was further confirmed by our previous studies [18,19,20,21,22,23,24,25,26,27,28] 29. However, the mechanism of ctDNA methylation in regulating prognosis for CRC patients requires further investigation. This work established an effective prognostic model based on two noninvasive ctDNA methylation markers, which can predict the 3-year disease-free survival (DFS) in CRC patients after radical surgery.
Results
Patient demographics and clinical characteristics
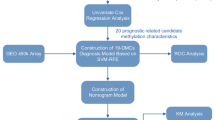
As described in our previous study [29], 248 CRC patients were enrolled between August 2016 and May 2018 to test the contribution of ctDNA methylation for early diagnosis of CRC. In the present study, the 3-year DFS and overall survival (OS) from 84 CRC patients in the training group and 56 patients in the validation group were analyzed (Fig. 1). The clinical characteristics for all patients included are summarized in Table 1. All 140 patients underwent a minimum follow-up of 3 years after radical surgery. Recurrence within 3 years was observed in 9 (10.71%) in the training group and 6 (10.71%) in the validation group. In addition, 5 (5.95%) patients and 3 (5.36%) patients died within 3 years after radical surgery in the training and validation data sets, respectively.
Limited contribution of current diagnostic biomarkers for prognostic prediction
With ctDNA methylation, our previous study has discovered valuable diagnostic markers for the early diagnosis of CRC [29]. Based on this study, we first set out to evaluate the contribution of these diagnostic markers for prognostic prediction. Time-dependent receiver operating characteristic (ROC) [30] analysis indicated that these diagnostic markers exhibited low to medium accuracy in 3-year DFS prediction (Additional file 1: Fig. S1). Moreover, Cox regression analysis was performed on these eleven methylation markers (cg11407741, cg00310855, cg15020425, cg01857475, cg11596863, cg24733262, cg22329423, cg25300584, cg01922936, cg26337020, and cg11320449) in the training group. The coefficients were used to calculate the prognostic index for 3-year DFS based on diagnostic markers (DPI) (Additional file 1: Table S1). Univariate Cox analysis showed that DPI was a significant factor related to 3-year DFS, with a hazard ratio (HR) of 7.413 [95% confidence interval (CI) 1.982–27.730, P = 0.003], and the Kaplan–Meier curves indicated that the risk of 3-year DFS was increased with higher DPI in the training group (P < 0.001). However, in the validation set, the Kaplan–Meier curves showed that DPI did not correlate with 3-year DFS (P = 0.550) (Fig. 2). Together, these results suggested that the diagnostic markers had limited contribution in recurrence prediction, highlighting an unmet clinical need to discover alternative ctDNA methylation markers for prognostic prediction.
Prognostic prediction of 3-year DFS in CRC patients based on diagnostic markers. Unsupervised hierarchical clustering of the diagnostic markers in the training (a) and validation group (b). Survival curves of CRC patients with low DPI and high DPI in the training group (c) and validation group (e). ROC curve and corresponding AUCs for 3-year DFS predicted by DPI in the training group (d) and validation group (f)
Discovery of ctDNA methylation markers for the 3-year DFS prediction
To identify ctDNA methylation markers for 3-year DFS prediction in CRC patients, a modified screening flowchart was used as previously described [31]. Based on a panel of 667 CRC-specific DNA methylation markers in our previous study [29], univariate Cox regression analysis was applied which identified 55 markers with a P value < 0.05 (Additional file 1: Table S2). Through time-dependent ROC analysis, 15 out of the 55 methylation biomarkers were identified with their receiver operating characteristic curve (AUC) greater than 0.7. To optimize the predictive model, multivariate Cox regression analysis was employed which showed that cg11186405 and cg17296166 were independent risk factors for 3-year DFS in CRC patients (Fig. 3a). Therefore, these two markers were selected for prognostic predictive model construction. The “Maxstat” package of R was used to generate the optimum cutoff scores for these two markers to classify patients into high-risk and low-risk groups. Kaplan–Meier curves manifested that the increased expression of either cg11186405 or cg17296166 was associated with poorer prognosis (Additional file 1: Table S3 and Fig. 3b, c).
