Abstract
Background
Despite an augmented research effort and scale-up of highly active antiretroviral therapy, a high prevalence of HIV-1-associated neurocognitive disorders (HAND) persists in the HIV-infected population. Nearly 50 % of all HIV-1-infected individuals suffer from a neurocognitive disorder due to neural and synaptodendritic damage. Challenges in HAND research, including limited availability of brain tissue from HIV patients, variation in HAND study protocols, and virus genotyping inconsistency and errors, however, have resulted in studies with insufficient power to delineate molecular mechanisms underlying HAND pathogenesis. There exists, therefore, a great need for a reliable and centralized resource specific to HAND research, particularly for epidemiological study and surveillance in resource-limited countries where severe forms of HAND persist.
Description
To address the aforementioned imperative need, here we present the HAND Database, a resource containing well-curated and up-to-date HAND virus information and associated clinical and epidemiological data. This database provides information on 5,783 non-redundant HIV-1 sequences from global HAND research published to date, representing a total of 163 unique individuals that have been assessed for HAND. A user-friendly interface allows for flexible searching, filtering, browsing, and downloading of data. The most comprehensive database of its kind, the HAND Database not only bolsters current HAND research by increasing sampling power and reducing study biases caused by protocol variation and genotyping inconsistency, it allows for comparison between HAND studies across different dimensions. Development of the HAND Database has also revealed significant knowledge gaps in HIV-driven neuropathology. These gaps include inadequate sequencing of viral genes beyond env, lack of HAND viral data from HIV epidemiologically important regions including Asian and Sub-Saharan African countries, and biased sampling toward the male gender, all factors that impede efforts toward providing an improved quality of life to HIV-infected individuals, and toward elimination of viruses in the brain.
Conclusion
Our aim with the HAND database is to provide researchers in both the HIV and neuroscience fields a comprehensive and rigorous data source toward better understanding virus compartmentalization and to help in design of improved strategies against HAND viruses. We also expect this resource, which will be updated on a regular basis, to be useful as a reliable reference for further HAND epidemiology studies. The HAND Database is freely available and accessible online at http://www.handdatabase.org.
Similar content being viewed by others
Background
Human immunodeficiency virus (HIV)-associated neurocognitive disorder (HAND) occurs due to damage to neurons and synapses by viral protein products, and due to a chemokine/cytokine imbalance in the brain, a pro-inflammatory response to HIV infection of macrophages and microglia [1–3]. HIV entry into the brain is an early event following infection [4], and presence of the blood brain barrier greatly limits entry of antiretroviral therapy into the brain. Our ability to control viral levels within and viral damage to the HIV-infected brain, therefore, remains highly limited. While the introduction of highly active antiretroviral therapy (HAART) brought about a decrease in the incidence of the most severe forms of HAND, i.e., HIV-associated dementia, the prevalence of milder forms has continued to increase [5–7]. In the recent HIV Anti-Retroviral Therapy Effects Research Study, nearly 50 % of all HIV-1 individuals exhibited some form of HAND, including deficits in motor function, verbal fluency, learning, memory, and attention [8]. HAND individuals experience difficulty performing day-to-day tasks, are less likely to adhere to medical treatments and other HIV-1 prevention practices, and ultimately suffer from around a threefold increased risk of death as compared to a mentally-healthy HIV-1 individual [9]. In addition, in resource-limited countries, the most severe forms of HAND continue to devastate the mental health of HIV individuals [9].
Delineating the underpinning molecular mechanisms of HAND development is critical to providing HIV-infected individuals an elevated quality of life, as well as toward clearance of the virus repertoire in the brain. Research in this area, however, has been largely limited by availability of samples from both the brain and from HAND-assessed individuals. In addition, a need to understand HAND progression across an HIV individual’s lifespan, coupled with difficulty in obtaining brain samples, has made cerebrospinal fluid (CSF) sampling a surrogate endpoint for assessing HAND development [10]. Both, small sample size from individual studies and indirect CSF inference have made it difficult to fully assess the complex interaction between viruses and the brain in the HAND setting. Additionally, variations in study methodologies and result interpretations have further confounded HAND studies, leading to conflicting findings in the field. To address these issues, there therefore exists a great need for a reliable HIV sequence resource, of adequate sample size, for HAND research.
