Abstract
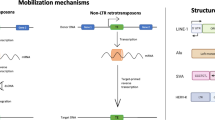
Researchers have long sought to understand the genetic basis of the cognitive differences between primates, with particular focus on the human brain. Although all mutational types have worked in concert with evolutionary forces to generate the current human brain, in this review we will explore the impact of mobile elements, specifically non-LTR retrotransposons. Non-LTR retrotransposons have contributed coding and regulatory sequences to the genome throughout evolution. During primate evolution there have been multiple waves of LINE retrotransposition as well as the birth of new mobile elements such as the SINEs Alu and SVA and we will explore what kinds of impacts these may have had on the evolving human brain.
Similar content being viewed by others
Genetic complexity of the human condition
The human species has developed art, literature, science, technology, agriculture, and grant cycles, all aspects of the human condition that are not observed in any other species of primate. Unique qualities of the human brain such as a relatively large cortical volume, surface area, and altered connectivity are often cited as key structural reasons for this increased complexity with respect to physiology. A yet unresolved question is the identification of the genetic modifications that underlie these physiological and cognitive complexities. Although all forms of mutation work together with selection and drift to produce the ever-dynamic phenotype, we will explore here the contribution of the repetitive portion of the genome to the evolution of the human brain, specifically that of non-LTR retrotransposons (RTs). By exploring the impact of RTs in disease, non-neuronal systems, and neuronal systems of model organisms we can gain insight into the still unresolved question of what role RTs might play in modifying neuronal function both throughout primate evolution and within the lifetime of a single individual human.
An evolutionary history of RTs during the expansion of the primate brain
RTs are present in most eukaryotic genomes and make up almost 40% of the human genome [1]. There are two major classes of active RTs in humans: long interspersed elements (LINEs) and short interspersed elements (SINEs). Of the RTs that are active in primates, there are two SINEs (SINE/Variable number tandem repeat/Alu (SVA) and Alu) and one LINE ( LINE-1 (L1)) that are commonly active in humans. The most ancient clades of eukaryotic RTs (GENIE, CRE, and R4) can be traced back at least 600 million years ago (mya) with the most ancient element identified within the primitive eukaryote Giardi lamblia [2,3,4]. Although the origin of non-LTR RTs can be traced to near the split of prokaryotes from eukaryotes, RTs did not expand to near present-day levels until the beginning of mammalian evolution, with the majority of currently fixed elements having inserted in the early primate genome between 12 and 40 mya [5,6,7,8]. While primate-specific RTs were beginning to populate the genome, these early ancestors were adapting to the changing conditions that would eventually generate the modern human, and with it the modern human brain.
To provide context for the following sections related to the functional impact of these RTs, we provide below a temporal map that intersects the integration timing of primate-specific RT subfamilies with major physiological changes that represent landmarks in the evolution of the primate brain (Fig. 1). It should be noted that this intersection is not meant to indicate that RTs directly caused each of these events, but the hope is to provide a general context with which to explore the connection between RTs and neuronal function.
Phylogenetic timeline of primate evolution. The major branches represent Strepsirrhini, Platyrrhini, and five genera of Catarrhini (Macaca, Pongo, Gorilla, Pan, and Homo), with branch points denoting the hypothesized most recent common ancestor (million years ago). Drawings underneath each branch represent the increased brain volume and cortical folding for each genus. The waves of retrotransposition that have been predicted to occur within the past 63 million years in primates are shown for two major families of retrotransposons; Alu (blue) and L1 (green). For each wave of retrotransposition the names of common active subfamilies are noted. New additions to the genome driven by each wave of retrotransposition are noted in red and coinciding changes in brain structure are noted in black
For example, during the Eocene (~35–65 mya), early prosimians proliferated. These ancestors of modern-day lemurs and tarsiers had a brain that was relatively small compared to their body mass and with relatively little gyrification [9, 10]. The early prosimian genome supported retrotransposition of ancient families of L1 (L1M, L1PA, and L1PB) and HERV. This period of time also witnessed the explosion of the extant Alu subfamilies AluJ and AluS, which had previously existed only as the smaller fragments FLAM and FRAM and which are present in all known primates. The split between platyrrhines and catarrhines occurred around 35 mya [11]. Platyrrhines are similar to prosimians in that they have a relatively small brain size as well as shallower sulci, or folds in the brain, than catarrhines; both of these features can impact higher cognitive functions [9, 12, 13]. The central sulcus, a prominent cortical fold, began to increase in size compared to the overall cortical surface area after the split of lesser apes from catarrhines [14, 15]. Around 25 mya AluY and the L1PA8, L1PA7, and L1PA6 families began to take over [5, 8]. At that time, the split occurred between hominoids and cercopithecoids [11]. In RT history, this was the time when SVAs first originated and the reign of the L1PA4/5 subfamilies began [16]. This time was also associated with an increase in size of the frontal cortex. The final point on our RT-brain map is the time of the largest increase in primate brain size, which occurred within the past ~5 my [17]. After the Pan-Homo split the human brain departed from the norm of allometric scaling [9]. The RTs that flourished during this period in Homo were the subfamilies L1Hs, AluYa5, AluYb8, and SVA, and these elements are still the most active elements in the human genome. The current estimated rates of germline insertion are ~1 in 20 births for Alu, 1 in 270 births for L1, and 1 in 916 births for SVA [18,19,20].
As with any mutation these insertions were more often than not likely to have neutral or even deleterious impacts on the population. However, in the odd case that a new insertion provided a selective advantage, the question becomes whether the proliferating subfamily of RT contained unique properties that enabled a more efficient and directed impact on neuronal evolution.
