Abstract
Background
Enhancer RNAs (eRNAs) are demonstrated to be closely associated with tumourigenesis and cancer progression. However, the role of eRNAs in lung adenocarcinoma (LUAD) remains largely unclear. Thus, a comprehensive analysis was constructed to identify the key eRNAs, and to explore the clinical utility of the identified eRNAs in LUAD.
Methods
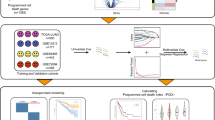
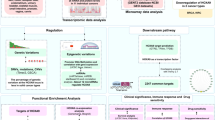
First, LUAD expression profile data from the Cancer Genome Atlas (TCGA) dataset and eRNA-relevant information were integrated for Kaplan-Meier survival analysis and Spearman’s correlation analysis to filtered the key candidate eRNAs that was associated with survival rate and their target genes in LUAD. Then, the key eRNA was selected for subsequent clinical correlation analysis. KEGG pathway enrichment analyses were undertaken to explore the potential signaling pathways of the key eRNA. Data from the human protein atlas (HPA) database were used to validate the outcomes and the quantitative real time-polymerase chain reaction (qRT-PCR) analysis was conducted to measure eRNA expression levels in tumor tissues and paired normal adjacent tissues from LUAD patients. Finally, the eRNAs were validated in pan-cancer.
Results
As a result, TBX5-AS1 was identified as the key eRNA, which has T-box transcription factor 5 (TBX5) as its regulatory target. KEGG analysis indicated that TBX5-AS1 may exert a vital role via the PI3K/AKT pathway, Ras signaling pathway, etc. Additionally, the qRT-PCR results and the HPA database indicated that TBX5-AS1 and TBX5 were significantly downregulated in tumour samples compared to matched-adjacent pairs. The pan-cancer validation results showed that TBX5-AS1 was associated with survival in four tumors, namely, adrenocortical carcinoma (ACC), LUAD, lung squamous cell carcinoma (LUSC), and uterine corpus endometrial carcinoma (UCEC). Correlations were found between TBX5-AS1 and its target gene, TBX5, in 26 tumor types.
Conclusion
Collectively, our results indicated that TBX5-AS1 may be a potential prognostic biomarker for LUAD patients and promote the targeted therapy of LUAD.
Similar content being viewed by others
Background
Lung cancer is recognized as one of the most common malignant carcinomas, and ranks first for morbidity and mortality among all malignant tumors in the world. Non-small cell lung cancer (NSCLC) is the most frequently diagnosed type of lung cancer, and comprises two major types—adenocarcinoma and squamous carcinoma [1, 2]. Because the early symptoms of lung cancer aren’t clear and are confused easily with those of other respiratory diseases, the rates of misdiagnosis and missed diagnosis are high, and the majority of patients are diagnosed at an intermediate or late stage, at which the effective treatment options are limited and the prognosis is usually poor. In recent years, with the development of gene detection technology and targeted therapies, great strides have been made in the treatment and prognosis of LUAD. However, the 5-year overall survival is still less than 20% [3, 4], highlighting the urgent need to identify effective prognosis-related biomarkers for LUAD; evaluating the prognosis of LUAD patients through biomarkers in the early stage of the disease is of great importance to improving the prognosis and survival rate of these patients.