Toward this effort, we developed a centralized HAND Database based on all HAND studies published to date. This resource database is freely accessible at: http://www.handdatabase.org. The HAND Database serves as the most comprehensive database in its field, and contains well-curated HAND virus information, epidemiology sampling data, patient clinical status, and therapy treatment information. All information was cross-validated using multiple resources, including the literature, GenBank entry, and author contact. Furthermore, all viral sequences have undergone stringent quality control examination, including genotyping validation, in order to minimize genotyping errors frequently seen in HIV subtype-based studies [11].
The only other published HIV database related to brain tissue, The HIV Brain Sequence Database [12], contains HIV env sequences from brain tissue, as well as from other tissues in patients with brain samples. In contrast, our database contains HAND-specific information with regards to virus sequences (genome coverage beyond env), epidemiology sampling information, clinical data, and treatment status, all factors important to the study of HAND pathogenesis. Unprecedented in its comprehensiveness of curated HAND HIV information, our HAND Database serves as a centralized gateway to study the role of HIV in the HAND setting.
Construction and content
Data sources
An extensive literature review was conducted to develop a comprehensive set of HAND-related research articles, from which we then extracted sequence data from HAND-assessed individuals. This literature search resulted in the use of data from 41 published studies. Publically available HIV-1 sequence data were collected from the GenBank (last accessed 3/2013) and the LANL HIV sequence database (last accessed 2/2014) [13, 14]. HIV-1 individual sampling and clinical information was collected from the relevant literature, the two aforementioned databases, and through communication with publication authors.
Sequence and clinical data filtering
All collected sequence data were validated through a series of quality control steps. We first employed the LANL quality control pipeline to check for potential problematic viruses with sequencing errors [13]. Amplification contamination was detected using BLASTn (v. 2.2.26) [15]. In addition, data regarding epidemiology sampling, clinical status, and treatment status were cross-referenced whenever available in more than one of the resources listed above.
Genotyping analysis
Genotyping of HIV sequence data is frequently inconsistent and error-prone [11].
Therefore, all filtered HIV sequences were re-genotyped. Here we applied the jumping profile Hidden Markov Model genotyping program (jpHMM), whose genotyping accuracy has been established [16–18]. In brief, following a hypermutation analysis [13], sequences greater than 300 nucleotides in length and with a hypermutation p-value of 0.05 or greater were subject to genotyping.
Database schema
The HAND Database was constructed using the relational database management system MySQL (v.5.6.17). MySQL was chosen for its ease of use, its high reliability, and as it is freely available. HIV-1 sequence and clinical data were compiled into one flat file, with annotations divided into three major categories: sequence and sequence descriptor data, HIV-1 patient descriptor data, and sample descriptor data (Table 1). Sequence data included the HIV-1 nucleotide sequence, sequence accession number, sequence genotype information, and sequence length. Epidemiology data included the geographical location and year at time of sampling, as well as tissue sampled. Patient data at time of sampling included patient age, risk factor, health status, CD4 count, viral load, HIV treatment information (treatment status, and when applicable, treatment type and duration), and patient HAND information (HAND status, the presence or absence of HAND, and when applicable, HAND type).
Utility
Database access and web query interface
The HAND Database was developed into a publically available, web accessible resource. The database website provides a home page with background information on HAND, as well as a help page to assist with database navigation (Fig. 1). The database itself allows for easy querying and downloading of user-defined data subsets. Researchers can perform a simple search using a keyword, or employ multiple column filters for a custom-made data subset. Selected entries can subsequently be downloaded into a variety of formats at the user’s discretion. Additional features include sorting by annotation of interest, as well as an option for viewing the complete record for any given entry.
