Abstract
Background
We aimed to identify differentially expressed pseudogenes and explore their potential functions in four types of common gynecological malignancies (e.g., cervical squamous cell carcinoma, ovarian serous cystadenocarcinoma, uterine corpus endometrial carcinoma, and uterine carcinosarcoma) using bioinformatics technology.
Materials and methods
We identified up-regulated and down-regulated pseudogenes and built a pseudogene-miRNA-mRNA regulatory network through public datasets to explore their potential functions in carcinogenesis and cancer prognosis.
Results
Among the 63 up-regulated pseudogenes identified, LDHAP5 demonstrated the greatest potential as a candidate pseudogene due to its significant association with poor overall survival in ovarian serous cystadenocarcinoma. KEGG pathway analysis revealed that LDHAP5 showed significant enrichment in MicroRNAs in cancer, Pathway in cancer and PI3K-AKT signaling pathway. Further analysis revealed that EGFR was the potential target mRNA of LDHAP5, which may play an important role in ovarian serous cystadenocarcinoma.
Conclusions
LDHAP5 was associated with the occurrence and prognosis of ovarian serous cystadenocarcinoma, and thus shows potential as a novel therapeutic target against such cancer.
Similar content being viewed by others
Background
Gynecological malignancies account for a large proportion of tumors in women and seriously endanger female health. It is estimated that there will be approximately 13,800 new cases of uterine cervical cancer, 65,620 cases of uterine corpus cancer, and 21,750 cases of ovarian cancer in the United States in 2020, and with 4290, 12,590 and 13,940 possible deaths, respectively [1]. Advanced gynecological malignancies usually exhibit poor prognosis due to a lack of effective treatment in controlling distant metastasis [2]. However, most current clinical drugs are non-specific, and their therapeutic effects are limited [3]. Therefore, the identification of novel biomarkers of gynecological tumors to improve drug efficacy and prolong survival remains urgent.
The term pseudogene was first conceived by Jacp et al. [4]. Pseudogenes usually originate from paralogous functional genes (“parent gene”), but have lost the capacity to encode functional proteins due to the accumulation of mutations (e.g., frameshift mutations, early or delayed stop codons) [5]. Pseudogenes initially received little attention until PTEN pseudogene 1 (PTENP1) was found to share the same microRNA response elements (MREs) as its homologous functional parent gene, PTEN [6].
With the advancement of next-generation sequencing (NGS), approximately 20,000 pseudogenes have been discovered in the human genome, and the role of pseudogenes as long non-coding RNAs (lncRNAs) in the development of disease has been revealed [7,8,9]. Current research suggests that pseudogenes mainly regulate gene expression at the post-transcriptional level through two pathways [10]. Firstly, pseudogenes can be used as competitive endogenous RNAs (ceRNAs) to competitively bind miRNAs with the coding gene, thereby positively regulating gene expression [11,12,13]. For example, PTENP1 can competitively bind miRNA-17, miRNA-21, miRNA-19, and other miRNAs through the ceRNA mechanism, thereby increasing parent gene (PTEN) expression by preventing miRNA-induced degradation [6]. Secondly, pseudogenes can play a negative role in the regulatory pathway, whereby they complete with their parent genes to destabilize RNA binding proteins (RBPs), resulting in a decrease in parent gene expression [14].
In the current study, we identified differentially expressed pseudogenes in four gynecological malignancies using the pseudogene database dreamBase, and then constructed a pseudogene-miRNA-mRNA regulatory network to further explore their potential functions and mechanisms in gynecological malignancies.
Materials and methods
Screening for dysregulated pseudogenes in four gynecological malignancies
We obtained RNA-seq data of pseudogenes in 32 human cancer from the online database dreamBase (http://rna.sysu.edu.cn/dreamBase/pancancer.php?SClade=mammal&SOrganism=hg38) [15] |Log2FC| > 2.0 was set as cutoff to identify differentially expressed pseudogenes. R v 3.5.1 and EXCEL v2016. were used to further analyze their expression landscape.