Over the past decade, the rapid application and development of high-throughput sequencing technology has opened a new window for noncoding RNAs, and accumulating evidence has shown that the occurrence and development of a series of major diseases are related to noncoding RNA dysregulation [5,6,7]. Among these, RNAs generated from enhancers (enhancer RNAs; eRNAs) have gained substantial interest owing to their underlying roles in the mediation of gene transcription and enhancer functions, and their frequent overlap with noncoding risk loci associated with diseases [8,9,10,11]. Enhancers are key cis-regulatory elements that promote the expression of eukaryotic genes [12, 13]. eRNAs are 500–5000 bp-long RNAs produced through enhancer transcription [14]. They can activate enhancer activity independently, and can also combine with other protein factors to promote the formation of the enhancer-promoter loop, thereby activating the expression of downstream genes [15,16,17]. eRNA dysregulation can affect biological processes such as the cell cycle and the growth of cancer cells, or change the expression of target genes, suggesting that eRNAs may serve as novel targets for tumour treatment [15, 18, 19]. A recent study found that, mutations in the eRNA element of ACTRT1 in basal cell carcinoma can weaken the enhancer’s activity as well as the expression of ACTRT1, resulting in the aberrant activation of Hedgehog signaling and, consequently, the promotion of carcinogenesis and tumour development [20]. In T-cell acute lymphoblastic leukemia (T-ALL), it was also reported that the eRNA ARIEL could recruit mediator proteins to the ARID5B enhancer, thereby boosting enhancer-promoter interactions and activating the expression of ARID5B. These activities led to the aberrant activation of the TAL1-induced transcriptional program and myc oncogenic signals [21]. NET1 is an oncogene of key importance in breast cancer. Related research has demonstrated that the NET1-associated eRNA (NET1e) could upregulate the expression of its target gene and promote breast cancer progression. In addition, overexpression of NET1e in situ can lead to the resistance of breast cancer cells to BEZ235 and octreotide [22]. Additionally, a separate research discovered an eRNA, AP001056.1, is able to serve to be a prognostic marker for head and neck squamous cell carcinoma (HNSCC) [23]. However, to date, no study has investigated the potential of using eRNAs as biomarkers to predict the survival of patients with LUAD.
Herein, we aimed at identifying prognostic eRNAs as well as their target genes within LUAD for predicting LUAD patients’ prognosis. Our results may facilitate the understanding of the potential value of the clinical application of LUAD-associated eRNAs in prognosis stratification and promote the targeted therapy of LUAD.
Materials and methods
Identification of predictive eRNAs in LUAD by comprehensive data analysis
Clinical data, RNA expression profile data, and survival information for 33 cancer types were downloaded from the TCGA database. All RNA expression profile data were transformed to log2 (FPKM+ 1). Ensemble transcript IDs were con-verted to their corresponding GENCODE using GTF annotation files for humans. We searched and comprehensively analysed the eRNA source literature to obtain the relevant eRNA information. The human GTF files were used to transform eRNA transcript IDs into gene symbols and the expression profiles of the eRNAs were extracted from the RNA expression profile of lung adenocarcinoma. Then, we combined the eRNA expression matrix with the lung adenocarcinoma survival data using the limma R software package.
The survival-associated eRNAs were screened using Kaplan-Meier methodology, taking the false discovery rate (FDR) adjusted P < 0.05 as standard cut-off values, we selected eRNAs that met this condition as survival-associated eRNAs. The patients were classified into low- and high-expression groups according to the median expression of each eRNA. Differences in survival were compared between the two groups using p < 0.05 as the cut-off and a survival curve for the eRNAs that met the conditions was drawn. Then, Spearman’s correlation analysis was performed to obtain candidate key eRNAs related to survival and associated with target genes with roles in LUAD; statistical significance was considered at cor > 0.4 and p < 0.001.
Gene enrichment analysis
Besides the predicted targets, Pearson’s correlation analysis was performed to identify other genes in LUAD that were significantly associated with the key eRNA (cor > 0.4, and p < 0.001). The target genes of the key eRNA were converted into their corresponding gene IDs for further analysis using the org. Hs.eg.db R software package. To investigate which functions and pathways were affected by the eRNA in tumorigenesis and metastasis, the clusterProfiler package in R was used for the pathway enrichment analyses of Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) www.kegg.jp/kegg/kegg1.html) [24,25,26] of the key eRNA target genes. Enriched GO terms with an adjusted p-value < 0.001 were considered as relevant biological processes.
Clinical LUAD sample collection
A total of 10 LUAD tumor tissues and para-tumor samples were collected from patients diagnosed with LUAD according to pathological analysis and who were undergoing surgery at the Second Xiangya Hospital of Central South University. After tumor resection, the tumor and adjacent tissues were immediately frozen in liquid nitrogen and stored at − 80°Cuntil total RNA extraction. All patients gave their informed consent for inclusion before surgery. The study was conducted in accordance with the Declaration of Helsinki, and the protocol was approved by the Ethics Committee of the Second Xiangya Hospital of Central South University (protocol code 2020038 and 07/01/2020).
