Abstract
Background
Community acquired pneumonia (CAP) is a high incidence disease resulting in about 260,000 hospital admissions per year in Germany, more than myocardial infarction or stroke. Worldwide, CAP is the most frequent infectious disease with high lethality ranging from 1.2 % in those 20–29 years old to over 10 % in patients older than 70 years, even in industrial nations. CAP poses numerous medical challenges, which the PROGRESS (Pneumonia Research Network on Genetic Resistance and Susceptibility for the Evolution of Severe Sepsis) network aims to tackle: Operationalization of disease severity throughout the course of disease, outcome prediction for hospitalized patients and prediction of transitions from uncomplicated CAP to severe CAP, and finally, to CAP with sepsis and organ failure as a life-threatening condition. It is a major aim of PROGRESS to understand and predict patient heterogeneity regarding outcome in the hospital and to develop novel treatment concepts.
Methods
PROGRESS was designed as a clinical, observational, multi-center study of patients with CAP requiring hospitalization. More than 1600 patients selected for low burden of co-morbidities have been enrolled, aiming at a total of 3000. Course of disease, along with therapy, was closely monitored by daily assessments and long-term follow-up. Daily blood samples allow in depth molecular-genetic characterization of patients. We established a well-organized workflow for sample logistics and a comprehensive data management system to collect and manage data from more than 50 study centers in Germany and Austria. Samples are stored in a central biobank and clinical data are stored in a central data base which also integrates all data from molecular assessments.
Discussion
With the PROGRESS study, we established a comprehensive data base of high quality clinical and molecular data allowing investigation of pressing research questions regarding CAP. In-depth molecular characterization will contribute to the discovery of disease mechanisms and establishment of diagnostic and predictive biomarkers. A strength of PROGRESS is the focus on younger patients with low burden of co-morbidities, allowing a more direct look at host biology with less confounding. As a resulting limitation, insights from PROGRESS will require validation in representative patient cohorts to assess clinical utility.
Trial registration
The PROGRESS study was retrospectively registered on May 24th, 2016 with ClinicalTrials.gov: NCT02782013
Similar content being viewed by others
Background
Community acquired pneumonia (CAP) is truly a high incidence disease (5–10 cases/1000 inhabitants, up to 80 cases/1000 nursing home residents) which results in about 260,000 hospital admissions per year in Germany with an overall mortality for hospitalized CAP of about 13 % [2, 19, 35]. CAP is the most frequent infectious disease worldwide with high lethality [37]. Pneumonia induced severe sepsis frequently jeopardizes successful patient outcome in many medical situations [1]. Severe sepsis and septic shock are the main causes of death on non-coronary intensive care units [1, 4, 7]. According to a recent study by Dremsizov et al., almost half of all patients with severe CAP developed sepsis in the PORT study [6]. In Germany, the SepNet consortium has collected important data on CAP as cause of sepsis in a study of prevalence of severe sepsis and septic shock in German Intensive Care Units (ICUs). In a prospective, observational, cross-sectional study (454 randomly selected ICUs in 310 hospitals in Germany, 3.877 patients screened), respiratory tract infections were identified in 62.9 % of patients with severe sepsis; 39.1 % of infections were considered as community acquired [7]. There is clear evidence that in adults incidence of CAP requiring hospitalization increases with age [2, 9]. Lethality shows a similar pattern: AQUA-data reveal that lethality of hospitalized CAP goes up from about 1.2 % in those 20–29 years old to 12.3 % in the group 70–79 years old and even 22.5 % in the group older than 90 years [2]. In consequence, the health and economic impact of pneumonia is substantial [35].
To date, substantial research efforts over many years have illuminated the origin of CAP, its epidemiology and potential new avenues for treatment [31]. Nevertheless, major challenges remain: While several scoring systems have been developed, their main strength lies in identifying patients not requiring treatment in a hospital [18]. Assessment of disease severity still mainly relies on clinical judgement and outcome prediction remains a challenge [3, 10, 21, 24]. The current lack of a validated operationalization of disease severity throughout the course of treatment in the hospital may hamper clinical research [18].
To meet these challenges and to better understand what determines individual immune responses to pneumonia we conduct a multi-centric observational study of hospitalized patients with CAP in Germany and Austria to establish a comprehensive database of high quality clinical and molecular data. Data obtained will provide the basis for clinical and molecular scoring systems helping clinicians to decide which CAP patient can stay on a regular ward and who is at high risk to progress to severe pneumonia or even septic shock with necessity for intensive care including artificial respiration. The systematic analytical discovery-driven approach chosen will help to identify yet unknown targets and molecular pathomechanisms in the complex dynamics of endothelial barrier function during acute inflammation of the lung [12, 22]. These may directly improve pneumonia management (diagnostics, therapy) and contribute to a better understanding of the pathophysiology of the pneumonia-to-sepsis progression, which in turn will accelerate the identification of new treatment options.