Prognostic analysis of up-regulated expressed pseudogenes
Gene Expression Profiling Interactive Analysis (GEPIA) (http://gepia.cancer-pku.cn/) was used to evaluate prognostic values (overall survival) of up-regulated pseudogenes in 32 kinds of common human cancer [16]. The group thresholds were as follows: the group cut-off was ‘Median’, the ‘cutoff-high’ and ‘cutoff-low’ were 50%, axis units were ‘Months’, and P value < 0.05 was considered statistically significant.
Screening for pseudogene- regulated miRNAs and miRNA-target mRNAs
The public online datasets of starBase v-2.0 and miRTarBase were used to identify pseudogene-binding miRNAs and miRNA-target mRNAs, respectively [17, 18]. The network of pseudogenes-miRNA-mRNA was constructed using Cytoscape v-3.7.2 [19].
KEGG pathways and gene oncology (GO) enrichment analysis of target mRNAs
The list of miRNA-target genes was imported into the STRING v-11.0, and the top five significantly GO terms and KEGG pathways were selected according to the values of false discovery rate (FDR), and then were visualized by GraphPad PRISM Version 6.02 [20].
Construction of protein–protein interaction network and identification of hub genes
STIRNG v-11.0 was used to construct the regulatory network of protein–protein, and then visualized by Centiscape plugin of Cytoscape v-3.7.2 [19,20,21]. The top 10 hub genes were identified according to the values of Degree unDir.
Hub genes expression and mutations analysis
Hub genes expression and mutations analysis in ovarian serous cystadenocarcinoma were analyzed using the online cBioPortal database [22]. 489 patients (TCGA, Nature 2011) with ovarian serous cystadenocarcinoma were selected for further analysis. The select genomic profiles were as follows: ‘Mutations’; ‘Putative copy-number alterations (GISTIC)’; ‘mRNA/miRNA expression Z-scores (all genes)’, and the Z-scores threshold were ± 2. Finally, OncoPrint was obtained under the guidance of online database at c-BioPortal.
Identification of potential target gene of LDHAP5
Pearson correlation analysis between LDHAP5 and the top 10 hub genes expression in ovarian serous cystadenocarcinoma was performed using GEPIA [16]. Kaplan–Meier overall survivals of target genes were analyzed by Kaplan–Meier Plotter [23]. The mRNA expression levels of 10 hub genes in TCGA patients were further measured using Oncomine Main database [24].
Results
Identification of dysregulated pseudogenes in four common gynecological malignancies
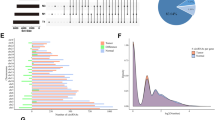
According to epidemiological statistics, cervical squamous cell carcinoma, ovarian serous cystadenocarcinoma, uterine corpus endometrial carcinoma, and uterine carcinosarcoma remain lethal diseases in women [1]. To explore the potential role of pseudogenes in carcinogenesis and cancer prognosis of four gynecological malignancies, we used the public dreamBase database to identify differentially expressed pseudogenes. As shown in Fig. 1a and Table 1, we identified 63 up-regulated and 0 down-regulated pseudogenes simultaneously in the four gynecological malignancies after preliminary screening. We then measured the expression levels of the 63 up-regulated pseudogenes in 32 types of human cancer (Fig. 1b). After removal of pseudogenes that were not highly expressed in the 32 types of human cancer, 40 pseudogenes were identified as playing potential roles in gynecological malignancies.