The susceptibility and resilience of brain genes to retrotransposition
The impact of a RT is largely dependent on its genomic location. Other than a small subset of regions (for example, the NF1 gene), there has been no strong evidence to suggest that there are specific hotspots of RT integration [21]. However, at random chance, RTs are more likely to reside within the introns of long genes. Importantly, long genes are enriched for genes that are expressed in the brain, specifically expressed within neuronal populations, and involved in synapse formation, cell adhesion, and other neuronal-specific processes [22,23,24]. Given their length, these neuronal genes have an increased susceptibility to have retrotransposons insert within the transcriptional boundaries (Fig. 2). However, the presence of a RT does not equal the function of that element; therefore, it is important to examine whether retrotransposition within these long introns has a significant impact on neuronal function. Although this information is not directly known for the human brain, the cross-species phenomenon of purifying selection of RTs within genes indicates that selection may be acting on these elements, although the influence of genetic drift can not be ruled out [25,26,27]. Furthermore, studies examining the comparative density of RTs within long versus short introns have identified that germline RTs are reduced in prevalence within genes and as they become closer to an exon, and this effect is more stringent for RTs in the antisense orientation [27, 28]; however, it is unclear whether this effect is consistent in long introns. As these reports are limited to examining the impact of germline RTs, the question remains open as to what extent insertions into introns directly impact neuronal function.
RT copy number as a function of gene length. RTs consisted of all repeat masked elements, only L1, or only Alu. Gene length was calculated as transcription start to transcription stop. The element count was normalized by the total number of elements across all genes. Genes were then subdivided into four bins as noted by diagonal lines and the top Gene Ontology term (molecular function) was noted along with the Benjamini corrected p value and top genes [93]. Note that, similar to findings from previous studies, the largest genes are commonly cell adhesion molecules, channel genes, and calcium ion binding genes that are important for neuronal function [23, 24]
One known consequence of intronic RTs that is heightened within the brain is RNA editing of the primate-specific Alu RT. Brain-related genes such as those that are important for neuronal excitability have an increased propensity to undergo RNA editing due to their enrichment of intronic Alu, which is a primary recruiting signal for the ADAR enzyme that catalyzes these A to I transitions [29,30,31,32,33,34]. These RT-driven editing events have also been shown to alter the function of a neuron and this is reviewed in detail in Rosenthal and Seeburg [29]. For example, in human cells, an AluJ element is present in an intron of the GABA receptor GABRA3. This element promotes A-to-I editing of a neighboring exon, converting an isoleucine to methionine, which then results in altered sensitivity and deactivation of GABRA3 receptors [32, 35, 36]. The functional impact of this AluJ element establishes the proof-of-principle that Alu RTs can supply an additional level of functional modulation to neuronal genes through A-to-I editing. Given the millions of primate-specific Alu loci and their propensity to undergo editing, it is likely that at least a small subset of these elements impact neuronal function beyond just the GABRA3 receptor.
RTs also bring regulatory elements to gene regions; a concept that was initially posed by Britten and Davidson [37] and which was recently reviewed comprehensively in Chuong, Elde, and Feschotte [38]. For example, RTs that were active before the split of Platyrrhines from Catarrhines distributed DNA binding elements that are important for development throughout the primate genome [39]. The functionality of these primate-specific elements remains unclear as even the impact of RTs that integrated within early mammalian lineages is still under debate. For example, work from the Noonan lab showed that mammalian neocortical enhancers were not likely the result of transposons having immediately taken on a functional role upon insertion as the repeats with enhancer activity did not show signatures of conservation [40]. This finding potentially opposes the Britten and Davidson hypothesis; however, the MER130 family seems to be at least one exception [41]. The Bejerano lab identified that MER130 elements present within a highly specific set of p300-bound neocortical enhancers indeed displayed enhancer activity using an in vitro assay [41]. Importantly, these two reports are not mutually exclusive as they indicate that although RTs may display neocortical enhancer activity, as a general rule they are not directly advantageous sans additional mutations.
Further reports on the impact of RTs in development focus on the retinoic acid response which is an integral component of neurogenesis [42,43,44,45,46]. In humans, more than 90% of retinoic acid response elements (DR2) are derived from Alu elements that were active both before the split from prosimians (AluS) and after the split between platyrrhines and catarrhines (AluY) [47, 48]. Although it is unclear whether these DR2-containing Alus are important in human neurogenesis, they do display function in the human stem cell. With the addition of retinoic acid, DR2-containing Alus are actively transcribed and subsequently broken down into small RNAs that are required for the proper regulation of human stem-cell proliferation [49, 50]. Given the importance of the retinoic acid response in neuronal differentiation [44,45,46], future studies could benefit from determining whether these primate-specific DR2-containing Alus are analogously important for human neurogenesis. An additional binding signal, the estrogen response element for estrogen receptor-alpha, is similarly present largely due to the expansion of the ancient AluS subfamily [51, 52]. Examination of these elements in vitro showed that they are indeed bound by ER-alpha and functionally modulate local transcription [51, 53]. However, a study examining estrogen response in breast cancer showed that Alus were not preferentially bound in this cell line, indicating that caution must be used when attempting to link the presence of a binding site with the functional utility of that site [54].
A recent discovery from our lab suggests that transposons may also have made previously unknown contributions to the proteome [55]. Primate L1s encode a third open reading frame, ORF0, in the antisense orientation. ORF0 dates back to the L1PA8 subfamily, which was active after the split between Catarrhines and Platyrrhines. What makes this ORF particularly interesting is the presence of splice donor sites within its coding sequence. These donor sites act in concert with splice acceptor sites within proximal exons to generate fusion proteins. Thus, L1s can generate insertion-site specific proteins in Catarrhines, including humans. Around 3000 ORF0 loci exist in the human and chimp genomes and their transcription is enriched in pluripotent cells. Considering that a number of ORF0 fusions are associated with neuronal genes, it is possible that ORF0 may contribute to primate-specific properties of the brain. Which of the fusion events have evolved functions is currently under investigation.