Current research supports the notion that broad as well as in-depth molecular analyses may contribute to a better understanding of disease mechanisms and also lead to diagnostic and prognostic biomarkers [14, 29, 38]. Patients with and without infection may be distinguishable by e.g. gene expression profiles [5, 16, 28, 32]. Even causal pathogens may be detectable by analyzing host biospecimen [25]. A strong association between death from infection in adoptees and their biological, but not adoptive parents suggested a genetic influence on the risk for and outcome from infection [30]. More recently, in the GenOSept study, analysis of data from three cohorts suggested genetic variants associated with survival from sepsis due to pneumonia [26]. It appears worthwhile to study pneumonia systematically and to collect clinical and concomitant molecular data as a basis for further insights.
We here present the design of the PROGRESS study which aims to provide new and in-depth clinical and molecular data helping to elucidate variation in host response to CAP. We hope to contribute to the development of an operationalization of disease severity, to the identification of clinical and biomarkers of pneumonia progression in hospitalized patients, and to new therapeutic concepts. Here, we describe clinical and molecular assessments of the PROGRESS study to facilitate future co-operation and complementary work related to the pressing research questions in CAP.
Methods/Design
Study design and assessments
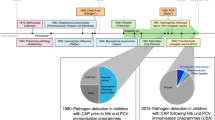
PROGRESS is a prospective multi-centric longitudinal observational study on patients hospitalized due to confirmed CAP (Fig. 1). Biomaterials are collected for broad molecular investigation. The study does not intend to influence treatment decisions; treatment is at the discretion of the attending physician. PROGRESS is designed to observe transitions from uncomplicated community acquired pneumonia (uCAP) to severe CAP (sCAP) to pneumogenic sepsis and septic shock (ssCAP). Patients are enrolled within 48 h of hospitalization with CAP. Routine clinical and laboratory data are collected for the day of hospitalization, the day of enrolment (d0), and the four following days (d1 through d4). Scores like CURB-65 [20] and PSI [11] are obtained at d0. Parameters for SOFA [33] and SIRS [27] are obtained daily on d0 through d4, along with information on treatment unit, respiratory support, medication, and newly observed clinical and microbiological findings. Blood samples for molecular analyses are collected daily on d0 through d3. At hospital discharge, information like vital status, number of days in intensive care or with respiratory support, and subsequent type of residence are documented. Patients are followed up regarding vital status, quality of life (EQ-5D [15]), and new episodes of pneumonia at days 28, 180, and 360 after enrolment. Baseline data comprise sociodemographic information, relevant aspects of medical history, and treatment prior to hospitalization.
Schematic view of the PROGRESS Study. Stars indicate the time periods or time points from which data originate. Data from study visits correspond to the time range between the previous and current study visit. Patients must be enrolled within 48 h after hospitalization. Study visits correspond to the time points when blood for biospecimen is drawn in the morning. If there would be less than 12 h between collection of biospecimen at enrolment and at visit 1, visit 1 is postponed to the following morning
Study population
PROGRESS aims at recruiting a total of 3000 CAP patients of different degrees of initial disease severity (see inclusion criteria). As PROGRESS aims to provide data on host parameters modulating the response to infection in patients whose immune system appears clinically uncompromised, several exclusion criteria apply (see exclusion criteria). PROGRESS enrolls adult male and female patients from all ethnic backgrounds.
Inclusion criteria
Study participants must be 18 years of age or older and hospitalized within the past 48 h due to CAP confirmed by a new pulmonary infiltrate on chest radiograph and more than one of the following signs and symptoms of lower respiratory tract infection: 1) Fever, 2) cough, 3) purulent sputum, 4) shortness of breath or need for respiratory support, or 5) crackling or rales on auscultation, dullness to percussion, or bronchial breathing.
Exclusion criteria
To ensure that the diagnosed pneumonia has not been contracted in a hospital and to avoid patients whose immune system may have been substantially compromised for other reasons, the following exclusion criteria are applied: Patients must not have been hospitalized for any reason within 28 days prior to hospitalization for the current episode of CAP. Patients cannot be enrolled with known HIV infection or AIDS or immunosuppressive treatments within the past six months including anti-tumor treatment, non-steroidal immunosuppressive therapy, radiation therapy, or therapy with corticosteroids ≥ 20 mg for ≥ 14 days. Further exclusion criteria comprise poststenotic pneumonia in conjunction with bronchial carcinoma, organ or bone marrow transplant, respiratory support at home via tracheostoma, cystic fibrosis, active tuberculosis, acute lung injury or acute respiratory distress syndrome for extrapulmonary reasons, massive aspiration, sepsis with extrapulmonary focus, acute pulmonary embolism, congestive heart failure NYHA-IV, or liver insufficiency Child-Pugh C stadium. Also, patients who are pregnant, breast feeding, previously participated in the PROGRESS study, or under limitation of therapy are excluded from the study.
Study centers
Patients are recruited at study centers throughout Germany and in Austria (Table 1), comprising emergency wards and wards with normal, intermediate, or intensive care. To qualify, a potential study center has to regularly treat CAP patients, have capabilities to process serum and plasma samples using a refrigerated centrifuge and store them at -20 °C, and have access to the internet. To date, more than 50 study centers have contributed over 1600 patients to PROGRESS.