Identification of differentially expressed pseudogenes in four types of gynecological malignancies. a Venn diagram of 63 up-regulated pseudogenes in four gynecological malignancies. b Heat map of 63 frequently up-regulated pseudogenes in 32 types of human cancer. Red represents up-regulated genes and green represents down-regulated genes. Values in boxes represent |log2 FC| values
Prognostic analysis of up-regulated pseudogenes in 32 types of human cancer
We next explored the prognostic values of the 40 up-regulated pseudogenes in the 32 kinds of human cancer using GEPIA. As shown in Fig. 2, KRT8P3, KRT8P45, and LDHAP5 predicted poor overall survival in ovarian serous cystadenocarcinoma (HR = 1.3, P = 0.046; HR = 1.3, P = 0.019; HR = 1.3, P = 0.03, respectively), FTLP14 predicted poor unfavorable prognosis in uterine corpus endometrioid carcinoma (HR = 2.6, P = 0.018) No other pseudogenes that were significantly correlated with poor prognosis in the four types of gynecological malignancies.
Prognostic values of 40 upregulated pseudogenes in 32 kinds of human cancer using GEPIA. Red represents poor outcome, green represents good prognosis, yellow represents neutral outcome (hazard ratio = 1), and light blue represents insufficient sample size at these custom thresholds. Values in boxes are P-values. P-values less than 0.05 were considered statistically significant. GEPIA: gene expression profiling interactive analysis
Investigation of pseudogene-miRNA-mRNA regulatory network
By searching the starBase v2.0 database, only LDHAP5 had its corresponding miRNAs. The specific characteristics of the nine retrieved miRNAs are shown in Table-S1. In addition, as shown in Table-S2, only hsa-miR-181d-5p, hsa-miR-181c-5p, hsa-miR-7-5p, hsa-miR-543, hsa-miR-151a-5p, and hsa-miR-181b-5p had their own target genes. In total, 148 miRNA target genes, which were validated by at least one of three robust method (i.e., reporter assay, western blot, and quantitative-real-time polymerase chain reaction (qRT-PCR)), were identified via miRTarBase. The pseudogene-miRNA-mRNA network was constructed using Cytoscape v_3.7.2 (Fig. 3a).
Regulatory pseudogene-miRNA-mRNA network and enrichment analysis of 148 miRNA target mRNAs. a Pseudogene-miRNA-mRNA network constructed by Cytoscape v-3.7.2. b 148 miRNA target mRNAs were divided into three functional groups: i.e., biological processes, cellular components, and molecular functions. Top five GO enriched terms are shown according to FDR values. c Top five KEGG pathways are shown according to the FDR values. GO Gene Oncology, FDR false discovery rate
KEGG pathway and gene oncology (GO) enrichment analysis of miRNA target mRNAs
The 148 miRNA target genes were imported into STRING v-11.0, with GO and KEGG pathway enrichment analysis then performed under the operational guidance of the website. We selected the top five significantly enriched GO terms and KEGG pathways according to false discovery rate (FDR) values. The top five Biological Process (BO), Molecular Function (MO) and Cellular Component (CO) and their corresponding FDR values are shown in Fig. 3b. The top five significantly enriched KEGG pathways were MicroRNAs in cancer (hsa05206, FDR = 4.32E−26), Pathway in cancer (hsa05200, FDR = 6.77E−18), PI3K-AKT signaling pathway (hsa04151, FDR = 9.95E−16), Endocrine resistance (hsa01522, FDR = 9.95E−16), and Foxo signaling pathway (hsa04068, FDR = 2.65E−15) (Fig. 3c). These findings confirmed that the LDHAP5 pseudogene may mediate the occurrence and progression of ovarian serous cystadenocarcinoma.