Quantitative real-time polymerase chain reaction (qRT-PCR) validation
TRIzol reagent was used to extract total RNA from tissue samples (Invitrogen, GrandIsland, NY, USA). The quality and quantity of RNA were estimated by NanoDropND-1000 (Thermo Scientific, Waltham, MA, USA), while the integrity of RNA were assessed by standard denaturing agarose gel electrophoresis. Then, the RNA was reverse-transcribed into cDNA by adopting SuperScript III Reverse Transcriptase (Invitrogen) in accordance with the instructions of the manufacturer. A 2× PCR Master Mix (Arraystar) and an Applied Biosystems ViiA 7 Real-Time PCR System were adopted for qRT-PCR following the instructions of the manufacturer. The relative eRNA and mRNA expression levels were calculated using the 2 − ΔΔCt method. The primers used and the sequences are listed in Table 1 and Table 2. The data represent the means of three experiments.
Human protein atlas analysis
The human protein atlas (HPA; https://www.Proteinatlas.org/) comprises an atlas of human protein expression patterns in normal and tumor tissues. In this study, we assessed the expression of T-box transcription factor 5 (TBX5) in the protein expression module of the HPA database and analyzed the immunohistochemical staining results for TBX5 in both tumor and normal tissues (antibody: HPA008786).
Verification in pan-Cancer
First, the expression data of TBX5-AS1 and its target gene TBX5 in pan-cancer were obtained using the R limma package, and then the expression matrix was combined with the pan-cancer survival data. The samples were classified into low- and high-expression groups according to the median value of the expression of TBX5-AS1, and then the Kaplan–Meier method was used to compare the survival difference between the two groups. p < 0.05 was considered statistically significant. A survival curve was plotted for TBX5-AS1 in tumours that met the criteria. Spearman’s coefficient was used to test the cor-relation between TBX5-AS1 and its corresponding target genes TBX5 in pan-cancer. A correlation coefficient > 0.4 and a p-value < 0.001 were considered statistically significant.
Results
Screening the key eRNA in LUAD
Using the Kaplan-Meier method, eRNAs with P-values adjusted according to the FDR cut-off of < 0.05 were extracted as survival-related eRNAs in LUAD. A total of 174 eRNAs were identified (Additional file 1: Table S1). Spearman’s correlation was then used to screen these 174 eRNAs to identify those with a significant correlation with their target genes that were associated with LUAD. Only 76 eRNAs met the condition (Spearman’s rank correlation coefficient r > 0.40, p < 0.001; Additional file 1: Table S2). Of these, TBX5-AS1 exhibited the highest cor value, and was therefore considered to be the most relevant eRNA associated with its target gene. Kaplan–Meier curve results for TBX5-AS1 showed that patients in the low-expression group survived longer than those in the high-expression group (p = 0.02998; Fig. 1a). In addition, there was a positive correlation between TBX5-AS1 and its target gene TBX5 (r = 0.89973, p = 2.2 × 10–15; Fig. 1b). We further investigated the connections between the clinical features of LUAD patients and TBX5-AS1 expression and found that the expression level of TBX5-AS1 in LUAD was significantly linked to T stage (T1 vs. T2, p = 0.00011), N stage (N0 vs. N2, p = 0.05), sex (p = 0.0061), and cancer status (p = 0.018) (Fig. 2a–d); however, other clinical characteristics, such as age and M stage, had no clear correlation with TBX5-AS1 expression (p > 0.05) (Fig. 2e, f).
Gene enrichment analysis
A total of 1407 transcripts presented a significant correlation with TBX5-AS1 (p < 0.05), including TBX5. GO enrichment analysis and KEGG pathway analysis of the 1407 target genes provided the basis for the biological study of these genes. The top 10 terms for molecular function (MF), biological process (BP) and cellular component (CC) are summarized in Fig. 3a. In BP, the terms were mainly related to extracellular matrix organization and extracellular structure organization; in CC, they were related to extracellular matrix, collagen-containing extracellular matrix, postsynaptic membrane, and adherens junction, among others. In MF, term enrichment mainly involved extracellular matrix structural constituent and glycosaminoglycan binding. The intensity of the colors indicates the p-value (stronger colors indicate lower p-values)—the lower the p-value, the more significant the GO term. The top 30 pathways are summarized in Fig. 3b. KEGG enrichment analysis showed that the most significantly enriched biological process was “PI3K/AKT signaling pathway” (adjusted p < 0.001). The PI3K/AKT signaling pathway-related genes with a Spearman’s correlation coefficient > 0.6 are depicted in Table 3.