Survival analysis based on selected biomarkers. a Workflow for screening methylation markers related to 3-year DFS. b–c Survival curves of CRC patients with low and high cg11186405/cg17296166 in the training group. d–i ROC curve and corresponding AUCs for 3-year DFS predicted by cg11186405, cg17296166, DFSPI, CEA and CA19-9
A new prognostic index based on 3-year DFS (DFSPI) was built using the above two markers. And time-dependent ROC analysis showed that DFSPI was better than the individual ones (Fig. 3d–f). As the routine monitoring biomarker in CRC patients, CEA, CA19-9, and a combination of these two markers both were demonstrated with poor performance in postoperative recurrence detection (Fig. 3g–i). Univariate Cox regression analysis revealed that DFSPI was significantly associated with 3-year DFS in CRC patients, with HR as 28.776 [95% CI 3.594–230.400, P = 0.002] (Table 2). Based on this cutoff, the Kaplan–Meier curve indicated that the median survival time in CRC patients with low risk was significantly longer than those with high-risk (P < 0.001) (Fig. 4a).
Establishment of prognostic prediction model based on ctDNA methylation markers. a Survival curves of CRC patients with low DFSPI or high DFSPI in the training group. b ROC and corresponding AUCs for 3-year DFS predicted by DFSPI and pTNM in the training group. c Nomogram for predicting DFS of CRC patients based on DFSPI and pTNM. d Calibration plot of nomogram for predicting three-year DFS in the training group. e Decision curve analysis of the prognostic prediction model
In addition, time-dependent ROC analysis revealed that the performance of DFSPI was better than pathological TNM stage in predicting 3-year DFS of CRC patients (AUC 0.862 vs. 0.698) (Fig. 4b). Moreover, a prediction nomogram based on DFSPI and pathological TNM stage was developed to predict DFS for CRC patients (Fig. 4c). The C-index for this prediction nomogram was 0.841, and calibration curve demonstrated good consistency between prediction and observation of the nomogram (Fig. 4d and Additional file 1: Fig. S2). Furthermore, decision curve analysis demonstrated that the nomogram performed well in predicting the 3-year DFS rate in CRC patients (Fig. 4e).
Validation of the ctDNA methylation marker-based prognostic prediction model
To confirm the prognostic value of the ctDNA methylation marker-based prognostic prediction model, we applied the two-marker classifier in the validation cohort. Results showed that 43 (76.79%) patients were of low risk and 13 (23.21%) were of high risk. Univariate Cox regression analysis indicated that DFSPI was a risk factor for 3-year DFS, with HR as 7.796 [95% CI 1.425–42.660, P = 0.018]. Kaplan–Meier curves found that DFSPI could indicate patient prognosis (P = 0.005) (Fig. 5a). Time-dependent ROC analysis revealed that DFSPI was of superior performance than pathological TNM stage (AUC 0.913 vs. 0.803) (Fig. 5b). Furthermore, the calibration curve demonstrated that the prediction nomogram based on DFSPI and TNM stage was reliable and accurate for prognostic prediction (Fig. 5c). The ctDNA methylation model was also shown to be superior to CEA and CA19-9 in the validation cohort (AUC 0.913 versus 0.675 and 0.543, respectively) (Fig. 5d). Taken together, these results showed that ctDNA methylation marker could contribute to prognostic prediction in CRC patients.
Validation of prognostic prediction model in CRC patients. a Survival curves of CRC patients separated by DFSPI in the validation group. b ROC and corresponding AUCs of DFSPI and pTNM in the validation group. c Calibration plot of nomogram for predicting 3-year DFS in the validation group. d ROC and corresponding AUCs for 3-year DFS predicted by DFSPI, CEA and CA19-9 in the validation group
ctDNA-based overall survival prediction for CRC
To further evaluate the predictive performance of ctDNA-based diagnostic markers, we used Cox regression analysis to construct a prediction model and calculate a prognostic index for 3-year OS (OPI) using the 11 diagnostic markers (Additional file 1: Table S4). Unsupervised hierarchical clustering was used to show the association between 11 diagnostic markers and OS in the training and validation cohorts (Additional file 1: Fig. S3a, b). A combination of 11 diagnostic ctDNA markers could separate CRC patients into good prognosis versus poor prognosis group based on OS data from CRC patients in the training but not validation set (Additional file 1: Fig. S3c, d). A similar pattern was found in the ROC analysis (Additional file 1: Fig. S3e, f). These findings revealed the limited contribution of diagnostic ctDNA markers for OS prediction.