Table 1 lists all study sites which so far contributed patients to the PROGRESS study. Contact to patients can be made through central study management (Petra Creutz: petra.creutz@charité.de or Peter Ahnert: peter.ahnert@imise.uni-leipzig.de).
The study further remains open for participation by interested hospitals. Before initiation of a study center, study personnel have to undergo a comprehensive training program making them familiar with the aims of the study, study procedures, electronic data entry, query management, and sample processing and logistics.
Data collection and processing
In PROGRESS, study data are collected through standardized web-based data sheets (eCRF) into a central database. PROGRESS aims to collect individual values for all parameters of interest: Scores and derived values are to be calculated centrally to avoid disparity in calculation procedures between study centers, increase opportunities for quality control, allow analysis of individual parameters, and to empower development of new scores and derivatives. Professional trial management and database software are employed for remote data entry and secure storage of clinical data, query management, and basic reporting. Plausibility of data is checked directly upon entry into the eCRF, by several hundred daily queries running on the central database, and in campaigns performed by the data management and biometry teams. Data management actively monitors resolution of identified problems, supported by clinicians from central study management if necessary. Data regarding sample collection, processing, logistics, and from molecular measurements in PROGRESS biospecimen are stored in the same database, ensuring a high level of confidence in sample identity and in matching biospecimen and data from molecular measurements with patient data.
Ethics and data protection
Written informed consent for participation in the study was obtained from all participants or, in case of critical illness, from patient’s legal representatives. The study protocol has been approved by the ethics committee of the University of Jena (registration number 2403-10/08) and all locally responsible ethics committees of all study centers. The study protocol adheres to the requirements of the Declaration of Helsinki [39] and to the ICH-GCP guideline [8].
To achieve the objective of the study, it is necessary to collect and process data and biospecimen of individual patients. Identification of patients is possible only in the study centers via a study specific patient pseudonym. All inquiries of data management to the study centers as part of quality assurance processes are using the pseudonym. Collected biospecimen are identified by a ten-digit code which is linked to the patient pseudonym at the central database. Identification of specific patients is not required at any time outside the study centers. To ensure that DNA samples cannot be linked to patient identity by anyone involved in the study and to also ensure that DNA samples can be destroyed at patient’s request, information linking DNA samples to patient identity are held by an external data trustee.
Sample collection and biobanking
Specimen to be collected daily (days d0 through d3) include serum, EDTA plasma with protease inhibitors, citrate plasma, and PAXgene blood for RNA. Samples are collected, processed, and stored at -20 °C for up to three months in the study centers. Samples are then shipped to the Integrated Biobank Jena (IBBJ), the central biobank of the PROGRESS network. A main goal was to minimize the number of pre-analytical steps in study centers to limit heterogeneity introduced by slight differences in personnel and procedures. Centralized and automated sample handling and processing helps to assure standardization and to minimize sample loss and mix-up, greatly contributing to high standards of quality. At the biobank, sample aliquots are kept at -80 °C in a fully automated sample store. Two vials of EDTA whole blood for extraction of DNA are obtained from each patient and shipped directly to the biobank at ambient temperature, then stored at -80 °C until processing.
Molecular assessments
Broad molecular analyses are an important focus of PROGRESS. All study participants will be genotyped using the CAP2 array, a customized array based on the Axiom platform (Affymetrix). The CAP2 array comprises the standard genome-wide content of the Axiom CEU array and about 60,000 custom SNPs. This custom content consists of SNPs selected from existing literature evidence, eQTL data bases, GWAS data bases, and data bases of functionally relevant mutations. Genetic data from the CAP2 array are suitable for imputation of genotype data with common genomic references. Genome-wide gene-expression measurements using the HumanHT-12v4 Expression BeadChip (Illumina) comprise 47.000 transcript features in RNA from stabilized whole blood (PAXgene Blood RNA System, PreAnalytiX). Proteome analyses involve gel-free LC-MS/MS focused measurements in EDTA plasma samples stabilized by protease inhibitors (BD P100 Blood Collection System for plasma protein preservation, Becton Dickinson) and depleted for highly abundant proteins. Collected biospecimen also allow measurements of other serum and plasma parameters as well as cytokines. As patient samples are available from consecutive study time points, time series data can be established which are amenable to temporal analysis of molecular response to infection.
Discussion
PROGRESS collects clinical and molecular data for investigating the role of host factors modulating innate immunity in patient’s response to community acquired pneumonia. Most patients hospitalized with CAP recover rather quickly while others will experience a severe course of disease, with some requiring ICU treatment including respiratory support. PROGRESS is designed to investigate host factors beyond old age and comorbidity which modulate the course of disease. Therefore, PROGRESS was designed as a unique, detailed, systematic, longitudinal study of CAP in Germany and Austria focusing on the progression of hospitalized community acquired pneumonia from mild CAP to very severe disease including severe sepsis and septic shock. Patients are closely followed for several days regarding clinical and laboratory parameters and several biospecimen are collected on these days. Inspiration for the design of PROGRESS came from CAPNETZ and its study on epidemiological and health research aspects of CAP in Germany [34, 36]. Main differences of PROGRESS compared to CAPNETZ are very stringent exclusion criteria excluding severe comorbidities, a specific focus on hospitalized CAP, more comprehensive documentation on disease course within the first five days in the hospital, and a stronger focus on molecular assessments. Another inspiration for the PROGRESS study was the “study on Genetic and Inflammatory Markers of Sepsis” (GenIMS) in CAP in the US [17]. However, criteria in PROGRESS are much stricter regarding exclusion of potentially immunocompromised patients and options for molecular assessments are broader. Inspiration regarding the assessment of severely ill CAP patients came from several SepNet [23] studies in Germany which, like the European GenOSept study [13, 26], focus on patients with sepsis of diverse origins, including pneumonia.