Data validation
qRT-PCR was used to measure the expression levels of TBX5-AS1 and TBX5 in 10 LUAD samples and paired adjacent samples. The outcomes indicated that the expression of TBX5-AS1 and TBX5 in LUAD tissues was significantly down-regulated when comparing it with adjacent tissues, which was consistent with its expression trend in the TCGA data set (p < 0.05; Fig. 4a-b). and a significantly positive correlation between TBX5-AS1 and TBX5 were seen in the samples (Spearman’s rank correlation coefficient r = 0.6893, p = 0.008) (Fig. 4c). Furthermore, immunohistochemistry data extracted from the HPA indicated that the protein expression of TBX5 was higher in non-tumour tissues compared with that in tumour tissues (Fig. 4d-e).
Pan-Cancer verification
To determine the prognostic role of our selected eRNA in pan-cancer and the correlation with its target gene, we con-ducted survival and correlation analyses. The results showed that TBX5-AS1 was associated with survival in four tumours, namely, adrenocortical carcinoma (ACC), LUAD, uterine corpus endometrial carcinoma (UCEC) and lung squamous cell carcinoma (LUSC). The survival curves for TBX5-AS1 in these four tumours are shown in Fig. 5a–d. In addition, we found that TBX5-AS1 and its target gene were associated with 26 types of tumours (Additional file 1: Table S3).
Discussion
Although it has been reported lncRNA TBX5-AS1 as antisense RNA of TBX5 could serve as potential diagnostic biomarkers for NSCLC and could inhibit the tumor progression of NSCLC through inactivating the PI3K/AKT pathway [27,28,29], the exact mechanism between TBX5-AS1 and TBX5 in LUAD has not been reported and there are no studies used eRNA as a biomarker for LUAD. In this study, we performed a comprehensive data analysis approach and found that TBX5-AS1 belong to a subclass of lncRNAs, termed “enhancer RNA”. Moreover, we also clarified the possible relationship between TBX5-AS1 and TBX5 and elaborated the involved pathways and possible molecular mechanism of TBX5-AS1 exert effects in LUAD.
eRNAs are noncoding RNAs that are transcribed from enhancers. Studies have shown that enhancers usually regulate the expression of target genes by forming a chromatin loop with the target promoter [8, 30]. For example, Rosenfeld et al. found that estrogen-regulated eRNAs can bind cohesin complex proteins, including RAD21 cohesin complex component (RAD21) and structural maintenance of chromosomes 3 (SMC3), and the authors proposed that eRNA/cohesin interactions serve to stabilize chromatin looping in breast cancer cells [31]. Additionally, mediator complex subunit 1 (MED1) in prostate cancer cells [21], heterogeneous nuclear ribonucleoprotein U (hnRNPU) in gastric cancer cells [19], CCCTC-binding factor (CTCF) in colon cancer cells [32], and mediator complex subunit 12 (MED12) in T-cell acute lymphoblastic leukaemia cells [33] have been shown to regulate chromatin loop stability through interactions with eRNAs, thereby modulating the expression of downstream target genes.
In the present study, we obtained RNA expression profile data and clinical information for LUAD from the TCGA database and integrated these data with eRNA-related information. Using Spearman’s correlation and Kaplan–Meier analyses, we identified 76 candidate eRNAs related to the prognosis of LUAD. We further identified TBX5-AS1 as the most critical eRNA candidate sequence in LUAD, and TBX5 as its regulatory target. Clinical correlation analysis showed that TBX5 was differentially expressed according to T stage, N stage, gender, and cancer status. KEGG pathway enrichment results suggested that TBX5 may affect the survival outcome of LUAD patients through the PI3K/AKT pathway, Ras signaling pathway, cGMP−PKG signaling pathway, MAPK signaling pathway, etc. PI3K/AKT pathway is one of the most enriched biological process. Our pan-cancer validation results revealed that TBX5-AS1 was associated with survival in four types of tumours (ACC, LUAD, LUSC, and UCEC). Furthermore, TBX5-AS1 expression was correlated with that of its target gene, TBX5, in 26 tumour types. Finally, we validated the expression of TBX5-AS1 in surgical specimens by qRT-PCR and examined the expression levels of TBX5 in LUAD using the HPA databases. Together, our results indicated that TBX5-AS1 could be used as an independent predictor of LUAD.