EGFR as target mRNA of LDHAP5 in ovarian serous cystadenocarcinoma
We used the Centiscape plugin of Cytoscape v-3.7.2 to visualize the regulatory protein–protein network constructed using STRING v-11.0 (Fig. 4). The top 10 hub genes (i.e., TP53, MYC, EGFR, PTEN, HRAS, SIRT1, TNF, RELA, KRAS, and CREB1) were then identified based on Degree unDir values (Table 2). We further explored the sequence mutations and copy-number alterations of the 10 hub genes in ovarian serous cystadenocarcinoma using cBioportal. The group (TCGA, Nature 2011) which contained 489 patients was selected. However, only 361 patients (64.6%) were suitable for further analysis. The mutation frequencies of the 10 hub genes were TP53 (96%), MYC (34%), EGFR (9%), PTEN (14%), HRAS (9%), KRAS (24%), SIRT1 (10%), TNF (24%), RELA (11%) and CREB1 (10%), respectively (Fig. 5). Pearson correlation analysis showed that EGFR (R = 0.16, P = 0.00072), PTEN (R = 0.098, P = 0.043), SIRT1 (R = 0.094, P = 0.013), RELA (R = 0.18, P = 0.00013) and CREB1 (R = 0.16, P = 0.00094) were significantly correlated with LDHAP5 expression in ovarian serous cystadenocarcinoma (Table 3). Using the Oncomine Main database, only EGFR (fold-change = 1.192, P = 0.001), PTEN (fold-change = 1.214, P = 0.007), and CREB1 (fold-change = 1.723, P = 1.66E−04) mRNAs were more highly expressed in TCGA ovarian patients (n = 594) than in normal patients (n = 8) (Fig. 6a). We further analyzed the prognostic values (overall survival) of the five hub genes in ovarian serous cystadenocarcinoma using Kaplan–Meier plotter (Table 4, Fig. 6b). Only EGFR was significantly correlated with poor outcome (HR = 1.51, 95% CI 1.15–2, P = 0.0033) in ovarian serous cystadenocarcinoma, whereas SIRT1 predicted a good outcome (HR = 0.75, 95% CI 0.57–1, P = 0.047). Thus, according to the pseudogene-miRNA-mRNA regulatory mechanism, we concluded that LDHAP5 may play potential roles in ovarian serous cystadenocarcinoma by targeting EGFR.
Discussion
With deepening research, we continue to gain a better understanding of pseudogenes. Currently, there are two major pseudogene classifications. Firstly, pseudogenes can be divided into three categories based on differences in structure and origin, i.e., duplicated, unitary, and processed pseudogenes, respectively. Duplicated pseudogenes are caused by mutations of the gene coding region or regulatory region in the process of genome DNA tandem replication or chromosome unequal exchange [25]. Unitary pseudogenes cannot be transcribed or translated because of spontaneous mutations in the coding or regulatory regions of a single copy gene with coding function [26]. Both duplicated and unitary pseudogenes are collectively called unprocessed pseudogenes. Processed pseudogenes are formed by the random integration of mRNA transcripts into cDNA and lose their normal functions due to improper insertion sites or sequence mutations [27, 28]. Secondly, pseudogenes can be classified based on their functions into pseudogenes that can be transcribed, pseudogenes that cannot be transcribed, and pseudogenes that can encode short-chain peptides or truncated proteins. These pseudogenes play important roles in carcinogenesis and cancer prognosis [29,30,31].
Centered on the ceRNA hypothesis, our research focused on pseudogenes that can be transcribed into mRNA. We used the pseudogene-miRNA-mRNA regulatory network to identify pseudogenes that may play potential roles in common gynecological malignancies and to explore their related mechanisms.
The initial goal of our study was to discover pseudogenes that were differentially expressed in four common gynecological malignancies. However, we only found three and one significantly up-regulated pseudogenes that predicted poor prognosis in ovarian serous cystadenocarcinoma and uterine corpus endometrioid carcinoma after Kaplan–Meier survival analysis. We selected LDHAP5 as the candidate pseudogenes as it had corresponding miRNAs. There are two reasons accounting for the lack of pseudogenes. Firstly, many pseudogenes remain unidentified. Initially, pseudogenes were considered as “junk” or “fossil” DNA, and many methods were developed to avoid their detection [32,33,34,35,36]. The second possibility is that the current ceRNA hypothesis is not yet perfect, and further analysis is needed to build a more comprehensive regulatory network [37].