Stringent inclusion criteria ensure that only patients with community acquired pneumonia are enrolled. A new infiltrate in chest X-ray along with at least two other clinical symptoms of pneumonia are required. Patients identified at later time points, or suffering from tuberculosis or neoplasia of the lung, for instance, are excluded from the study. To further ensure disease homogeneity, patients who may have acquired pneumonia in medical care facilities (hospital or health care acquired pneumonia, HAP) or under respiratory support (ventilator acquired pneumonia, VAP) are excluded. Numerous exclusion criteria apply in an attempt to avoid patients whose immune systems may be substantially compromised by diseases with direct impact on the immune system, other severe diseases likely to modulate the course of pneumonia, or immunosuppressive treatments. Along these lines, patients for whom a therapy limitation is assumed are excluded. Decisions on limitation of therapy that are made after enrolment are documented. Hence, patients enrolled in PROGRESS are not representative of the overall population of hospitalized CAP patients. Rather, PROGRESS patients will likely be younger and generally healthier except for their acute pneumonia. While our approach aims to get closer to unconfounded biological heterogeneity regarding CAP progression, this also requires that insights from PROGRESS have to be validated in more representative patient cohorts before generalizations regarding all CAP patients can be drawn.
By definition, it is impossible to observe patients hospitalized with CAP without the influence of the hospital environment. To come as close as possible to an analysis of pure CAP, we require enrolment of patients within 48 h of hospitalization and are limiting detailed observation of patients to the first five to six days after arrival in the hospital. At later time points, it cannot be ruled out that hospital acquired infection may superimpose on the initial CAP. However, major events and substantial changes in patients’ health state are recorded until hospital discharge to gain insight into the overall course of disease in the hospital. Patients are followed up by telephone interview at days 28, 180, and 360 after enrolment for vital status, recurrence of pneumonia, housing situation, and quality of life. Thus, questions regarding long term consequences of CAP can also be addressed.
In the PROGRESS study, microbiological data are documented as they become available from routine clinical materials, especially if materials were obtained between hospital admission and day 4 of the study. This includes culture from airway materials and blood as a well as influenza rapid test and urine antigen tests for legionella viruses and pneumococci. Results from molecular techniques such as PCR are also documented. Microbiological assessment will likely be available for only a part of PROGRESS patients and disease causing agents will be detected for only a fraction of patients. We hope to supplement this information on disease causing agents in the long run by molecular analyses of available biospecimen. These analyses can be validated with documented microbiological data.
The main goal of PROGRESS is to contribute to the improvement of clinical decision making and management of patients with pneumonia and sepsis and of therapeutic strategies. This will be achieved by clinical and molecular investigation of host factors modulating outcome of CAP and by identifying clinical scores and molecular signatures for diagnosis, risk stratification, prognosis, and outcome prediction.
Abbreviations
AIDS, acquired immunodeficiency syndrome; AQUA, Institute for Applied Quality Improvement and Research in Health Care, Germany; CAP, community acquired pneumonia; DNA, deoxyribonucleic acid; eCRF, electronic case report form; EDTA, ethylenediaminetetraacetic acid; eQTL, expression quantitative trait locus; GenIMS, genetic and inflammatory markers of sepsis study; GWAS, Genome Wide Association Study; HAP, Hospital or Health care Acquired Pneumonia; HIV, human immunodeficiency virus; IBBJ, integrated biobank jena; ICU, intensive care unit; LC-MS/MS, liquid chromatography coupled tandem mass spectroscopy; PCR, polymerase chain reaction; PROGRESS, Pneumonia Research Network on Genetic Resistance and Susceptibility for the Evolution of Severe Sepsis; RNA, ribonucleic acid; sCAP, severe CAP; SepNet, Study Group Competency Network Sepsis; ssCAP, severe CAP with pneumogenic sepsis and septic shock; uCAP, uncomplicated CAP; VAP, ventilator acquired pneumonia; WHO, World Health Organization
References
Angus DC, Marrie TJ, Obrosky DS, Clermont G, Dremsizov TT, Coley C, et al. Severe community-acquired pneumonia: use of intensive care services and evaluation of American and British Thoracic Society Diagnostic criteria. Am J Respir Crit Care Med. 2002;166(5):717–23. doi:10.1164/rccm.2102084.
AQUA-Institut. Bundesauswertung zum Erfassungsjahr 2014. PNEU – Ambulant erworbene Pneumonie. Qualitätsindikatoren. 2015.
Brown JS. Community-acquired pneumonia. Clin Med (London, England). 2012;12(6):538–43.