In light of our above findings, we subsequently explored the potential of alternative methylation markers in predicting 3-year OS for CRC patients (Additional file 1: Table S5). Using the Cox regression model, the two-marker panel (cg05173737 and cg25224568) was used to construct a prognostic index for overall survival (OSPI) (Additional file 1: Table S6, Additional file 1: Fig. S4 and Fig. 6a). Univariate Cox regression analysis revealed that the OSPI was significantly associated with a higher risk of death within 3 years, with HR as 13.442 [95% CI 1.502–120.300, P = 0.020]. Kaplan–Meier curves showed that CRC patients with different OSPI had significantly different prognoses (training cohort: P = 0.003; validation cohort: P = 0.009) (Fig. 6b, c). Among the clinicopathological variables, the pathological T stage (P = 0.030) was significantly associated with OS in CRC patients (Table 3). Time-dependent ROC analysis showed that OSPI was better than pathological T stage in predicting the prognosis of CRC patients (training cohort: AUC, 0.821 vs. 0.782; validation cohort: AUC, 0.972 vs. 0.549) (Fig. 6d, e).
Prognostic prediction of 3-year OS based on ctDNA methylation markers. a Workflow for exploring 3-year OS related methylation markers. b–c 3-year OS curves of CRC patients with low OSPI and high OSPI in the training (b) and validation group (c). d–e ROC and corresponding AUCs for 3-year OS predicted by OSPI and pT in the training (d) and validation group (e)
Discussion
Despite current progress in cancer prognostic evaluation, the present prognostic assessment tool for CRC patients is still far from optimum due to limited efficacy. Computed tomography is routinely used for disease monitor, but it cannot detect minimal residual disease; the radiation characteristic further limits its frequent use as an instant test. These current situations highlight the need to identify new biomarkers that can effectively monitor the prognosis of CRC patients after radical surgery.
DNA methylation plays a vital role in tumorigenesis; it has been shown that ctDNA methylation status can be applied in the early diagnosis of multiple malignancies. The methylation of oncogenes and tumor suppressors presented at early stages of malignant transformation has been proved to exhibit significant values in cancer detection and diagnosis [31,32,33,34,35,36]. Our previous study [29] has established a promising non‐invasive diagnostic model based on the ctDNA methylation pattern (AUC: 0.91, sensitivity: 83.9%, specificity: 85.7%) for CRC patients. These findings suggest that DNA methylation patterns may play a pivotal role in malignancy recurrence prediction.
Considering the potential of ctDNA methylation in prognostic prediction, this work identified two markers for prognostic 3-year DFS prediction in CRC patients. According to their AUC values for 3-year DFS, it is conceivable that cg11186405 and cg17296166 might work together in CRC progression. The genomic locations of cg11186405 and cg17296166 indicate that they may affect the transcription of SOX1-OT and PNMA8A, respectively (Additional file 1: Table S7). Due to the lack of known functions of SOX1-OT and PNMA8A in CRC development, whether these two markers are connected with biological functions in CRC remains unknown. More importantly, it is of great clinical significance to explore the association between these markers and therapeutic resistance, which needs further investigation.
In this study, we constructed a prognostic prediction model (DFSPI) based on two ctDNA methylation markers (cg11186405 and cg17296166), which could stratify CRC patients with different prognoses. CEA and CA19-9 have been extensively utilized for routine surveillance in CRC patients. However, significant variations in CEA level can occur, and values exceeding 10 ng/mL can occasionally be observed without a clinically evident cause [37]. In our study, DFSPI demonstrated satisfactory performance in prognostic monitoring compared with CEA, CA19-9 and their combination (Fig. 3d–i). Consistent with previous studies [38, 39], the pathological TNM stage was identified as a predictor for 3-year DFS in this study. Moreover, the prognostic predictive power of DFSPI was superior to the pathological TNM stage. On the basis of these, a nomogram consisting of DFSPI and pathological TNM stage was constructed, which could contribute to identifying CRC patients who need more aggressive treatment and surveillance. The prediction accuracy of this nomogram was further supported by the satisfactory C-index and calibration.