TBX5, a member of the T-box gene family, plays an important role in heart development [34,35,36]. Members of this family are involved in a variety of processes, including cell–cell signalling, proliferation, apoptosis, and migration [37]. Studies have shown that TBX5 dysregulation plays an important role in breast cancer [37], gastric cancer [38], oesophageal carcinoma [39, 40], and colon cancer [41]. For example, the TBX5 mRNA expression level is significantly downregulated in colorectal cancer cells; its methylation rate is significantly higher in cancer tissues than in normal tissues; and overexpressing TBX5 can inhibit the growth of colorectal cancer cells, promote the apoptosis of cancer cells, and reduce the migration rate of cancer cells [41]. In this study, we found that TBX5 was highly expressed in normal paracancerous tissues and cell lines, whereas its expression was downregulated in LUAD tissue and cells. These findings revealed that the level of TBX5 is negatively correlated with lung cancer progression, which is consistent with the regulation of TBX5 in colon cancer [41]. Besides, Ma et al. (2017) found that TBX5 may induce early and late apoptosis in NSCLC cells by activating key molecules in the apoptotic pathway, including cleaved caspase-3, cleaved caspase-8, CDKN2, and PARP. Moreover, the overexpression of TBX5 can inhibit cell proliferation, clone formation, and invasion, as well as induce apoptosis, in a NSCLC cell line [42]. To our knowledge, the specific interaction mechanism of TBX5-AS1 and TBX5 in LUAD has not been reported previously. Our results suggest that eRNA TBX5-AS1 might exert its antitumour effects through the following way: TBX5-AS1 may regulate chromatin loop stability by binding to specific protein factors, which in turn promotes the expression of its target gene TBX5. And then, TBX5 suppress the invasion lung adenocarcinoma by affecting PI3K/AKT pathway, Ras signaling pathway, cGMP−PKG signaling pathway, MAPK signaling pathway, etc. In summary, the present findings suggested that TBX5-AS1 has potential as a diagnostic biomarker and therapeutic target for LUAD. In the future, in the early stage of disease, we could be detect the expression level of eRNA TBX5-AS1 in blood or other body fluids to determine whether the patients has LUAD. For patients who have been diagnosed with LUAD, we can predict the prognosis of lung adenocarcinoma patients based on TNM stage and the expression level of TBX5-AS1 in blood. This type of inspection would be particularly popular among patients with LUAD. Although lung biopsy histological examination is the gold standard for LUAD diagnosis and disease surveillance, histopathological examination is invasive, labour intensive and expensive which may impose a heavy financial burden on patients. Therefore, once our candidate biomarkers have been validated in blood and other body fluid samples, we can confirm or exclude lung adenocarcinoma through a simple and inexpensive blood test which would be significant.
Although we identified the key eRNA in LUAD and investigated its possible role in this disease, our study had several limitations. For instance, a larger clinical cohort is needed to further assess the diagnostic and prognostic potential of TBX5-AS1 in LUAD, while the potential function of TBX5-AS1 in LUAD also requires further investigation. Moreover, we demonstrated that TBX5-AS1 has roles in multiple cancer types, including ACC, LUAD, LUSC, and UCEC, and the emerging roles of TBX5-AS1 in these cancers also need to be characterized.
Conclusion
In conclusion, we showed for the first time that TBX5-AS1 is a prognostic eRNA for LUAD, and through further prospective analysis, our study may help to evaluate prognosis of patients with LUAD, contribute to the early diagnosis of LUAD and provide treatment options for developing clinical interventions.
Availability of data and materials
The data supporting the findings of this study are available from the corresponding author on reasonable request. The datasets generated during the current study are available in the Cancer Genome Atlas (TCGA) http://portal.gdc.cancer.gov/), the The Human Protein Atlas (HPA) https://www.proteinatlas.org/).