In our study, 148 potential target mRNAs were identified. Functional enrichment analysis showed the top five significantly enriched gene sets were MicroRNAs in cancer (hsa05206), Pathway in cancer (hsa05200), PI3K-AKT signaling pathway (hsa04151), Endocrine resistance (hsa01522), and Foxo signaling pathway (hsa04068). Interestingly, epithelial ovarian cancer, bladder cancer, lung cancer, and colorectal cancer were enriched in the MicroRNAs in cancer pathway (hsa05206). The PI3K-AKT signaling pathway has been researched extensively and plays an important role in a variety of cancers. Studies have shown that activated AKT mediates various downstream reactions, including cell survival, growth, proliferation, cell migration, and angiogenesis via phosphorylation of a range of intracellular proteins [38, 39]. More significantly, studies have shown that EGFR is dysregulated in many solid tumors, and PI3K-AKT signaling can be used as a downstream regulatory pathway for EGFR to mediate the occurrence and progression of disease, as confirmed in many cancers [40, 41].
Our research has several limitations. Specially, our conclusions are primarily based on the analysis of existing databases. To further confirm the role of the LDHAP5 pseudogene at the in vivo and in vitro level, we need to construct ovarian cancer cell lines that differentially express LDHAP5, with clinical pathological specimens from ovarian cancer patients also used to verify our findings. EGFR antagonists (e.g., gefitinib, lapatinib, erlotinib) have been used in a variety of cancers, including pancreatic, small cell lung, and colorectal cancer [42,43,44]. Once our research is successfully validated, it may be used in ovarian cancer in the future. With continuing research, more pseudogene functions and corresponding mechanisms will be revealed, which could help in the identification of novel biomarkers, development of specific drug design, and the adoption of personalized treatment in the future.
Conclusions
This study is the first to report on the high expression of the LDHAP5 pseudogene in ovarian serous cystadenocarcinoma, which may lead to poor prognosis via its targeting of EGFR. Thus, LDHAP5 may serve as a new therapeutic target, and improve the prognosis of patients with ovarian cancer in the future.
Availability of data and materials
Not applicable.
Abbreviations
- MREs:
-
MicroRNA response elements
- lncRNAs:
-
Long non-coding RNAs
- ceRNAs:
-
Competitive endogenous RNAs
- RBPs:
-
RNA binding proteins
- GEPIA:
-
Gene expression profiling interactive analysis
- GO:
-
Gene oncology
- FDR:
-
False discovery rate
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA. 2020;70(1):7–30.
Fotopoulou C, Neumann U, Kraetschell R, Schefold JC, Weidemann H, Lichtenegger W, et al. Long-term clinical outcome of pelvic exenteration in patients with advanced gynecological malignancies. J Surg Oncol. 2010;101(6):507–12.
Diaz-Padilla I, Duran I, Clarke BA, Oza AM. Biologic rationale and clinical activity of mTOR inhibitors in gynecological cancer. Cancer Treat Rev. 2012;38(6):767–75.
Jacq C, Miller JR, Brownlee GG. A pseudogene structure in 5S DNA of Xenopus laevis. Cell. 1977;12(1):109–20.
Ding W, Lin L, Chen B, Dai J. L1 elements, processed pseudogenes and retrogenes in mammalian genomes. IUBMB Life. 2006;58(12):677–85.
Poliseno L, Salmena L, Zhang J, Carver B, Haveman WJ, Pandolfi PP. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature. 2010;465(7301):1033–8.
Pink RC, Wicks K, Caley DP, Punch EK, Jacobs L, Carter DR. Pseudogenes: pseudo-functional or key regulators in health and disease? RNA. 2011;17(5):792–8.
Sen K. Ghosh T. Pseud Compos: Delving in the ‘debris’ of human genome. Briefings in functional genomics; 2013. p. 12.