Another prediction model (OSPI) was constructed using another two-marker panel (cg05173737 and cg25224568) to predict the 3-year OS in CRC patients. The OSPI was proved to be a prognostic risk factor to effectively distinguish CRC patients with different prognoses. The stratification potential of OSPI was superior to pathological T stage and conventional serum markers CEA and CA19-9 (Additional file 1: Fig. S4).
This study demonstrated that ctDNA methylation markers were associated with recurrence risk and survival in CRC patients after radical surgery. Based on these findings, a nomogram integrating pathological T stage and ctDNA methylation markers was developed to calculate the recurrence risk for each patient. This ctDNA methylation-based model is valuable for colorectal surgeons, which may help refine the postoperative recurrence risk calculation in CRC patients after radical surgery.
There are some inevitable limitations in this study. Firstly, the results of this study should be carefully interpreted due to the limited sample size. Secondly, the nomogram was established based on data from a single institution. Thirdly, a follow-up evaluation of these two markers is unavailable in the present work, and related studies are needed in future work. Further validation work from more medical centers would be ideal and necessary before the application in clinical practice.
Conclusions
In summary, our findings show that the ctDNA methylation marker-based prognostic model can effectively stratify CRC patients into low and high recurrence risk groups. This model may add prognostic values to traditional clinicopathological risk factors in prognostic evaluation for CRC patients.
Materials and methods
Detailed procedures are provided in Supplementary materials.
Patients and samples
All specimens, including plasma, fresh–frozen tissues, and formalin-fixed paraffin-embedded (FFPE) tissues were collected at The Sixth Affiliated Hospital of Sun Yat-sen University (Guangzhou, China) from August 2016 to May 2018. The methylation data were assembled from 313 tissue samples [139 normal, 30 advanced adenomas (AA), and 144 CRC] and 577 plasma samples [169 healthy controls, 44 non-advanced adenomas (NAA), 76 AA, and 288 CRC], which were generated in our previous study [29]. All procedures were performed with the approval of the Institutional Review Board of the Sixth Affiliated Hospital of Sun Yat-sen University.
Inclusion and exclusion criteria
The inclusion criteria were: (1) patients with CRC; (2) patients who underwent radical surgical resection in our center; and (3) patients' plasma undergoing DNA methylation sequencing. The exclusion criteria included: (1) patients with missing follow-up data; (2) patients with other cancers; (3) patients with distant metastases; (4) patients with follow-up < 3 years; and (5) patients with missing clinical data exceeding 20% of variables. Finally, 140 eligible CRC patients were included in this study and were randomly divided into a training group and a validation group at a 3:2 ratio.
Definition and variables
Clinical variables and demographic were defined as follows: gender, age at the time of surgery, body mass index (BMI), elevated CEA level (> 100 ug/L), elevated CA19-9 level (> 37 U/mL), and pathological TNM staging.
Statistical analysis
Frequencies were used to describe continuous factors, and means and standard deviations were applied to categorical factors. The chi-square test for categorical variables and a two-tailed t test for continuous variables were used to test baseline balance between the training and validation groups. The Cox regression model was used to analyze the correlation between ctDNA methylation markers and DFS or OS. Time-dependent ROC analysis was used to shrink the number of biomarkers. Univariate Cox analysis was applied to screen prognostic-related clinical indicators. The Cox regression model was utilized in the univariate survival analysis, and Cox regression coefficients were used for nomogram construction. Decision curve analysis was conducted to determine the effectiveness of the nomogram by quantifying the net benefits at different threshold probabilities. The P value less than 0.05 was considered statistically significant. The R software (version 4.2.1; http://www.r-project.org) was used for all analysis.