Abbreviations
- ERNAs:
-
Enhancer RNAs
- LUAD:
-
Lung adenocarcinoma
- TCGA:
-
The Cancer Genome Atlas
- HPA:
-
The human protein atlas
- qRT-PCR:
-
quantitative real time-polymerase chain reaction
- TBX5:
-
T-box transcription factor 5
- ACC:
-
Adrenocortical carcinoma
- LUSC:
-
Lung squamous cell carcinoma
- UCEC:
-
Uterine corpus endometrial carcinoma
- NSCLC:
-
Non-small cell lung cancer
- T-ALL:
-
T-cell acute lymphoblastic leukemia
- HNSCC:
-
Head and neck squamous cell carcinoma
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- GO:
-
Gene Ontology
- MF:
-
Molecular function
- BP:
-
Biological process
- CC:
-
Cellular component
- SMC3:
-
Structural maintenance of chromosomes 3
- MED1:
-
Mediator complex subunit 1
- HnRNPU:
-
Heterogeneous nuclear ribonucleoprotein U
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70(1):7–30. https://doi.org/10.3322/caac.21590.
Wang L, Yu C, Liu Y, Wang J, Li C, Wang Q, et al. Lung Cancer mortality trends in China from 1988 to 2013: new challenges and opportunities for the government. Int J Environ Res Public Health. 2016;13(11):1052. https://doi.org/10.3390/ijerph13111052.
Herbst RS, Heymach JV, Lippman SM. Lung cancer. N Engl J Med. 2008;359(13):1367–80. https://doi.org/10.1056/NEJMra0802714.
Comprehensive molecular profiling of lung adenocarcinoma. Nature. 2014;511(7511):543–50. https://doi.org/10.1038/nature13385.
Djebali S, Davis CA, Merkel A, Dobin A, Lassmann T, Mortazavi A, et al. Landscape of transcription in human cells. Nature. 2012;489(7414):101–8. https://doi.org/10.1038/nature11233.
Hangauer MJ, Vaughn IW, McManus MT. Pervasive transcription of the human genome produces thousands of previously unidentified long intergenic noncoding RNAs. PLoS Genet. 2013;9(6):e1003569. https://doi.org/10.1371/journal.pgen.1003569.
Lee JH, Xiong F, Li W. Enhancer RNAs in cancer: regulation, mechanisms and therapeutic potential. RNA Biol. 2020;17(11):1550–9. https://doi.org/10.1080/15476286.2020.1712895.
Natoli G, Andrau JC. Noncoding transcription at enhancers: general principles and functional models. Annu Rev Genet. 2012;46(1):1–19. https://doi.org/10.1146/annurev-genet-110711-155459.
Andersson R, Gebhard C, Miguel-Escalada I, et al. An atlas of active enhancers across human cell types and tissues. Nature. 2014;507(7493):455–61. https://doi.org/10.1038/nature12787.
Li W, Notani D, Rosenfeld MG. Enhancers as non-coding RNA transcription units: recent insights and future perspectives. Nat Rev Genet. 2016;17(4):207–23. https://doi.org/10.1038/nrg.2016.4.
Chen H, Li C, Peng X, Zhou Z, Weinstein JN, Liang H, et al. A pan-Cancer analysis of enhancer expression in nearly 9000 patient samples. Cell. 2018;173(2):386–99. https://doi.org/10.1016/j.cell.2018.03.027.
Grosschedl R, Birnstiel ML. Spacer DNA sequences upstream of the T-A-T-A-A-A-T-A sequence are essential for promotion of H2A histone gene transcription in vivo. Proc Natl Acad Sci U S A. 1980;77(12):7102–6. https://doi.org/10.1073/pnas.77.12.7102.
Benoist C, Chambon P. In vivo sequence requirements of the SV40 early promoter region. Nature. 1981;290(5804):304–10. https://doi.org/10.1038/290304a0.
Andersson R. Promoter or enhancer, what’s the difference? Deconstruction of established distinctions and presentation of a unifying model. Bioessays. 2015;37(3):314–23. https://doi.org/10.1002/bies.201400162.
Lam MT, Cho H, Lesch HP, Gosselin D, Heinz S, Tanaka-Oishi Y, et al. Rev-Erbs repress macrophage gene expression by inhibiting enhancer-directed transcription. Nature. 2013;498(7455):511–5. https://doi.org/10.1038/nature12209.
Lin YC, Benner C, Mansson R, Heinz S, Miyazaki K, Miyazaki M, et al. Global changes in the nuclear positioning of genes and intra- and interdomain genomic interactions that orchestrate B cell fate. Nat Immunol. 2012;13(12):1196–204. https://doi.org/10.1038/ni.2432.