Korrodi-Gregorio L, Abrantes J, Muller T, Melo-Ferreira J, Marcus K, da Cruz e Silva OA, et al. Not so pseudo: the evolutionary history of protein phosphatase 1 regulatory subunit 2 and related pseudogenes. BMC Evol Biol. 2013;13:242.
Muro EM, Mah N, Andrade-Navarro MA. Functional evidence of post-transcriptional regulation by pseudogenes. Biochimie. 2011;93(11):1916–21.
Poliseno L, Marranci A, Pandolfi PP. Pseudogenes in human cancer. Front Med. 2015;2:68.
Poliseno L. Pseudogenes: newly discovered players in human cancer. Sci Signal. 2012;5(242):re5.
Salmena L, Poliseno L, Tay Y, Kats L, Pandolfi PP. A ceRNA hypothesis: the Rosetta Stone of a hidden RNA language? Cell. 2011;146(3):353–8.
Hu X, Yang L, Mo YY. Role of pseudogenes in tumorigenesis. Cancers. 2018;10(8):256.
Zheng LL, Zhou KR, Liu S, Zhang DY, Wang ZL, Chen ZR, Yang JH, Qu LH. dreamBase: DNA modification, RNA regulation and protein binding of expressed pseudogenes in human health and disease. Nucleic Acids Res. 2018;46(D1):D85–91.
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017;45(W1):W98–102.
Li JH, Liu S, Zhou H, Qu LH, Yang JH. StarBase v2 0: decoding miRNA-ceRNA, miRNA-ncRNA and protein–RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014;42(Database issue):D92–7.
Chou CHSS, Yang CD, et al. miRTarBase update 2018: a resource for experimentally validated microRNA-target interactions. Nucleic Acids Res. 2018;46(D1):D296–302.
Shannon PMA, Ozier O, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13(11):498–2504.
Szklarczyk D, Morris JH, Cook H, et al. The STRING database in 2017: quality-controlled protein–protein association networks, made broadly accessible. Nucleic Acids Res. 2017;45(D1):D362–8.
Scardoni G, Petterlini M, Laudanna C. Analyzing biological network parameters with CentiScaPe. Bioinformatics. 2009;25(21):2857–9.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012;2(5):401–4.
Győrffy B, Lánczky A, Szállási Z. Implementing an online tool for genome-wide validation of survival-associated biomarkers in ovarian-cancer using microarray data from 1287 patients. Endocr Relat Cancer. 2012;19(2):197–208.
Rhodes DR, Kalyana-Sundaram S, Mahavisno V, Varambally R, Yu J, Briggs BB, Barrette TR, Anstet MJ, Kincead-Beal C, Kulkarni P, Varambally S. Oncomine 3.0: genes, pathways, and networks in a collection of 18,000 cancer gene expression profiles. Neoplasia. 2007;9(2):166–80.
Mighell AJ, Smith NR, Robinson PA, Markham AF. Vertebrate pseudogenes. FEBS Lett. 2000;468(2–3):109–14.
Zhang ZD, Frankish A, Hunt T, Harrow J, Gerstein M. Identification and analysis of unitary pseudogenes: historic and contemporary gene losses in humans and other primates. Genome Biol. 2010;11(3):R26.
Maestre J, Tchenio T, Dhellin O, Heidmann T. mRNA retroposition in human cells: processed pseudogene formation. EMBO J. 1995;14(24):6333–8.
D’Errico I, Gadaleta G, Saccone C. Pseudogenes in metazoa: origin and features. Brief Funct Geno Proteomics. 2004;3(2):157–67.
Tam OH, Aravin AA, Stein P, Girard A, Murchison EP, Cheloufi S, et al. Pseudogene-derived small interfering RNAs regulate gene expression in mouse oocytes. Nature. 2008;453(7194):534–8.