Availability of data and materials
The datasets used and analyzed during the current study are available from the corresponding author upon reasonable request.
Abbreviations
- CRC:
-
Colorectal cancer
- DFS:
-
Disease-free survival
- HR:
-
Hazard ratio
- CI:
-
Confidence interval
- CEA:
-
Carcinoembryonic antigen
- CA19-9:
-
Carbohydrate antigen19-9
- ctDNA:
-
Circulating tumor DNA
- FFPE:
-
Formalin-fixed paraffin-embedded
- AA:
-
Advanced adenoma
- NAA:
-
Non-advanced adenoma
- BMI:
-
Body mass index
- OS:
-
Overall survival
- ROC:
-
Receiver operating characteristic
- DPI:
-
Prognostic index for DFS by diagnostic markers
- AUC:
-
Area under receiver operating characteristic curve
- DFSPI:
-
Prognostic index of the 3-year DFS
- OPI:
-
Prognostic index for OS by diagnostic markers
- OSPI:
-
Prognostic index of overall survival
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424.
Guraya SY. Pattern, stage, and time of recurrent colorectal cancer after curative surgery. Clin Colorectal Cancer. 2019;18(2):e223–8.
Hashiguchi Y, Muro K, Saito Y, Ito Y, Ajioka Y, Hamaguchi T, et al. Japanese society for cancer of the colon and rectum (JSCCR) guidelines 2019 for the treatment of colorectal cancer. Int J Clin Oncol. 2020;25(1):1–42.
Lan YT, Chang SC, Yang SH, Lin CC, Wang HS, Jiang JK, et al. Comparison of clinicopathological characteristics and prognosis between early and late recurrence after curative surgery for colorectal cancer. Am J Surg. 2014;207(6):922–30.
Suzuki Y, Ogura A, Uehara K, Aiba T, Ohara N, Murata Y, et al. The carcinoembryonic antigen ratio is a potential predictor of survival in recurrent colorectal cancer. Int J Clin Oncol. 2021;26(7):1264–71.
Kim CG, Ahn JB, Jung M, Beom SH, Heo SJ, Kim JH, et al. Preoperative serum carcinoembryonic antigen level as a prognostic factor for recurrence and survival after curative resection followed by adjuvant chemotherapy in stage III colon cancer. Ann Surg Oncol. 2017;24(1):227–35.
Konishi T, Shimada Y, Hsu M, Tufts L, Jimenez-Rodriguez R, Cercek A, et al. Association of preoperative and postoperative serum carcinoembryonic antigen and colon cancer outcome. JAMA Oncol. 2018;4(3):309–15.
Auclin E, André T, Taieb J, Banzi M, Van Laethem JL, Tabernero J, et al. Association of post-operative CEA with survival and oxaliplatin benefit in patients with stage II colon cancer: a post hoc analysis of the MOSAIC trial. Br J Cancer. 2019;121(4):312–7.
Huang SH, Tsai WS, You JF, Hung HY, Yeh CY, Hsieh PS, et al. Preoperative carcinoembryonic antigen as a poor prognostic factor in stage I–III colorectal cancer after curative-intent resection: a propensity score matching analysis. Ann Surg Oncol. 2019;26(6):1685–94.
Takagawa R, Fujii S, Ohta M, Nagano Y, Kunisaki C, Yamagishi S, et al. Preoperative serum carcinoembryonic antigen level as a predictive factor of recurrence after curative resection of colorectal cancer. Ann Surg Oncol. 2008;15(12):3433–9.
Meyerhardt JA, Mayer RJ. Follow-up strategies after curative resection of colorectal cancer. Semin Oncol. 2003;30(3):349–60.
Wang WS, Lin JK, Chiou TJ, Liu JH, Fan FS, Yen CC, et al. CA19-9 as the most significant prognostic indicator of metastatic colorectal cancer. Hepatogastroenterology. 2002;49(43):160–4.
Jones PA. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat Rev Genet. 2012;13(7):484–92.
Esteller M. Epigenetics in cancer. N Engl J Med. 2008;358(11):1148–59.