Advantages of RTs within the nervous system
The contribution of RTs to the health and normal function of the human brain is still readily debated. Progress is inherently slowed by the same impediments that plague all genomics research, such as small effect sizes and complex phenotypes. However, disease studies that have been reviewed extensively elsewhere [56, 57] have established links between RTs and neurological phenotypes. Although generalized contributions such as splicing, methylation, and A-to-I editing are known, experiments are needed that directly link specific RT loci to neuronal function and behavior. So far, examples of the advantageous nature of RTs in the human brain are limited, but two cases point to an evolved role as functional non-coding RNAs (ncRNA). For example, a composite Alu/L1 sequence within the SLC7A2 gene generates an ncRNA that is vital for human brain development and results in infantile encephalopathy when mutated [58]. Similarly, a monomeric Alu present after the split of prosimians and anthropoids now encodes the functional BCYRN1 ncRNA (aka Bc200) [59, 60]. The function of BCYRN1 is to aid protein synthesis in neuronal dendrites and, interestingly, this function has been replicated in the mouse genome through convergent evolution of mouse Bc1 from the rodent SINE B2 [60,61,62,63]. Future research should shed further light on these instances of advantageous adaptation of RTs. As increasing numbers of neurotypical individuals are sequenced, the RT component underlying normal human phenotypic variation should begin to be revealed and provide a library of locations to examine experimentally. Through these tools, we will begin to answer the lingering question: to what extent do RTs impact normal human phenotypic variation?
Although the focus of this review is on non-LTR RTs in humans, evidence from non-LTR RTs and DNA transposons in non-human species can provide helpful insight into the potential for mobile elements to impact neuronal function. One piece of evidence that non-LTR RTs can take on functional roles comes from a SINE that integrated over 170 mya and which currently functions as a tissue-specific enhancer in hypothalamic neurons [64]. Mobile elements can also impact on neuronal function by the generation of new genes such as POGZ, a Pogo element that when mutated can lead to microcephaly, intellectual disability, and autism spectrum disorders [65,66,67,68]. Furthermore, the immediate early gene, Arc, which is vital for long-term memory formation, is thought to have been generated from a Ty3/Gypsy LTR [69, 70]. These, as well as numerous other examples, indicate that mobile elements, including non-LTR RTs, have impacted the evolution of neuronal function in a way that is relevant to the human brain [71,72,73,74,75].
Somatic retrotransposition in the human brain
Genomic changes in response to retrotransposition also persist on the much shorter scale of a single human lifetime. Over the past decade, interest in brain somatic retrotransposition has steadily risen, largely in response to the findings, including evidence from our lab, that RTs can mobilize in neural progenitor cells [76,77,78]. Interest was further buoyed by the repeated findings of genetic mosaicism in adult human neurons [79,80,81,82], including an increased number of RTs in the prefrontal cortex of individuals diagnosed with schizophrenia [83]. This has led to a consortium of 15 institutions, including our lab, gathering in the ‘Brain Somatic Mosaicism Network’ to tackle the issue of identifying the diverse types of mosaicism present within the human brain, including retrotransposition [84].
This continued interest from labs around the world has resulted in independent confirmation that retrotransposition is indeed a phenomenon acting not only on the timescale of human evolution but also throughout the development of a single human brain. Recent studies, including our own, using single-cell sequencing approaches have found that both L1 insertions and L1-mediated deletions are prevalent in the human brain [82, 85,86,87]. These studies estimate a rate of retrotransposition between ~0.6 and 13 somatic insertions per neuron [86, 87]. It is likely that the true rate of neuronal somatic retrotransposition in humans is somewhere between those two bounds, with 0.6 representing a lower bound due to methodological limits of detection and 13 being the upper bound limited by artifacts created during whole genome amplification. Importantly, since the experiments that calculated the lower bound have high rates of validation, it is reasonable to take 0.6 insertions per neuron as approximating a true minimum. Although 0.6 elements per neuron seems small, with approximately 86 billion neurons in the adult human brain, that comes to approximately 51 billion somatic retrotransposition events within a given individual. Even under a conservative mutational hypothesis, where a majority of new insertions are neutral, it is easy to imagine a scenario where these somatic events modulate a portion of functional heterogeneity within the human brain.
Importantly, mosaic mutations would have the potential to have a large effect on the cell that they reside in but relatively small effect on cells that are independent of that founder cell. Therefore, the more cells within a tissue rely on each other for proper function, the more likely it will be that a single mutation alters the function of that tissue. Considering the highly networked state of the brain, neurons are a particularly useful system to study the impact of somatic mosaicism since a small number of functional mutations could have far-reaching effects on neuronal circuitry. In fact, studies examining the impact of individual neurons in rodents have shown that disrupting the firing of a single neuron can affect rodent behavior [88, 89], indicating that the function of individual neurons can have a profound impact on behavioral diversity. If the clues from selection on RTs are any sign of their impact on function, then it is possible that new RT insertions can alter genetic and phenotypic heterogeneity within a brain within a single generation.
Insights and outlook
Cognitive differences exist between humans and nonhuman primates that allow for the development of sophisticated behaviors such as language, self-awareness, symbolic thought, and cultural learning [88]. Although great advances have been made since the first discovery of RTs in humans, the functional contribution of RTs to these differences is still largely unknown. While previous efforts attempted to home in on RT function by examining signatures of selection, current efforts to increase our understanding are beginning to take advantage of high-throughput sequencing approaches to incorporate information from thousands of individuals as well as across multiple species. These future efforts will be aided by a more detailed and comprehensive approach to cataloguing and sharing this information. While RTs can influence the host independently of mobility, the number of active elements and their genomic locations will be instrumental in understanding their role in human biology, especially in somatic cells. In the future, we will have to move away from simply annotating transposable elements and take their context and activity state into consideration. Only a multi-species, comprehensive analysis and catalog of transposable elements will allow a true understanding of the influence of transposable elements on human brain evolution.