The HAND Database Search Interface. The HAND Database provides flexible searching, filtering, and browsing capabilities. Sequence entries and annotations of interest can be exported into a variety of file formats for further use. In addition, a website navigation bar allows easy access to help, contact, and background information pages
Database content
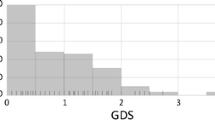
The HAND Database currently contains 5,783 HIV-1 sequences, representing a total of 163 unique individuals assessed for HAND status. For the 87 individuals with age information available, ages ranged from 19 to 63 years, with the largest proportion of individuals between 30 and 49 years of age (69 %) (Fig. 2). Gender information was available for 64 individuals, the majority of whom were males (77 %). HAND status, the absence or presence of HAND, was obtained for almost all database individuals (96 %), and indicated a close split between non-HAND (44 %) and HAND (52 %) patients. The top three reported HAND types in HAND-positive individuals were HIV-1-associated dementia (HAD, 54 %), HIV-1 encephalopathy (HIVE, 35 %), and AIDS dementia complex (ADC, 8 %) (Fig. 3). HIV treatment status information, whether or not an individual had received HIV treatment prior to sampling, was available for 67 % of individuals, and the majority of individuals with treatment information had received some form of treatment (49 %). Nearly half of all treated individuals had received HAART (46 %) prior to sampling, while the rest had received one or more forms of HIV monotherapy (54 %) (Fig. 4).
Distribution of HAND Database Entries By HAND Status And HAND Type. The top chart shows HAND status distribution across all database individuals, and the bottom chart shows HAND type distribution across database individuals for whom this information was available. The majority of individuals with HAND had HIV-associated dementia (HAD), followed by HIV-encephalitis (HIVE), AIDS dementia complex (ADC), and minor cognitive-motor disorder (MCMD). HAND type designations were obtained from the literature, and for some individuals, more than one HAND type had been assigned
Distribution of HAND Database Entries By HIV Therapy Status And HIV Therapy Type. The top chart shows HIV therapy status distribution across all database individuals, and the bottom chart shows HIV therapy type distribution across database individuals for whom this information was available. Nearly half of all treated individuals had received HAART. Therapy type designations were as we found to be reported in the literature, and for some individuals, more than one HIV therapy type had been assigned
Geographical region sampling information was available for 156 patients, with the top three sampling regions being North America (60 %), Europe (25 %), and Asia (6.4 %) (Fig. 5). Samples were derived from 20 different tissue types, with the top three sampling tissues being brain (47 %), lymph node (14 %), and CSF (7 %).
Five HIV-1 genes were represented in our database, gag, pol, env, tat, and nef, with the majority of sequence coverage in the env gene (Fig. 6). This result was expected due the known role of env in macrophage tropism, viral replication, and activation of pro-inflammatory responses toward neuronal injury [19–21]. Of all archived sequences, 79 % of sequences that underwent genotyping validation were of the pure B subtype, and all non-recombinant sequences were confirmed as having been correctly reported in the literature. Sixteen sequences were found to have undergone recombination events not reported in either the source literature or databases.
HAND Database HIV-1 Genome Coverage And Sequencing Depth. The top panel displays HAND Database sequencing depth across the HXB2 reference sequence, and the bottom panel displays HIV-1 gene location across the HXB2 reference sequence. The env gene was the HIV genomic region with the greatest sequencing depth. HXB2 accession number: K03455
Discussion
Despite increased HAND research and treatment efforts, the persistent prevalence of HAND continues to pose a great challenge to the HIV research and patient communities. Investigation in this area is limited by small sample sizes, primarily due to difficulty in obtaining tissue samples, and by variation in study protocols and result interpretation. Furthermore, errors and inconsistency in HIV genotyping compound the complexity in delineating viral mechanisms toward neuropathology. The HAND Database described here serves to narrow these research gaps and addresses the need for a reliable and centralized HAND data source for advanced research purposes.
The HAND database contains up-to-date and well-curated HAND virus and patient information. All sequence data have been subject to stringent quality control examination and re-genotyping, thereby laying a solid foundation toward elucidation of viral mechanisms driving neuropathology under various epidemiology settings.