Aucamp J, Bronkhorst AJ, Badenhorst CPS, Pretorius PJ. The diverse origins of circulating cell-free DNA in the human body: a critical re-evaluation of the literature. Biol Rev Camb Philos Soc. 2018;93(3):1649–83.
Francis G, Stein S. Circulating cell-free tumour DNA in the management of cancer. Int J Mol Sci. 2015;16(6):14122–42.
Stroun M, Maurice P, Vasioukhin V, Lyautey J, Lederrey C, Lefort F, et al. The origin and mechanism of circulating DNA. Ann N Y Acad Sci. 2000;906:161–8.
Alizadeh-Sedigh M, Fazeli MS, Mahmoodzadeh H, Sharif SB, Teimoori-Toolabi L. Methylation of FBN1, SPG20, ITF2, RUNX3, SNCA, MLH1, and SEPT9 genes in circulating cell-free DNA as biomarkers of colorectal cancer. Cancer Biomark. 2022;34(2):221–50.
Barault L, Amatu A, Siravegna G, Ponzetti A, Moran S, Cassingena A, et al. Discovery of methylated circulating DNA biomarkers for comprehensive non-invasive monitoring of treatment response in metastatic colorectal cancer. Gut. 2018;67(11):1995–2005.
Bedin C, Enzo MV, Del Bianco P, Pucciarelli S, Nitti D, Agostini M. Diagnostic and prognostic role of cell-free DNA testing for colorectal cancer patients. Int J Cancer. 2017;140(8):1888–98.
Cai G, Cai M, Feng Z, Liu R, Liang L, Zhou P, et al. A multilocus blood-based assay targeting circulating tumor DNA methylation enables early detection and early relapse prediction of colorectal cancer. Gastroenterology. 2021;161(6):2053–6.
Gallardo-Gómez M, Moran S, de la Páez Cadena M, Martínez-Zorzano VS, Rodríguez-Berrocal FJ, Rodríguez-Girondo M, et al. A new approach to epigenome-wide discovery of non-invasive methylation biomarkers for colorectal cancer screening in circulating cell-free DNA using pooled samples. Clin Epigenet. 2018;10:53.
Jensen SØ, Øgaard N, Ørntoft MW, Rasmussen MH, Bramsen JB, Kristensen H, et al. Novel DNA methylation biomarkers show high sensitivity and specificity for blood-based detection of colorectal cancer-a clinical biomarker discovery and validation study. Clin Epigenet. 2019;11(1):158.
Li J, Zhou X, Liu X, Ren J, Wang J, Wang W, et al. Detection of colorectal cancer in circulating cell-free DNA by methylated CpG tandem amplification and sequencing. Clin Chem. 2019;65(7):916–26.
Luo H, Zhao Q, Wei W, Zheng L, Yi S, Li G, et al. Circulating tumor DNA methylation profiles enable early diagnosis, prognosis prediction, and screening for colorectal cancer. Sci Transl Med. 2020;12(524):eaax7533.
Petit J, Carroll G, Gould T, Pockney P, Dun M, Scott RJ. Cell-free DNA as a diagnostic blood-based biomarker for colorectal cancer: a systematic review. J Surg Res. 2019;236:184–97.
Sui J, Wu X, Wang C, Wang G, Li C, Zhao J, et al. Discovery and validation of methylation signatures in blood-based circulating tumor cell-free DNA in early detection of colorectal carcinoma: a case-control study. Clin Epigenet. 2021;13(1):26.
Zhang Y, Wu Q, Xu L, Wang H, Liu X, Li S, et al. Sensitive detection of colorectal cancer in peripheral blood by a novel methylation assay. Clin Epigenet. 2021;13(1):90.
Wu X, Zhang Y, Hu T, He X, Zou Y, Deng Q, et al. A novel cell-free DNA methylation-based model improves the early detection of colorectal cancer. Mol Oncol. 2021;15(10):2702–14.
Heagerty PJ, Lumley T, Pepe MS. Time-dependent ROC curves for censored survival data and a diagnostic marker. Biometrics. 2000;56(2):337–44.