The prediction that RTs might have consequences for human brain development can inform experiments to be performed with new cell culture techniques, providing a powerful tool to probe the impact of RTs on human brain evolution. Currently, the vast amount of information available for comparative studies between humans and our closest relatives comes from DNA/RNA samples extracted from preserved (post-mortem) tissues. These samples do not always fairly represent the function of a region due to confounding effects of environment and development. Ideally, the identification of differences in genetic makeup between related species should be translated into phenotypic divergence in a controlled setting. Cell culture models utilizing neurons derived from non-human primate induced pluripotent stem cells could provide new insights into human adaptation features and could be genetically modified to determine the effects of individual loci of species-specific RTs. For example, in cell culture, RT loci that are predicted to have a functional impact can be mutated with gene editing technology (for example, CRISPR/Cas9 or the TALEN system), thereby enabling a direct study of the impact of RTs across neurons from different primate species. Work from our lab deriving induced pluripotent stem cells from primate lineages has begun to aid in these types of experiments [90,91,92]. Despite the findings presented here that clearly show that RTs have impacted the mammalian, primate, and human nervous system, the direct impact of RTs on the human brain currently remains under debate. Therefore, future studies, such as those using these induced pluripotent stem cell models, will help to define the role of RTs in the function of the human neuron.
References
Lander E, Linton L, Birren B, Nusbaum C, Zody M, Baldwin J, Devon K, Dewar K, Doyle M, FitzHugh W, Funke R, Gage D, Harris K, Heaford A, Howland J, Kann L, Lehoczky J, LeVine R, McEwan P, McKernan K, Meldrim J, Mesirov J, Miranda C, Morris W, Naylor J, Raymond C, Rosetti M, Santos R, Sheridan A, Sougnez C, Stange-Thomann N, Stojanovic N, Subramanian A, Wyman D, Rogers J, Sulston J, Ainscough R, Beck S, Bentley D, Burton J, Clee C, Carter N, Coulson A, Deadman R, Deloukas P, Dunham A, Dunham I, Durbin R, French L, Grafham D, Gregory S, Hubbard T, Humphray S, Hunt A, Jones M, Lloyd C, McMurray A, Matthews L, Mercer S, Milne S, Mullikin J, Mungall A, Plumb R, Ross M, Shownkeen R, Sims S, Waterston R, Wilson R, Hillier L, McPherson J, Marra M, Mardis E, Fulton L, Chinwalla A, Pepin K, Gish W, Chissoe S, Wendl M, Delehaunty K, Miner T, Delehaunty A, Kramer J, Cook L, Fulton R, Johnson D, Minx P, Clifton S, Hawkins T, Branscomb E, Predki P, Richardson P, Wenning S, Slezak T, Doggett N, Cheng J, Olsen A, Lucas S, Elkin C, Uberbacher E, Frazier M, Gibbs R, Muzny D, Scherer S, Bouck J, Sodergren E, Worley K, Rives C, Gorrell J, Metzker M, Naylor S, Kucherlapati R, Nelson D, Weinstock G, Sakaki Y, Fujiyama A, Hattori M, Yada T, Toyoda A, Itoh T, Kawagoe C, Watanabe H, Totoki Y, Taylor T, Weissenbach J, Heilig R, Saurin W, Artiguenave F, Brottier P, Bruls T, Pelletier E, Robert C, Wincker P, Rosenthal A, Platzer M, Nyakatura G, Taudien S, Rump A, Yang H, Yu J, Wang J, Huang G, Gu J, Hood L, Rowen L, Madan A, Qin S, Davis R, Federspiel N, Abola A, Proctor M, Myers R, Schmutz J, Dickson M, Grimwood J, Cox DR, Olson M, Kaul R, Shimizu N, Kawasaki K, Minoshima S, Evans G, Athanasiou M, Schultz R, Roe B, Chen F, Pan H, Ramser J, Lehrach H, Reinhardt R, McCombie W, De La Bastide M, Dedhia N, Blocker H, Hornischer K, Nordsiek G, Agarwala R, Aravind L, Bailey J, Bateman A, Batzoglou S, Birney E, Bork P, Brown D, Burge C, Cerutti L, Chen H, Church D, Clamp M, Copley R, Doerks T, Eddy SR, Eichler E, Furey T, Galagan J, Gilbert J, Harmon C, Hayashizaki Y, Haussler D, Hermjakob H, Hokamp K, Jang W, Johnson L, Jones T, Kasif S, Kaspryzk A, Kennedy S, Kent W, Kitts P, Koonin E, Korf I, Kulp D, Lancet D, Lowe T, McLysaght A, Mikkelsen T, Moran J, Mulder N, Pollara V, Ponting C, Schuler G, Schultz J, Slater G, Smit A, Stupka E, Szustakowki J, Thierry-Mieg D, Thierry-Mieg J, Wagner L, Wallis J, Wheeler R, Williams A, Wolf Y, Wolfe K, Yang S, Yeh R, Collins F, Guyer MS, Peterson J, Felsenfeld A, Wetterstrand K, Patrinos A, Morgan M, Conso IHGS. Initial sequencing and analysis of the human genome. Nature. 2001;409(6822):860–921.
Burke WD, Malik HS, Rich SM, Eickbush TH. Ancient lineages of non-LTR retrotransposons in the primitive eukaryote, Giardia lamblia. Mol Biol Evol. 2002;19(5):619–30.
Malik HS, Burke WD, Eickbush TH. The age and evolution of non-LTR retrotransposable elements. Mol Biol Evol. 1999;16(6):793–805.
Goodwin TJD, Ormandy JE, Poulter RTM. L1-like non-LTR retrotransposons in the yeast Candida albicans. Curr Genet. 2001;39(2):83–91.
Batzer MA, Deininger PL. Alu repeats and human genomic diversity. Nat Rev Genet. 2002;3(5):370–9.
Mager DL, Freeman JD. HERV-H endogenous retroviruses: presence in the New World branch but amplification in the Old World primate lineage. Virology. 1995;213(2):395–404.
Ostertag EM, Goodier JL, Zhang Y, Kazazian HH. Report SVA Elements Are Nonautonomous Retrotransposons that Cause Disease in Humans. Am J Hum Genet. 2003;73:1444–51.