Brunkhorst FM; Engel C, Bone HG, Brunkhorst R, Gerlach H, Grond S et al. Epidemiology of severe sepsis and septic shock in Germany - Results from the German Prevalence Study. Infection. 2005;33(Suppl. 1):49.
Buhimschi CS, Bhandari V, Hamar BD, Bahtiyar MO, Zhao G, Sfakianaki AK, et al. Proteomic profiling of the amniotic fluid to detect inflammation, infection, and neonatal sepsis. PLoS Med. 2007;4(1):e18. doi:10.1371/journal.pmed.0040018.
Dremsizov T, Clermont G, Kellum JA, Kalassian KG, Fine MJ, Angus DC. Severe sepsis in community-acquired pneumonia: when does it happen, and do systemic inflammatory response syndrome criteria help predict course? Chest. 2006;129(4):968–78. doi:10.1378/chest.129.4.968.
Engel C, Brunkhorst FM, Bone HG, Brunkhorst R, Gerlach H, Grond S, et al. Epidemiology of sepsis in Germany: results from a national prospective multicenter study. Intensive Care Med. 2007;33(4):606–18. doi:10.1007/s00134-006-0517-7.
European Medicines Agency (EMEA). ICH Topic E6 (R1) Guideline for Good Clinical Practice. NOTE FOR GUIDANCE ON GOOD CLINICAL PRACTICE CPMP/ICH/135/95 2006. 2002. Online available at http://www.ema.europa.eu/docs/en_GB/document_library/Scientific_guideline/2009/09/WC500002874.pdf, last accessed January 2016.
Ewig S, Birkner N, Strauss R, Schaefer E, Pauletzki J, Bischoff H, et al. New perspectives on community-acquired pneumonia in 388 406 patients. Results from a nationwide mandatory performance measurement programme in healthcare quality. Thorax. 2009;64(12):1062–9. doi:10.1136/thx.2008.109785.
Feldman C. Prognostic scoring systems: which one is best? Curr Opin Infect Dis. 2007;20(2):165–9. doi:10.1097/QCO.0b013e328014da75.
Fine MJ, Hanusa BH, Lave JR, Singer DE, Stone RA, Weissfeld LA, et al. Comparison of a disease-specific and a generic severity of illness measure for patients with community-acquired pneumonia. J Gen Intern Med. 1995;10(7):359–68.
Froy O. Regulation of mammalian defensin expression by Toll-like receptor-dependent and independent signalling pathways. Cell Microbiol. 2005;7(10):1387–97. doi:10.1111/j.1462-5822.2005.00590.x.
Gordon AC, Lagan AL, Aganna E, Cheung L, Peters CJ, McDermott MF, et al. TNF and TNFR polymorphisms in severe sepsis and septic shock: a prospective multicentre study. Genes Immun. 2004;5(8):631–40. doi:10.1038/sj.gene.6364136.
Hartlova A, Krocova Z, Cerveny L, Stulik J. A proteomic view of the host-pathogen interaction: The host perspective. Proteomics. 2011;11(15):3212–20. doi:10.1002/pmic.201000767.
Hurst NP, Kind P, Ruta D, Hunter M, Stubbings A. Measuring health-related quality of life in rheumatoid arthritis: validity, responsiveness and reliability of EuroQol (EQ-5D). Br J Rheumatol. 1997;36(5):551–9.
Johnson SB, Lissauer M, Bochicchio GV, Moore R, Cross AS, Scalea TM. Gene expression profiles differentiate between sterile SIRS and early sepsis. Ann Surg. 2007;245(4):611–21. doi:10.1097/01.sla.0000251619.10648.32.
Kellum JA, Kong L, Fink MP, Weissfeld LA, Yealy DM, Pinsky MR, et al. Understanding the inflammatory cytokine response in pneumonia and sepsis: results of the Genetic and Inflammatory Markers of Sepsis (GenIMS) Study. Arch Intern Med. 2007;167(15):1655–63. doi:10.1001/archinte.167.15.1655.
Kolditz M, Ewig S, Höffken G. Management-based risk prediction in community-acquired pneumonia by scores and biomarkers. Eur Respir J. 2013;41(4):974–84. doi:10.1183/09031936.00104412.
Lim WS, Baudouin SV, George RC, Hill AT, Jamieson C, Le Jeune I, et al. BTS guidelines for the management of community acquired pneumonia in adults: update 2009. Thorax. 2009;64 Suppl 3:iii1–55. doi:10.1136/thx.2009.121434.
Lim WS, van der Eerden MM, Laing R, Boersma WG, Karalus N, Town GI, et al. Defining community acquired pneumonia severity on presentation to hospital: an international derivation and validation study. Thorax. 2003;58(5):377–82.
Marti C, Garin N, Grosgurin O, Poncet A, Combescure C, Carballo S, et al. Prediction of severe community-acquired pneumonia: a systematic review and meta-analysis. Crit Care (London, England). 2012;16(4):R141. doi:10.1186/cc11447.
Müller-Redetzky HC, Suttorp N, Witzenrath M. Dynamics of pulmonary endothelial barrier function in acute inflammation: mechanisms and therapeutic perspectives. Cell Tissue Res. 2014;355(3):657–73. doi:10.1007/s00441-014-1821-0.