In creating this resource we noted a number of sequencing and sampling biases that currently limit research in the area, and have developed a set of potential research directions that may greatly benefit the HAND research community. First, although prior studies have indicated the role of multiple HIV proteins, including Nef, Vpr, and Tat [22–29], toward HAND development, the majority of research in the area has focused on the gp120 envelope glycoprotein. This sequencing bias is largely due to interest in Env for its role in conferring viral tropism for microglia and macrophage cells [30–33], its role in non-neuronal cell replication [34], and for its potential as an HIV therapeutic target [35]. A shortage of sequence data beyond the env gene, however, limits our ability to perform data-driven HAND research on the complete viral genome, and therefore an increase in sequencing efforts in other areas of the genome would provide insight into the role of regulatory and accessory proteins toward HAND pathogenesis. Second, there is a distinct lack of sequence data from HIV epidemiologically important regions including many Asian and Sub-Saharan African countries (Fig. 5). Limited access to HAART contributes to an increased vulnerability of HIV individuals in these geographical regions to the most severe forms of HAND. Recent studies indicate HIV-associated dementia (HAD) affects over 25 % of HIV individuals in several Sub-Saharan African countries [36–38]. In addition, research on treatment-naïve HIV-1-individuals in Thailand has greatly contributed to our understanding of HAND pathogenesis [39]. Finally, we noted a bias toward sequencing of male individuals. Research beyond the HIV field has implicated gender as playing a role in determining those genetic processes leading to neurocognitive deficiencies [40, 41]. A lack of information on HAND females, however, currently proves an obstacle in determining potential gender differences in HAND pathogenesis.
Conclusions
Developing a better understanding of mechanisms underlying the development of neurocognitive disorders is crucial toward providing the HIV patient community with a higher quality of life, and toward prevention of enhanced transmission. Through consolidation and validation of data from multiple data sources, here we have developed the HAND Database, a single, intuitive platform from which researchers can launch their high-throughput HAND sequencing projects. The HAND database contains up-to-date and curated HAND HIV virus and HIV-infected individual information, providing a solid foundation toward the elucidation of viral mechanisms driving this neuropathology. In particular, we anticipate this database will be of great use in increasing HAND research efforts in resource-limited countries. We plan to continue expanding the HAND Database as new HAND viral sequence data become publically available.
Availability and requirements
All records are freely available and accessible at www.handdatabase.org.
References
Johnson T, Nath A. Immune reconstitution inflammatory syndrome and the central nervous system. Curr Opin Neurol. 2011;24(3):284–90.
Marcondes MCG, Burudi E, Huitron-Resendiz S, Sanchez-Alavez M, Watry D, Zandonatti M, et al. Highly activated CD8+ T cells in the brain correlate with early central nervous system dysfunction in simian immunodeficiency virus infection. J Immunol. 2001;167(9):5429–38.
Zheng J, Zhuang W, Yan N, Kou G, Peng H, McNally C, et al. Classification of HIV-1-Mediated neuronal dendritic and synaptic damage using multiple criteria linear programming. Neuroinformatics. 2004;2(3):303–26.
Resnick L, Berger JR, Shapshak P, Tourtellotte WW. Early penetration of the blood–brain-barrier by HIV. Neurol. 1988;38(1):9–14.
McArthur JC, Steiner J, Sacktor N, Nath A. HIV-associated neurocognitive disorders: Mind the gap. Ann Neurol. 2010;67(6):699–714.
Tozzi V, Balestra P, Lorenzini P, Bellagamba R, Galgani S, Corpolongo A, et al. Prevalence and risk factors for human immunodeficiency virus-associated neurocognitive impairment, 1996 to 2002: Results from an urban observational cohort. J Neurovirol. 2005;11(3):265–73.
Sacktor N, McDermott MP, Marder K, Schifitto G, Selnes OA, McArthur JC, et al. HIV-associated cognitive impairment before and after the advent of combination therapy. J Neurovirol. 2002;8(2):136–42.