Khan H, Smit A, Boissinot S. Molecular evolution and tempo of amplification of human LINE-1 retrotransposons since the origin of primates. Genome Res. 2006;16(1):78–87.
Gilbert SL, Dobyns WB, Lahn BT. Genetic links between brain development and brain evolution. Nat Rev Genet. 2005;6(7):581–90.
Zilles K, Armstrong E, Moser KH, Schleicher A, Stephan H. Gyrification in the cerebral cortex of primates. Brain Behav Evol. 1989;34(3):143–50.
Schrago CG, Russo CAM. Timing the origin of New World monkeys. Mol Biol Evol. 2003;20(10):1620–5.
Stevens NJ, Seiffert ER, O’Connor PM, Roberts EM, Schmitz MD, Krause C, Gorscak E, Ngasala S, Hieronymus TL, Temu J. Palaeontological evidence for an Oligocene divergence between Old World monkeys and apes. Nature. 2013;497(7451):611–4.
Redmond JC. Cranial capacity and performance on delay-response task correlated with principal sulcus length in monkeys. Am J Phys Anthropol. 1999;109(1):33–40.
Hopkins WD, Meguerditchian A, Coulon O, Bogart S, Mangin J-F, Sherwood CC, Grabowski MW, Bennet AJ, Pierre PJ, Woods R, Hof PR, Vauclair J. Evolution of the central sulcus morphology in primates. Brain Behav Evol. 2014;84(1):19–30.
Clark DA, Mitra PP, Wang SS-H. Scalable architecture in mammalian brains. Nature. 2001;411:189.
Wang H, Xing J, Grover D, Hedges DJ, Han K, Walker JA, Batzer MA. SVA elements: a hominid-specific retroposon family. J Mol Biol. 2005;354(4):994–1007.
Semendeferi K, Lu A, Schenker N, Damasio H. Humans and great apes share a large frontal cortex. Nat Neurosci. 2002;5(3):272.
Cordaux R, Hedges DJ, Herke SW, Batzer MA. Estimating the retrotransposition rate of human Alu elements. Gene. 2006;373:134–7.
Ewing AD, Kazazian HH. High-throughput sequencing reveals extensive variation in human-specific L1 content in individual human genomes. Genome Res. 2010;20(9):1262–70.
Xing J, Zhang Y, Han K, Salem AH, Sen SK, Huff CD, Zhou Q, Kirkness EF, Levy S, Batzer MA, Jorde LB. Mobile elements create structural variation: analysis of a complete human genome. Genome Res. 2009;19(9):1516–26.
Wimmer K, Callens T, Wernstedt A, Messiaen L. The NF1 gene contains hotspots for L1 endonuclease-dependent De Novo insertion. Plos Genet. 2011;7(11), e1002371.
Gabel HW, Kinde B, Stroud H, Gilbert CS, Harmin DA, Kastan NR, Hemberg M, Ebert DH, Greenberg ME. Disruption of DNA-methylation-dependent long gene repression in Rett syndrome. Nature. 2015;522:89–93.
Zylka MJ, Simon JM, Philpot BD. Gene length matters in neurons. Neuron. 2015;86(2):353–5.
Sibley CR, Emmett W, Blazquez L, Faro A, Haberman N, Briese M, Trabzuni D, Ryten M, Weale ME, Hardy J, Modic M, Curk T, Wilson SW, Plagnol V, Ule J. Recursive splicing in long vertebrate genes. Nature. 2015;521(7552):371–5.
Baucom RS, Estill JC, Leebens-Mack J, Bennetzen JL. Natural selection on gene function drives the evolution of LTR retrotransposon families in the rice genome. Genome Res. 2009;19(2):243–54.
Pereira V. Insertion bias and purifying selection of retrotransposons in the Arabidopsis thaliana genome. Genome Biol. 2004;5(10):R79.
Zhang Y, Mager DL. Gene properties and chromatin state influence the accumulation of transposable elements in genes. PLoS One. 2012;7(1), e30158.
Zhang Y, Romanish MT, Mager DL. Distributions of transposable elements reveal hazardous zones in mammalian introns. Rigoutsos I, editor. Plos Comput Biol. 2011;7(5):13.
Rosenthal JJC, Seeburg PH. A-to-I RNA Editing: Effects on Proteins Key to Neural Excitability. Neuron. 2012;74:432–9.
Athanasiadis A, Rich A, Maas S. Widespread A-to-I RNA editing of Alu-containing mRNAs in the human transcriptome. Plos Biol. 2004;2(12), e391.
Iizasa H, Nishikura K. A new function for the RNA-editing enzyme ADAR1. Nat Immunol. 2009;10(1):16–8.
Daniel C, Silberberg G, Behm M, Ohman M. Alu elements shape the primate transcriptome by cis-regulation of RNA editing. Genome Biol. 2014;15:R28.
Ramaswami G, Lin W, Piskol R, Tan MH, Davis C, Li JB. Accurate identification of human Alu and non-Alu RNA editing sites. Nat Methods. 2012;9(6):579–81.
Li JB, Church GM. Deciphering the functions and regulation of brain-enriched A-to-I RNA editing. Nat Neurosci. 2013;16:1518–22.
Ohlson J, Pedersen JS, Haussler D, Ohman M. Editing modifies the GABA(A) receptor subunit alpha3. RNA. 2007;13:698–703.
Nimmich ML, Heidelberg LS, Fisher JL. RNA editing of the GABA(A) receptor alpha3 subunit alters the functional properties of recombinant receptors. Neurosci Res. 2009;63(4):288–93.
Britten RJ, Davidson EH. Gene Regulation for Higher Cells: A Theory. Science (80- ). 1969;165(3891):349.
Chuong EB, Elde NC, Feschotte C. Regulatory activities of transposable elements: from conflicts to benefits. Nat Rev Genet. 2017;18:71–86.
Gifford WD, Pfaff SL, Macfarlan TS. Transposable elements as genetic regulatory substrates in early development. Trends Cell Biol. 2013;23(5):218–26.