Oppert M, Engel C, Brunkhorst FM, Bogatsch H, Reinhart K, Frei U, et al. Acute renal failure in patients with severe sepsis and septic shock--a significant independent risk factor for mortality: results from the German Prevalence Study. Nephrol Dial Transplant. 2008;23(3):904–9. doi:10.1093/ndt/gfm610.
Pereira JM, Paiva JA, Rello J. Assessing severity of patients with community-acquired pneumonia. Semin Respir Crit Care Med. 2012;33(3):272–83. doi:10.1055/s-0032-1315639.
Ramilo O, Allman W, Chung W, Mejias A, Ardura M, Glaser C, et al. Gene expression patterns in blood leukocytes discriminate patients with acute infections. Blood. 2007;109(5):2066–77.
Rautanen A, Mills TC, Gordon AC, Hutton P, Steffens M, Nuamah R, et al. Genome-wide association study of survival from sepsis due to pneumonia: an observational cohort study. Lancet Respir Med. 2015;3(1):53–60. doi:10.1016/S2213-2600(14)70290-5.
Reinhart K, Brunkhorst FM, Bloos F, Bone HG; Gerlach H, Gründling M et al. Diagnose und Therapie der Sepsis. S2 Leitlinien der Deutschen Sepsis-Gesellschaft e.V. und der Deutschen Interdisziplinären Vereinigung für Intensiv- und Notfallmedizin. 2010. Online available at http://www.awmf.org/uploads/tx_szleitlinien/079-001l_S2k_Sepsis_2010-abgelaufen.pdf, last accessed February 2016.
Scicluna BP, Klouwenberg K, Peter MC, van Vught LA, Wiewel MA, Ong DSY, et al. A molecular biomarker to diagnose community-acquired pneumonia on intensive care unit admission. Am J Respir Crit Care Med. 2015;192(7):826–35. doi:10.1164/rccm.201502-0355OC.
Serkova NJ, Standiford TJ, Stringer KA. The emerging field of quantitative blood metabolomics for biomarker discovery in critical illnesses. Am J Respir Crit Care Med. 2011;184(6):647–55. doi:10.1164/rccm.201103-0474CI.
Sørensen TI, Nielsen GG, Andersen PK, Teasdale TW. Genetic and environmental influences on premature death in adult adoptees. N Engl J Med. 1988;318(12):727–32. doi:10.1056/NEJM198803243181202.
Steel HC, Cockeran R, Anderson R, Feldman C. Overview of community-acquired pneumonia and the role of inflammatory mechanisms in the immunopathogenesis of severe pneumococcal disease. Mediat Inflamm. 2013;2013:490346. doi:10.1155/2013/490346.
Tang BMP, McLean AS, Dawes IW, Huang SJ, Lin RCY. Gene-expression profiling of peripheral blood mononuclear cells in sepsis. Crit Care Med. 2009;37(3):882–8. doi:10.1097/CCM.0b013e31819b52fd.
Vincent JL, Moreno R, Takala J, Willatts S, de Mendonça A, Bruining H, et al. The SOFA (Sepsis-related Organ Failure Assessment) score to describe organ dysfunction/failure. On behalf of the Working Group on Sepsis-Related Problems of the European Society of Intensive Care Medicine. Intensive Care Med. 1996;22(7):707–10.
Welte T, Suttorp N, Marre R. CAPNETZ-community-acquired pneumonia competence network. Infection. 2004;32(4):234–8. doi:10.1007/s15010-004-3107-z.
Welte T, Torres A, Nathwani D. Clinical and economic burden of community-acquired pneumonia among adults in Europe. Thorax. 2012;67(1):71–9. doi:10.1136/thx.2009.129502.
Welte T, Köhnlein T. Global and local epidemiology of community-acquired pneumonia: the experience of the CAPNETZ Network. Semin Respir Crit Care Med. 2009;30(2):127–35. doi:10.1055/s-0029-1202941.
WHO. WHO report on global burden of disease 2000-2012, update 2014. Health statistics and information systems Global Health Estimates (GHE). 2016. Online available at http://www.who.int/healthinfo/global_burden_disease/en/, last accessed February 2016.
Wong HR, Cvijanovich N, Allen GL, Lin R, Anas N, Meyer K, et al. Genomic expression profiling across the pediatric systemic inflammatory response syndrome, sepsis, and septic shock spectrum. Crit Care Med. 2009;37(5):1558–66. doi:10.1097/CCM.0b013e31819fcc08.
World Medical Association Declaration of Helsinki. Recommendations guiding physicians in biomedical research involving human subjects. Cardiovascular research. 1997;35(1):2–3.
Acknowledgements
We thank all patients for their participation in the PROGRESS study and their altruistic contribution to science. We also thank all physicians, study nurses, scientists, and all others without whom this study would not be possible.
In particular, we thank Marlies Herold for her continuous effort in providing support and help for all study centers to ensure high quality of data and biospecimen, Kerstin Wieland for programming and maintaining the remote data entry tools and the central PROGRESS database, Katrin Ludewig, Kay Stoetzer, and Julia Koehler for providing consistent support with study materials and biobanking.