Heaton RK, Clifford DB, Franklin Jr DR, Woods SP, Ake C, Vaida F, et al. HIV-associated neurocognitive disorders persist in the era of potent antiretroviral therapy: CHARTER Study. Neurol. 2010;75(23):2087–96.
Horn, T. Update on the Neurological Manifestations of HIV. In: The PRN Notebook. Physicians’ ResearchNetwork. 2005. http://www.prn.org/images/pdfs/74_mcarthur_justin.pdf.
Ellis RJ, Moore DJ, Childers ME, Letendre S, McCutchan JA, Wolfson T, et al. Progression to neuropsychological impairment in human immunodeficiency virus infection predicted by elevated cerebrospinal fluid levels of human immunodeficiency virus RNA. Arch Neurol. 2002;59(6):923–8.
Zhang M, Foley B, Schultz AK, Macke JP, Bulla I, Stanke M, et al. The role of recombination in the emergence of a complex and dynamic HIV epidemic. Retrovirology. 2010;7:25.
Holman AG, Mefford ME, O’Connor N, Gabuzda D. HIVBrainSeqDB: A database of annotated HIV envelope sequences from brain and other anatomical sites. AIDS Res Ther. 2010;43.
Los Alamos HIV Sequence Database. Los Alamos National Laboratory. http://www.hiv.lanl.gov. Accessed Feb 2014.
Benson DA, Clark K, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW. GenBank. Nucleic Acids Res. 2014;42(Database issue):D32–7.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215(3):403–10.
Zhang M, Schultz AK, Calef C, Kuiken C, Leitner T, Korber B, et al. jpHMM at GOBICS: A web server to detect genomic recombinations in HIV-1. Nucleic Acids Res. 2006;34(Web Server issue):W463–5.
Schultz AK, Zhang M, Bulla I, Leitner T, Korber B, Morgenstern B, et al. jpHMM: Improving the reliability of recombination prediction in HIV-1. Nucleic Acids Res. 2009;37(Web Server issue):W647–51.
Schultz AK, Zhang M, Leitner T, Kuiken C, Korber B, Morgenstern B, et al. A jumping profile hidden markov model and applications to recombination sites in HIV and HCV genomes. BMC Bioinformatics. 2006;7:265.
Toggas SM, Masliah E, Rockenstein EM, Rall GF, Abraham CR, Mucke L. Central nervous system damage produced by expression of the HIV-1 coat protein gp120 in transgenic mice. Nature. 1994;367(6459):188–93.
Kaul M, Lipton SA. Experimental and potential future therapeutic approaches for HIV-1 associated dementia targeting receptors for chemokines, glutamate and erythropoietin. Neurotox Res. 2005;8(1–2):167–86.
Fontana G, Valenti L, Raiteri M. Gp120 can revert antagonism at the glycine site of NMDA receptors mediating GABA release from cultured hippocampal neurons. J Neurosci Res. 1997;49(6):732–8.
Lamers SL, Salemi M, Galligan DC, Morris A, Gray R, Fogel G, et al. Human immunodeficiency virus-1 evolutionary patterns associated with pathogenic processes in the brain. J Neurovirol. 2010;16(3):230–41.
Lamers SL, Poon AF, McGrath MS. HIV-1 Nef protein structures associated with brain infection and dementia pathogenesis. PLoS ONE. 2011;6(2):e16659.
Thomas ER, Dunfee RL, Stanton J, Bogdan D, Kunstman K, Wolinsky SM, et al. High frequency of defective vpu compared with tat and rev genes in brain from patients with HIV type 1-associated dementia. AIDS Res Hum Retroviruses. 2007;23(4):575–80.
Gonzalez-Scarano F, Martin-Garcia J. The neuropathogenesis of AIDS. Nat Rev Immunol. 2005;5(1):69–81.
Nath A. Human immunodeficiency virus (HIV) proteins in neuropathogenesis of HIV dementia. J Infect Dis. 2002;186 Suppl 2:S193–8.
Kaul M, Lipton SA. Mechanisms of neuronal injury and death in HIV-1 associated dementia. Curr HIV Res. 2006;4(3):307–18.