Eichenlaub MP, Ettwiller L. De novo genesis of enhancers in vertebrates. PLoS Biol. 2011;9(11):e1001188.
Kandouz M, Bier A, Carystinos GD, Alaoui-Jamali MA, Batist G. Connexin43 pseudogene is expressed in tumor cells and inhibits growth. Oncogene. 2004;23(27):4763–70.
Andersson JO, Andersson SG. Pseudogenes, junk DNA, and the dynamics of Rickettsia genomes. Mol Biol Evol. 2001;18(5):829–39.
Yeo C, Brody J. Does this band make sense? Limits to expression based cancer studies. Cancer Lett. 2008;271:81–4.
Brent MR. Steady progress and recent breakthroughs in the accuracy of automated genome annotation. Nat Rev Genet. 2008;9:62–73.
Britten RJ, Davidson EH. Gene regulation for higher cells: a theory. Science. 1969;165(3891):349–57.
Britten RJ, Davidson EH. Repetitive and non-repetitive DNA sequences and a speculation on the origins of evolutionary novelty. Q Rev Biol. 1971;46(2):111–38.
Cheetham SW, Faulkner GJ. Overcoming challenges and dogmas to understand the functions of pseudogenes. Nat Rev Genet. 2020;21(3):191–201.
Hoxhaj G, Manning BD. The PI3K–AKT network at the interface of oncogenic signalling and cancer metabolism. Nat Rev Cancer. 2019;20(2):74–88.
Murugan AK. Special issue: PI3K/Akt signaling in human cancer. Semin Cancer Biol. 2019;59:1–2.
Fu X, Cui G, Liu S, Zhao S. Linc01014 regulates gefitinib resistance in oesophagus cancer via EGFR-PI3K-AKT-mTOR signalling pathway. J Cell Mol Med. 2020;24(2):1670–5.
Zhang F, Xu M, Yin X, Guo H, Zhang B, Wang Y, et al. TWEAK promotes hepatic stellate cell migration through activating EGFR/Src and PI3K/AKT pathways. Cell Biol Int. 2020;44(1):278–85.
Wang Z, Cheng Y, An T, Gao H, Wang K, Zhou Q, et al. Detection of EGFR mutations in plasma circulating tumour DNA as a selection criterion for first-line gefitinib treatment in patients with advanced lung adenocarcinoma (BENEFIT): a phase 2, single-arm, multicentre clinical trial. Lancet Respir Med. 2018;6(9):681–90.
Pyrotinib Tops Lapatinib in Metastatic Breast Cancer. Cancer discovery. 2019; 9(11): 3.
Xiong L, Li R, Sun J, Lou Y, Zhang W, Bai H, et al. Erlotinib as neoadjuvant therapy in stage IIIA (N2) EGFR mutation-positive non-small cell lung cancer: a prospective, single-arm. Phase II Study. Oncologist. 2019;24(2):157.
Acknowledgements
Not applicable.
Funding
This work was supported by National Natural Science Foundation of China (81630060), and the research-oriented clinician funding program of Tongji Medical College, Huazhong University of Science and Technology.
Author information
Authors and Affiliations
Contributions
PW was responsible for the study concept and design; SL, CC, PW, PG, WZ, TP were involved in data collection, data screening and statistical analysis; SL wrote the manuscript, and YM took charge of supervising the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethical approval and informed consent
Not applicable.
Consent for publication
Not applicable.
Competing interests
The author(s) declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Additional file 1: Table S1.
miRNAs targeting LDHAP5 were predicted by starBase v2.0.
Additional file 2: Table S2.
Numbers of miRNA target gene identified by miRTarBase.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Lin, S., Meng, Y., Cao, C. et al. Comprehensive analysis of LDHAP5 pseudogene expression and potential pathogenesis in ovarian serous cystadenocarcinoma. Cancer Cell Int 20, 229 (2020). https://doi.org/10.1186/s12935-020-01324-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12935-020-01324-6