Responsibilities in the PROGRESS consortium are: NS principle investigator, NS, PC, PA, and HS (formerly) clinical study coordinator, ML, MS, PA Biometry and data bases, MK biobanking, TC and HH microbiology, UV proteomics, MS and HH genomics, and MB and MS transcriptomics. Conception of the study was further supported by Frank Martin Brunkhorst, Tobias Welte, Katrin Ludewig, and Lothar Jänsch.
Members of the PROGRESS study group 2015
Andreas Liebrich: St. Vincenz und Elisabeth Hospital Mainz, Innere Medizin, Mainz, Germany. Ingrid Maria Stelzmüller: Allgemeines Krankenhaus der Stadt Linz, Abteilung Lungenheilkunde, Linz, Austria. Barbara Khanavkar, Aida Avdic: Altmark-Klinikum, Krankenhaus Salzwedel, Salzwedel, Germany. Deborah Wehde, Barabara Hauptmeier: Berufsgenossenschaftl. Universitätsklinikum Bergmannsheil, Klinik f. Pneumologie Allergologie u. Schlafmedizin, Bochum, Germany. Katharina Edle v. Schmädel, Florian Steiner, Yeji Kim, Lea Drobbe, Michaela Niebank, Katharina Dobrovits: Charité - Universitätsmedizin Berlin, Medizinische Klinik m. S. Infektiologie und Pneumologie, Berlin, Germany. Hermann Druckmiller, Galina Schott: Christliches Krankenhaus Quakenbrück, Med. Klinik Abtl. Pneumologie Allergologie Schlafmedizin, Quakenbrück, Germany. Tom Schaberg, Iris Hering: Diakoniekrankenhaus Rotenburg (Wümme), Zentrum für Pneumologie, Rotenburg (Wümme), Germany. Klaus Thomas, Codruta Ursu, Florijan Ameti: Evangelisches Fachkrankenhaus für Atemwegserkrankungen, Innere Medizin, Neustadt/Südharz, Germany. Martin Buchenroth, Oliver Kanwar, Bernhard Schöffend: Evangelische Kliniken Bonn - Betriebsstätte Johanniter Krankenhaus, Innere Medizin II, Bonn, Germany. Lena Kappauf, Constanze Steiner: Evangelisches Krankenhaus Kalk, Innere Medizin/Pneumologie, Köln, Germany. Andreas Bundkirchen: Franziskus Krankenhaus Linz, Innere Abteilung, Linz, Germany. Christian Grah, Adrian Steinmetz: Gemeinschaftskrankenhaus Havelhöhe, Kardio-Pneumologie, Berlin, Germany. Christian Wrede, Wolfgang Stahl: HELIOS Klinikum Berlin-Buch, Interdisziplinäres Notfallzentrum mit Rettungsstelle, Berlin, Germany. Antonia Wohlers, Gabriele Wöbker: HELIOS Klinikum Wuppertal, Klinik für Intensivmedizin, Wuppertal, Germany. Markus Leidag, Peter Kaufmann, Kurt Rasche: HELIOS Klinikum Wuppertal, Pneumologie - Bergisches Lungenzentrum, Wuppertal, Germany. Andre Althoff, Sabine Schröder-Hergt: Hochtaunus-Kliniken, Medizinische Klinik III, Bad Homburg, Germany. Carola Hobler, Simone Hamberger, Thomas Müller: Kliniken d. Main-Taunus-Kreises, Klinik f. Pneumologie u. Allg. Innere Medizin, Hofheim, Germany. Bernhard Schaaf, Markus Unnewehr, Ann-Kathrin Silber: Klinikum Dortmund, Medizinische Klinik Nord Pneumologie Infektiologie und internistische Intensivmedizin, Dortmund, Germany. Bernhard Ruf, Tobias Marcello: Klinikum St. Georg, Klinik für Infektiologie/Tropenmedizin Nephrologie und Rheumatologie, Leipzig, Germany. Brigitte Mayer: Klinkum Heidenheim, Medizinische Klinik II & Innere Intensiv, Heidenheim, Germany. Jan Pluta, Christoph Arntzen: Krankenhaus Angermünde, Klinik f. Innere Medizin/Pneumologie, Angermünde, Germany. Walter Knüppel: Krankenhaus Bad Arolsen, Innere Medizin - Herz- Kreislauf- u. Lungendiagnostik, Bad Arolsen, Germany. Christina Priegnitz: Krankenhaus Bethanien, Abt. Pneumologie, Solingen, Germany. Thomas Spinner, Verena Sautermeister, Christine Krollmann, Malte Müller: Krankenhaus München-Neuperlach, Klinik für Kardiologie Pneumologie und Internistische Intensivmedizin, München, Germany. Hubert Pilgram, Thomas Glück: Kreisklinik Trostberg, Innere Abteilung, Trostberg, Germany. Holger Flick, Michael Palfner, Martin Hönigl, Gudrun Wakonigg: LKH-Univ. Klinikum Graz, UKIM Pulmologie, Graz, Austria. Karsten Hartung, Barbara Wagener: Lungenklinik Ballenstedt/Harz, Fachkrankenhaus f. Lungenkrankheiten, Ballenstedt, Germany. Sebastian Bubel, Sebastian Nuding: Universitätsklinikum Halle (Saale), Klinik für Innere Medizin III - Kardiologie Pneumologie Intensivmedizin Angiologie, Halle (Saale), Germany. Ayhan Berber, Petra Hoffknecht, Nicolas Dickgreber: Mathias-Spital Rheine, Klinik für Pneumologie und Thoraxonkologie, Rheine, Germany. 