Dunfee R, Thomas ER, Gorry PR, Wang J, Ancuta P, Gabuzda D. Mechanisms of HIV-1 neurotropism. Curr HIV Res. 2006;4(3):267–78.
King JE, Eugenin EA, Buckner CM, Berman JW. HIV tat and neurotoxicity. Microbes Infect. 2006;8(5):1347–57.
Chesebro B, Wehrly K, Nishio J, Perryman S. Mapping of independent V3 envelope determinants of human immunodeficiency virus type 1 macrophage tropism and syncytium formation in lymphocytes. J Virol. 1996;70(12):9055–9.
O’Brien WA, Koyanagi Y, Namazie A, Zhao JQ, Diagne A, Idler K, et al. HIV-1 tropism for mononuclear phagocytes can be determined by regions of gp120 outside the CD4-binding domain. Nature. 1990;348(6296):69–73.
Shioda T, Levy JA, Cheng-Mayer C. Small amino acid changes in the V3 hypervariable region of gp120 can affect the T-cell-line and macrophage tropism of human immunodeficiency virus type 1. PNAS. 1992;89(20):9434–8.
Jordan CA, Watkins BA, Kufta C, Dubois-Dalcq M. Infection of brain microglial cells by human immunodeficiency virus type 1 is CD4 dependent. J Virol. 1991;65(2):736–42.
Toohey K, Wehrly K, Nishio J, Perryman S, Chesebro B. Human immunodeficiency virus envelope V1 and V2 regions influence replication efficiency in macrophages by affecting virus spread. Virology. 1995;213(1):70–9.
Wu X, Yang ZY, Li Y, Hogerkorp CM, Schief WR, Seaman MS, et al. Rational design of envelope identifies broadly neutralizing human monoclonal antibodies to HIV-1. Science. 2010;329(5993):856–61.
Joska JA, Westgarth-Taylor J, Myer L, Hoare J, Thomas KG, Combrinck M, et al. Characterization of HIV-Associated Neurocognitive Disorders among individuals starting antiretroviral therapy in South Africa. AIDS Behav. 2011;15(6):1197–203.
Perriens JH, Mussa M, Luabeya MK, Kayembe K, Kapita B, Brown C, et al. Neurological complications of HIV-1-seropositive internal medicine inpatients in Kinshasa, Zaire. J Acquir Immune Defic Syndr. 1992;5(4):333–40.
Howlett WP, Nkya WM, Mmuni KA, Missalek WR. Neurological disorders in AIDS and HIV disease in the northern zone of Tanzania. AIDS. 1989;3(5):289–96.
Shiramizu B, Ratto-Kim S, Sithinamsuwan P, Nidhinandana S, Thitivichianlert S, Watt G, et al. HIV DNA and dementia in treatment-naive HIV-1-infected individuals in Bangkok, Thailand. Int J Med Sci. 2007;4(1):13–8.
Gatz M, Fiske A, Reynolds CA, Wetherell JL, Johansson B, Pedersen NL. Sex differences in genetic risk for dementia. Behav Genet. 2003;33(2):95–105.
Artero S, Ancelin ML, Portet F, Dupuy A, Berr C, Dartigues JF, et al. Risk profiles for mild cognitive impairment and progression to dementia are gender specific. J Neurol Neurosurg Psychiatry. 2008;79(9):979–84.
Acknowledgements
TZG was funded by a Presidential Scholarship from the University of Georgia. MZ was supported by NIH R03AI104258, R03AI120203, and UGA Faculty Research Grant 1025GR793002. We also thank two anonymous reviewers for their valuable critiques and suggestions.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
MZ and YJM conceived the infectious and chronic disease ideas that led to this study. TZG and WLK collected the data. TZG, YJM and MZ designed the study and analyzed the data. All authors contributed to the manuscript.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Griffin, T.Z., Kang, W., Ma, Y. et al. The HAND Database: a gateway to understanding the role of HIV in HIV-associated neurocognitive disorders. BMC Med Genomics 8, 70 (2015). https://doi.org/10.1186/s12920-015-0143-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12920-015-0143-8