Emera D, Yin J, Reilly SK, Gockley J, Noonan JP. Origin and evolution of developmental enhancers in the mammalian neocortex. Proc Natl Acad Sci. 2016;113(19):E2617–26.
Notwell JH, Chung T, Heavner W, Bejerano G. A family of transposable elements co-opted into developmental enhancers in the mouse neocortex. Nat Commun. 2015;6:6644.
Walsh T, McClellan JM, McCarthy SE, Addington AM, Pierce SB, Cooper GM, Nord AS, Kusenda M, Malhotra D, Bhandari A, Stray SM, Rippey CF, Roccanova P, Makarov V, Lakshmi B, Findling RL, Sikich L, Stromberg T, Merriman B, Gogtay N, Butler P, Eckstrand K, Noory L, Gochman P, Long R, Chen Z, Davis S, Baker C, Eichler EE, Meltzer PS, Nelson SF, Singleton AB, Lee MK, Rapoport JL, King M-C, Sebat J. Rare structural variants disrupt multiple genes in neurodevelopmental pathways in schizophrenia. Science (80- ). 2008;320(5875):539–43.
Andrews PW. Retinoic acid induces neuronal differentiation of a cloned human embryonal carcinoma cell line in vitro. Dev Bio. 1984;103(2):285–93.
Shiotsugu J. Multiple points of interaction between retinoic acid and FGF signaling during embryonic axis formation. Development. 2004;131(11):2653–67.
Maden M, Gale E, Kostetskii I, Zile M. Vitamin A-deficient quail embryos have half a hindbrain and other neural defects. Current biology. 1996;6(4):417–26.
Mccaffery P, Drager UC. High levels of a retinoic acid-generating dehydrogenase in the meso-telencephalic dopamine system. Proc Natl Acad Sci U S A. 1994;91(16):7772–6.
Vansant G, Reynolds WF. The consensus sequence of a major Alu subfamily contains a functional retinoic acid response element. Proc Natl Acad Sci U S A. 1995;92(18):8229–33.
Laperriere D, Wang T-T, White JH, Mader S. Widespread Alu repeat-driven expansion of consensus DR2 retinoic acid response elements during primate evolution. BMC Genomics. 2007;8:23.
Hu Q, Tanasa B, Trabucchi M, Li W, Zhang J, Ohgi KA, Rose DW, Glass CK, Rosenfeld MG. DICER- and AGO3-dependent generation of retinoic acid–induced DR2 Alu RNAs regulates human stem cell proliferation. Nat Struct Mol Biol. 2012;19(11):1168–75.
Morales-Hernández A, González-Rico FJ, Román AC, Rico-Leo E, Alvarez-Barrientos A, Sánchez L, Macia A, Heras SR, Garcyá-Perez JL, Merino JM, Fernandez-Salguero PM. Alu retrotransposons promote differentiation of human carcinoma cells through the aryl hydrocarbon receptor. Nucleic Acids Res. 2016;44(10):4665–83.
Norris J, Fan D, Aleman C, Marks JR, Futreal PA, Wiseman RW, Iglehart JD, Deininger PL, McDonnell DP. Identification of a new subclass of Alu DNA repeats which can function as estrogen receptor-dependent transcriptional enhancers. J Biol Chem. 1995;270(39):22777–82.
Polak P, Domany E. Alu elements contain many binding sites for transcription factors and may play a role in regulation of developmental processes. BMC Genomics. 2006;7(1):133.
Mason CE, Shu FJ, Wang C, Session RM, Kallen RG, Sidell N, Yu T, Liu MH, Cheung E, Kallen CB. Location analysis for the estrogen receptor-α reveals binding to diverse ERE sequences and widespread binding within repetitive DNA elements. Nucleic Acids Res. 2010;38(7):2355–68.
Lin C-Y, Ström A, Vega VB, Li Kong S, Li Yeo A, Thomsen JS, Chan WC, Doray B, Bangarusamy DK, Ramasamy A, Vergara LA, Tang S, Chong A, Bajic VB, Miller LD, Gustafsson J-Å, Liu ET. Discovery of estrogen receptor α target genes and response elements in breast tumor cells. Genome Biol. 2004;5(9):R66.
Denli AM, Narvaiza I, Kerman BE, Pena M, Benner C, Marchetto MCN, Diedrich JK, Aslanian A, Ma J, Moresco JJ, Moore L, Hunter T, Saghatelian A, Gage FH. Primate-specific ORF0 contributes to retrotransposon-mediated diversity. Cell. 2015;163(3):583–93.
Belancio VP, Hedges DJ, Deininger P. Mammalian non-LTR retrotransposons: for better or worse, in sickness and in health. Genome Res. 2008;18(3):343–58.
Hancks DC, Kazazian HH. Roles for retrotransposon insertions in human disease. Mob DNA. 2016;7:9.
Cartault F, Munier P, Benko E, Desguerre I, Hanein S, Boddaert N, Bandiera S, Vellayoudom J, Krejbich-Trotot P, Bintner M, Hoarau JJ, Girard M, Genin E, De Lonlay P, Fourmaintraux A, Naville M, Rodriguez D, Feingold J, Renouil M, Munnich A, Westhof E, Fahling M, Lyonnet S, Henrion-Caude A. Mutation in a primate-conserved retrotransposon reveals a noncoding RNA as a mediator of infantile encephalopathy. Proc Natl Acad Sci U S A. 2012;109(13):1–6.
Kuryshev VY, Skryabin BV, Kremerskothen J, Jurka J, Brosius J. Birth of a gene: locus of neuronal BC200 snmRNA in three prosimians and human BC200 pseudogenes as archives of change in the Anthropoidea lineage. J Mol Biol. 2001;309(5):1049–66.
Skryabin BV, Kremerskothen J, Vassilacopoulou D, Disotell TR, Kapitonov VV, Jurka J, Brosius J. The BC200 RNA gene and its neural expression are conserved in Anthropoidea (Primates). J Mol Evol. 1998;47(6):677–85.