Sanyal A, Lajoie BR, Jain G, Dekker J. The long-range interaction landscape of gene promoters. Nature. 2012;489(7414):109–13. https://doi.org/10.1038/nature11279.
Melo CA, Drost J, Wijchers PJ, van de Werken H, de Wit E, Oude Vrielink JAF, et al. eRNAs are required for p53-dependent enhancer activity and gene transcription. Mol Cell. 2013;49(3):524–35. https://doi.org/10.1016/j.molcel.2012.11.021.
Hsieh CL, Fei T, Chen Y, Li T, Gao Y, Wang X, et al. Enhancer RNAs participate in androgen receptor-driven looping that selectively enhances gene activation. Proc Natl Acad Sci U S A. 2014;111(20):7319–24. https://doi.org/10.1073/pnas.1324151111.
Bal E, Park HS, Belaid-Choucair Z, Kayserili H, Naville M, Madrange M, et al. Mutations in ACTRT1 and its enhancer RNA elements lead to aberrant activation of hedgehog signaling in inherited and sporadic basal cell carcinomas. Nat Med. 2017;23(10):1226–33. https://doi.org/10.1038/nm.4368.
Tan SH, Leong WZ, Ngoc P, et al. The enhancer RNA ARIEL activates the oncogenic transcriptional program in T-cell acute lymphoblastic leukemia. Blood. 2019;134(3):239–51. https://doi.org/10.1182/blood.2018874503.
Zhang Z, Lee JH, Ruan H, Ye Y, Krakowiak J, Hu Q, et al. Transcriptional landscape and clinical utility of enhancer RNAs for eRNA-targeted therapy in cancer. Nat Commun. 2019;10(1):4562. https://doi.org/10.1038/s41467-019-12543-5.
Gu X, Wang L, Boldrup L, Coates P, Fahraeus R, Sgaramella N, et al. AP001056.1, A Prognosis-Related Enhancer RNA in Squamous Cell Carcinoma of the Head and Neck. Cancers. 2019;11(3):347. https://doi.org/10.3390/cancers11030347.
Kanehisa M, Goto S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000;28(1):27–30. https://doi.org/10.1093/nar/28.1.27.
Kanehisa M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 2019;28(11):1947–51. https://doi.org/10.1002/pro.3715.
Kanehisa M, Furumichi M, Sato Y, Ishiguro-Watanabe M, Tanabe M. KEGG: integrating viruses and cellular organisms. Nucleic Acids Res. 2021;49(D1):D545–51. https://doi.org/10.1093/nar/gkaa970.
Shih JH, Chen HY, Lin SC, Yeh YC, Shen R, Lang YD, et al. Integrative analyses of noncoding RNAs reveal the potential mechanisms augmenting tumor malignancy in lung adenocarcinoma. Nucleic Acids Res. 2020;48(3):1175–91. https://doi.org/10.1093/nar/gkz1149.
Zhao T, Khadka VS, Deng Y. Identification of lncRNA biomarkers for lung cancer through integrative cross-platform data analyses. Aging (Albany NY). 2020;12(14):14506–27. https://doi.org/10.18632/aging.103496.
Qu QH, Jiang SZ, Li XY. LncRNA TBX5-AS1 regulates the tumor progression through the PI3K/AKT pathway in non-small cell lung Cancer. Onco Targets Ther. 2020;13:7949–61. https://doi.org/10.2147/OTT.S255195.
Schmitt AD, Hu M, Ren B. Genome-wide mapping and analysis of chromosome architecture. Nat Rev Mol Cell Biol. 2016;17(12):743–55. https://doi.org/10.1038/nrm.2016.104.
Li W, Notani D, Ma Q, Tanasa B, Nunez E, Chen AY, et al. Functional roles of enhancer RNAs for oestrogen-dependent transcriptional activation. Nature. 2013;498(7455):516–20. https://doi.org/10.1038/nature12210.
Jiao W, Chen Y, Song H, Li D, Mei H, Yang F, et al. HPSE enhancer RNA promotes cancer progression through driving chromatin looping and regulating hnRNPU/p300/EGR1/HPSE axis. Oncogene. 2018;37(20):2728–45. https://doi.org/10.1038/s41388-018-0128-0.