10.1186/s12890-016-0255-8 Jessica Rademacher, Medizinische Hochschule Hannover, Klinik für Pneumologie, Hannover, Germany. Barbara Schröder, Berthold Jany: Missionsärztliche Klinik, Innere Medizin, Würzburg, Helmut Wunderlich, Susanne Lang, Johannes Ortmann: SRH Wald-Klinikum Gera, Medizinische Klinik 2, Gera, Germany. Michael Simpfendörfer: St.Vincentius-Kliniken, Med. Klinik IV/Pneumologie, Karlsruhe, Germany. Marianne Schelle, Mathias Plauth, Judith Pannier: Städt. Klinikum Dessau, Innere Medizin, Dessau-Roßlau, Germany. Claus Peckelsen, Patrick Schmidt-Schridde: Städtisches Klinikum München - Klinikum Harlaching, Klinik für Gastroent. Pneum. intern. Akut- u. Intensivmedizin, München, Germany. Alexander Heine: Uniklinikum Greifswald, Zentrum für Innere Medizin B, Greifswald, Germany. Lea Deterding, Hubert Wirtz, Eva Koch: Universitätsklinikum Leipzig, Innere Medizin Neurologie Dermatologie und Pneumologie, Leipzig, Germany. Martin Kolditz: Universitätsklinikum Carl Gustav Carus, Medizinische Klinik 1 - Abteilung Pneumologie, Dresden, Germany. Christian Lensch, Robert Bals: Universitätsklinikum des Saarlandes, Innere Medizin V, Homburg/Saar, Germany. Mathias Pletz, Janina Frosinski, Anne Moeser, John Paul Fobiwe: Universitätsklinikum Jena, Zentrum für Infektionsmedizin und Krankenhaushygiene, Jena, Germany. Katrin Ludewig: Universitätsklinikum Jena, Klinik für Anästhesiologie und Intensivtherapie, Jena, Germany. Klaus Dalhoff, Jan Rupp: Universitätsklinikum Schleswig-Holstein - Campus Lübeck, Med. Klinik III Pneumologie, Lübeck, Germany. Cornelia Kropf-Sanchen, Stefan Rüdiger: Universitätsklinikum Ulm, Klinik für Innere Medizin II - Sektion Pneumologie, Ulm, Germany. Ursula Hoffmann: Universitätsmedizin Mannheim, 1. Medizinische Klinik, Mannheim, Germany. Josefa Lehmke: Humboldt-Klinikum Vivantes, Kardiologie und kons. Intensivmedizin, Berlin, Germany. Henning Kahnert: Vivantes Klinikum Spandau, Kard. Pneum. und kons. Intensivmedizin, Berlin, Wulf Pankow, Dorina Thiemig, Achim Lies: Vivantes Klinikum Neukölln, Klinik f. Innere Med. - Pneumologie u. Infektiologie – Thoraxzentrum, Berlin, Germany.
Funding
The PROGRESS study is supported by the German Federal Ministry of Education and Research, grant numbers 01KI07110 (Giessen), 01KI07111 (Jena), 01KI07113 (Leipzig), 01KI07114 (Berlin), 01KI1010I (Leipzig), and 01KI1010D (Greifswald). The German Federal Ministry of Education and Research has no influence on the design of the study, on collection, analysis, and interpretation of data, or on writing the manuscript.
Availability of data and material
Data and biomaterials from the PROGRESS study will be available once the study has been finished. Applications for data require a formal application and will be decided upon by the board of the PROGRESS study group.
Authors’ contributions
All authors have made substantial contributions to the conception and design of the PROGRESS study, were actively involved in implementing it, and support the ongoing effort: The study was designed by NS, ML, MS, CE, PA, MB, MK, HH, TC, SH, and HS. The manuscript was drafted by PA, PC, and MS and was critically revised by ML and NS. All authors read and approved the final manuscript.
Competing interests
The authors declare that they have no competing interests.
Consent for publication
Not applicable.
Ethics approval and consent to participate
Written informed consent for participation in the study was obtained from all participants or, in case of critical illness, from patient’s legal representatives. The study protocol has been approved by the ethics committee of the University of Jena (registration number 2403-10/08) and all locally responsible ethics committees of all study centers (see Table 1).
Author information
Authors and Affiliations
Consortia
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Ahnert, P., Creutz, P., Scholz, M. et al. PROGRESS – prospective observational study on hospitalized community acquired pneumonia. BMC Pulm Med 16, 108 (2016). https://doi.org/10.1186/s12890-016-0255-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12890-016-0255-8