Tiedge H, Fremeau RT, Weinstock PH, Arancio O, Brosius J. Dendritic location of neural BC1 RNA. Proc Natl Acad Sci U S A. 1991;88(March):2093–7.
Wang H, Iacoangeli A, Popp S, Muslimov IA, Imataka H, Sonenberg N, Lomakin IB, Tiedge H. Dendritic BC1 RNA: functional role in regulation of translation initiation. J Neurosci. 2002;22(23):10232–41.
Kondrashov AV, Kiefmann M, Ebnet K, Khanam T, Muddashetty RS, Brosius J. Inhibitory effect of naked neural BC1 RNA or BC200 RNA on eukaryotic in vitro translation systems is reversed by poly(A)-binding protein (PABP). J Mol Biol. 2005;353(1):88–103.
Santangelo AM, De Souza FSJ, Franchini LF, Bumaschny VF, Low MJ, Rubinstein M. Ancient exaptation of a CORE-SINE retroposon into a highly conserved mammalian neuronal enhancer of the proopiomelanocortin gene. PLoS Genet. 2007;3(10):1813–26.
Tan B, Zou Y, Zhang Y, Zhang R, Ou J, Shen Y, Zhao J, Luo X, Guo J, Zeng L, Hu Y, Zheng Y, Pan Q, Liang D, Wu L. A novel de novo POGZ mutation in a patient with intellectual disability. J Hum Genet. 2016;61(4):357–9.
Bartholomeeusen K, Christ F, Hendrix J, Rain JC, Emiliani S, Benarous R, Debyser Z, Gijsbers R, de Rijck J. Lens epithelium-derived growth factor/p75 interacts with the transposase-derived DDE domain of pogZ. J Biol Chem. 2009;284(17):11467–77.
Stessman HAF, Willemsen MH, Fenckova M, Penn O, Hoischen A, Xiong B, Wang T, Hoekzema K, Vives L, Vogel I, Brunner HG, Van Der Burgt I, Ockeloen CW, Schuurs-Hoeijmakers JH, Klein Wassink-Ruiter JS, Stumpel C, Stevens SJC, Vles HS, Marcelis CM, Van Bokhoven H, Cantagrel V, Colleaux L, Nicouleau M, Lyonnet S, Bernier RA, Gerdts J, Coe BP, Romano C, Alberti A, Grillo L, Scuderi C, Nordenskjöld M, Kvarnung M, Guo H, Xia K, Piton A, Gerard B, Genevieve D, Delobel B, Lehalle D, Perrin L, Prieur F, Thevenon J, Gecz J, Shaw M, Pfundt R, Keren B, Jacquette A, Schenck A, Eichler EE, Kleefstra T. Disruption of POGZ Is Associated with Intellectual Disability and Autism Spectrum Disorders. Am J Hum Genet. 2016;98(3):541–52.
White J, Beck CR, Harel T, Posey JE, Jhangiani SN, Tang S, Farwell KD, Powis Z, Mendelsohn NJ, Baker JA, Pollack L, Mason KJ, Wierenga KJ, Arrington DK, Hall M, Psychogios A, Fairbrother L, Walkiewicz M, Person RE, Niu Z, Zhang J, Rosenfeld JA, Muzny DM, Eng C, Beaudet AL, Lupski JR, Boerwinkle E, Gibbs RA, Yang Y, Xia F, Sutton VR. POGZ truncating alleles cause syndromic intellectual disability. Genome Med. 2016;8(1):3.
Zhang W, Wu J, Ward MD, Yang S, Chuang YA, Xiao M, Li R, Leahy DJ, Worley PF. Structural basis of arc binding to synaptic proteins: Implications for cognitive disease. Neuron. 2015;86(2):490–500.
Messaoudi E, Kanhema T, Soule J, Tiron A, Dagyte G, da Silva B, Bramham CR. Sustained Arc/Arg3.1 Synthesis Controls Long-Term Potentiation Consolidation through Regulation of Local Actin Polymerization in the Dentate Gyrus In Vivo. J Neurosci. 2007;27(39):10445–55.
Toth M, Grimsby J, Buzsaki G, Donovan GP. Epileptic seizures caused by inactivation of a novel gene, jerky, related to centromere binding protein–B in transgenic mice. Nat Genet. 1995;11(1):71–5.
Liu W, Seto J, Donovan G, Toth M. Jerky, a Protein Deficient in a Mouse Epilepsy Model, Is Associated with Translationally Inactive mRNA in Neurons. J Neurosci. 2002;22(1):176–82.
Irie M, Yoshikawa M, Ono R, Iwafune H, Furuse T, Yamada I, Wakana S, Yamashita Y, Abe T, Ishino F, Kaneko-Ishino T. Cognitive Function Related to the Sirh11/Zcchc16 Gene Acquired from an LTR Retrotransposon in Eutherians. PLoS Genet. 2015;11(9), e1005521.
Pavelitz T, Gray LT, Padilla SL, Bailey AD, Weiner AM. PGBD5: a neural-specific intron-containing piggyBac transposase domesticated over 500 million years ago and conserved from cephalochordates to humans. Mob DNA. 2013;4(1):23.
Henssen AG, Henaff E, Jiang E, Eisenberg AR, Carson JR, Villasante CM, Ray M, Still E, Burns M, Gandara J, Feschotte C, Mason CE, Kentsis A. Genomic DNA transposition induced by human PGBD5. eLife. 2015;4:e10565.
Coufal NG, Garcia-Perez JL, Peng GE, Yeo GW, Mu Y, Lovci MT, Morell M, O’Shea KS, Moran JV, Gage FH. L1 retrotransposition in human neural progenitor cells. Nature. 2009;460(7259):1127–31.
Muotri AR, Chu VT, Marchetto MCN, Deng W, Moran JV, Gage FH. Somatic mosaicism in neuronal precursor cells mediated by L1 retrotransposition. Nature. 2005;435(7044):903–10.