Xiang JF, Yin QF, Chen T, Zhang Y, Zhang XO, Wu Z, et al. Human colorectal cancer-specific CCAT1-L lncRNA regulates long-range chromatin interactions at the MYC locus. Cell Res. 2014;24(5):513–31. https://doi.org/10.1038/cr.2014.35.
Zhu T, Qiao L, Wang Q, Mi R, Chen J, Lu Y, et al. T-box family of transcription factor-TBX5, insights in development and disease. Am J Transl Res. 2017;9(2):442–53.
Zhang XL, Qiu XB, Yuan F, et al. TBX5 loss-of-function mutation contributes to familial dilated cardiomyopathy. Biochem Biophys Res Commun. 2015;55:1417–25.
Khan P, Linkhart B, Simon HG. Different regulation of T⁃box genes Tbx4 and Tbx5 during limb development and limb regeneration. Dev Biol. 2002;250(2):383–92. https://doi.org/10.1006/dbio.2002.0801.
Kim J, Siverly AN, Chen D, Wang M, Yuan Y, Wang Y, et al. Ablation of miR-10b suppresses oncogene-induced mammary Tumourigenesis and metastasis and reactivates tumor-suppressive pathways. Cancer Res. 2016;76(21):6424–35. https://doi.org/10.1158/0008-5472.CAN-16-1571.
Zheng Y, Li Y, Wang W, et al. High expression level of T-box transcription factor 5 predicts unfavorable survival in stage I and II gastric adenocarcinoma. Oncol Lett. 2015;10(4):2021–6. https://doi.org/10.3892/ol.2015.3515.
Becker J, May A, Gerges C, Anders M, Schmidt C, Veits L, et al. The Barrett⁃associated variants at GDF7 and TBX5 also increase esophageal adenocarcinoma risk. Cancer Med. 2016;5(5):888–91. https://doi.org/10.1002/cam4.641.
Palles C, Chegwidden L, Li X, Findlay JM, Farnham G, Castro Giner F, et al. Polymorphisms near TBX5 and GDF7 are associated with increased risk for Barrett′s esophagus. Gastroenterology. 2015;148(2):367–78. https://doi.org/10.1053/j.gastro.2014.10.041.
Yu J, Ma X, Cheung KF, Li X, Tian L, Wang S, et al. Epigenetic inactivation of T⁃box transcription factor 5, a novel tumor suppressor gene, is associated with colon cancer. Oncogene. 2010;29(49):6464–74. https://doi.org/10.1038/onc.2010.370.
Ruoting M, Yu Y, Qiuyun T, et al. Overexpression of T-box transcription factor 5 (TBX5) inhibits proliferation and invasion in non-small cell lung carcinoma cells. Oncol Res. 2017;25:1495–504.
Acknowledgments
We acknowledge TCGA database for providing their platforms and contributors for uploading their meaningful datasets.
Funding
This work was supported by the National Natural Science Foundation of China (No.81873169) and Hunan Provincial Natural Science Foundation of China (No. 2020JJ4803).
Author information
Authors and Affiliations
Contributions
Conception and design: WP; Administrative support: KN, BC; Provision of study materials or patients: TH; Collection and assembly of data: LC, BC; Data analysis and interpretation: LC, TH; Manuscript writing: LC; Final approval of manuscript: All authors.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The study was conducted according to the guidelines of the Declaration of Helsinki, and approved by the Ethics Committee of the Second Xiangya Hospital of Central South University (protocol code 2020038 and 07/01/2020) and all patients gave their informed consent for inclusion before surgery.
Consent for publication
Informed consent was obtained from all subjects involved in the study for the publication of the data.
Competing interests
The authors declare no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Table S1.
List of 174 survival-related eRNAs in lung adenocarcinoma. Table S2. List of the 76 eRNAs with a significant correlation with their target gene in lung adenocarcinoma. Table S3. List of the 26 types of tumours associated with TBX5-AS1 and TBX5.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Cheng, L., Han, T., Chen, B. et al. TBX5-AS1, an enhancer RNA, is a potential novel prognostic biomarker for lung adenocarcinoma. BMC Cancer 21, 794 (2021). https://doi.org/10.1186/s12885-021-08517-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12885-021-08517-w