Xu RH, Wei W, Krawczyk M, Wang W, Luo H, Flagg K, et al. Circulating tumour DNA methylation markers for diagnosis and prognosis of hepatocellular carcinoma. Nat Mater. 2017;16(11):1155–61.
Chen X, Zhang J, Ruan W, Huang M, Wang C, Wang H, et al. Urine DNA methylation assay enables early detection and recurrence monitoring for bladder cancer. J Clin Invest. 2020;130(12):6278–89.
Hulbert A, Jusue-Torres I, Stark A, Chen C, Rodgers K, Lee B, et al. Early detection of lung cancer using DNA promoter hypermethylation in plasma and sputum. Clin Cancer Res. 2017;23(8):1998–2005.
Jiao X, Zhang S, Jiao J, Zhang T, Qu W, Muloye GM, et al. Promoter methylation of SEPT9 as a potential biomarker for early detection of cervical cancer and its overexpression predicts radioresistance. Clin Epigenet. 2019;11(1):120.
Molnár B, Galamb O, Kalmár A, Barták BK, Nagy ZB, Tóth K, et al. Circulating cell-free nucleic acids as biomarkers in colorectal cancer screening and diagnosis - an update. Expert Rev Mol Diagn. 2019;19(6):477–98.
Worm Ørntoft MB. Review of blood-based Colorectal cancer screening: How far are circulating cell-free DNA methylation markers from clinical implementation? Clin Colorectal Cancer. 2018;17(2):e415–33.
Denstman F, Rosen L, Khubchandani IT, Sheets JA, Stasik JJ, Riether RD. Comparing predictive decision rules in postoperative CEA monitoring. Cancer. 1986;58(9):2089–95.
Miyoshi N, Ohue M, Yasui M, Noura S, Shingai T, Sugimura K, et al. Novel prognostic prediction models for patients with stage IV colorectal cancer after concurrent curative resection. ESMO Open. 2016;1(3): e000052.
Zakaria S, Donohue JH, Que FG, Farnell MB, Schleck CD, Ilstrup DM, et al. Hepatic resection for colorectal metastases: value for risk scoring systems? Ann Surg. 2007;246(2):183–91.
Acknowledgements
We are grateful for the sincere cooperation of the patients enrolled in the study for voluntarily providing samples and clinical information.
Funding
This work was supported by the National Key R&D Program of China (No. 2017YFC1308800, 2017YFC1309002), National Natural Science Foundation of China (No. 81870383), Guangdong Natural Science Foundation (No. 2017A030313785, 2016A030310187), Science and Technology Planning Project of Guangzhou City (No. 201804010014), Science and Technology Planning Project of Guangdong Province (No. 2015B020229001, 20160916, 2017B020226005), Key-Area Research and Development Program of Guangdong Province (2019B020229002), Scheme of Guangzhou Economic and Technological Development District for Leading Talents in Innovation and Entrepreneurship (No. 2017-L152), Scheme of Guangzhou for Leading Talents in Innovation and Entrepreneurship (No.2016007), Scheme of Guangzhou for Leading Team in Innovation (No.201909010010), National Key Clinical Discipline and the program of Guangdong Provincial Clinical Research Center for Digestive Diseases (2020B1111170004), National Natural Science Foundation of China (82200569), China Postdoctoral Science Foundation (2021M703723), and Guangdong Basic and Applied Basic Research Foundation (2021A1515111011).
Author information
Authors and Affiliations
Contributions
XY, XFW, QG, and YFZ analyzed and organized the data together, while ZHL, ZXL, QW, WMR, ZJY, HW, ZWC, and JBF were responsible for acquiring and managing patients. XY wrote the main manuscript text. XRW, HSL, and PL designed the study and revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This study was approved by the ethics committee of the Sixth Affiliated Hospital of Sun Yat-sen University.
Consent for publication
Not applicable.
Competing interests
All authors declare no competing interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Yang, X., Wen, X., Guo, Q. et al. Predicting disease-free survival in colorectal cancer by circulating tumor DNA methylation markers. Clin Epigenet 14, 160 (2022). https://doi.org/10.1186/s13148-022-01383-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13148-022-01383-8