Muotri AR, Marchetto MC, Coufal NG, Oefner R, Yeo G, Nakashima K, Gage FH. L1 retrotransposition in neurons is modulated by MeCP2. Nature. 2010;468(7322):443–6.
Singer T, McConnell MJ, Marchetto MCN, Coufal NG, Gage FH. LINE-1 retrotransposons: Mediators of somatic variation in neuronal genomes? Trends Neurosci. 2010;33(8):345–54.
McConnell MJ, Lindberg MR, Brennand KJ, Piper JC, Voet T, Cowing-Zitron C, Shumilina S, Lasken RS, Vermeesch JR, Hall IM, Gage FH. Mosaic copy number variation in human neurons. Science. 2013;342(6158):632–7.
Wei PC, Chang AN, Kao J, Du Z, Meyers RM, Alt FW, Schwer B. Long Neural Genes Harbor Recurrent DNA Break Clusters in Neural Stem/Progenitor Cells. Cell. 2016;164(4):644–55.
Baillie JK, Barnett MW, Upton KR, Gerhardt DJ, Richmond TA, De Sapio F, Brennan PM, Rizzu P, Smith S, Fell M, Talbot RT, Gustincich S, Freeman TC, Mattick JS, Hume DA, Heutink P, Carninci P, Jeddeloh JA, Faulkner GJ. Somatic retrotransposition alters the genetic landscape of the human brain. Nature. 2011;479(7374):534–7.
Bundo M, Toyoshima M, Okada Y, Akamatsu W, Ueda J, Nemoto-Miyauchi T, Sunaga F, Toritsuka M, Ikawa D, Kakita A, Kato M, Kasai K, Kishimoto T, Nawa H, Okano H, Yoshikawa T, Kato T, Iwamoto K. Increased L1 retrotransposition in the neuronal genome in schizophrenia. Neuron. 2014; 81(2):306–13.
McConnell MJ, Moran J V., Abyzov A, Akbarian S, Bae T, Cortes-Ciriano I, Erwin JA, Fasching L, Flasch DA, Freed D, Ganz J, Jaffe AE, Kwan KY, Kwon M, Lodato MA, Mills RE, Paquola ACM, Rodin RE, Rosenbluh C, Sestan N, Sherman MA, Shin JH, Song S, Straub RE, Thorpe J, Weinberger DR, Urban AE, Zhou B, Gage FH, Lehner T, Senthil G, Walsh CA, Chess A, Courchesne E, Gleeson JG, Kidd JM, Park PJ, Pevsner J, Vaccarino FM. Intersection of diverse neuronal genomes and neuropsychiatric disease: The Brain Somatic Mosaicism Network. Science. 2017;356(6336):eaal1641.
Erwin JA, Paquola ACM, Singer T, Gallina I, Novotny M, Quayle C, Bedrosian TA, Alves FIA, Butcher CR, Herdy JR, Sarkar A, Lasken RS, Muotri AR, Gage FH. L1-associated genomic regions are deleted in somatic cells of the healthy human brain. Nat Neurosci. 2016;9(12):1583–91.
Upton KR, Gerhardt DJ, Jesuadian JS, Richardson SR, Sanchez-Luque FJ, Bodea GO, Ewing AD, Salvador-Palomeque C, Van Der Knaap MS, Brennan PM, Vanderver A, Faulkner GJ. Ubiquitous L1 mosaicism in hippocampal neurons. Cell. 2015;161(2):228–39.
Evrony GD, Cai X, Lee E, Hills LB, Elhosary PC, Lehmann HS, Parker JJ, Atabay KD, Gilmore EC, Poduri A, Park PJ, Walsh CA. Single-neuron sequencing analysis of l1 retrotransposition and somatic mutation in the human brain. Cell. 2012;151(3):483–96.
Houweling AR, Brecht M. Behavioural report of single neuron stimulation in somatosensory cortex. Neuroforum. 2008;14(1):174–5.
Li C-YT, Poo M-M, Dan Y. Burst spiking of a single cortical neuron modifies global brain state. Science. 2009;324(5927):643–6.
Hrvoj-Mihic B, Marchetto MCN, Gage FH, Semendeferi K, Muotri AR. Novel tools, classic techniques: Evolutionary studies using primate pluripotent stem cells. Biol Psychiatry. 2014;75:929–35. Elsevier USA.
Otani T, Marchetto MC, Gage FH, Simons BD, Livesey FJ. 2D and 3D Stem Cell Models of Primate Cortical Development Identify Species-Specific Differences in Progenitor Behavior Contributing to Brain Size. Cell Stem Cell. 2016;18(4):467–80.
Marchetto MCN, Narvaiza I, Denli AM, Benner C, Lazzarini TA, Nathanson JL, Paquola ACM, Desai KN, Herai RH, Weitzman MD, Yeo GW, Muotri AR, Gage FH. Differential L1 regulation in pluripotent stem cells of humans and apes. Nature. 2013;503(7477):525–9.
Huang DW, Lempicki RA, Sherman BT. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4(1):44–57.
Acknowledgments
We would like to thank Mary Lynn Gage for her assistance in editing and Jamie Simon for his help on the figure design. We would also like to thank Cedric Feschotte and John McCormick for their helpful insight during the preparation of this manuscript. We further thank our funding sources: NIH R01 MH095741, NIH U01 MH106882, The G. Harold and Leila Y. Mathers Charitable Foundation, The Leona M. and Harry B. Helmsley Charitable Trust (grant #2012-PG-MED00), The Engman Foundation, Annette C. Merle-Smith, Lynn and Ed Streim, and Takeda Pharmaceutical International.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Linker, S.B., Marchetto, M.C., Narvaiza, I. et al. Examining non-LTR retrotransposons in the context of the evolving primate brain. BMC Biol 15, 68 (2017). https://doi.org/10.1186/s12915-017-0409-z
Published:
DOI: https://doi.org/10.1186/s12915-017-0409-z