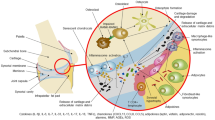
Chapter summary
The role of matrix metalloproteinases in the degradative events invoked in the cartilage and bone of arthritic joints has long been appreciated and attempts at the development of proteinase inhibitors as potential therapeutic agents have been made. However, the spectrum of these enzymes orchestrating connective tissue turnover and general biology is much larger than anticipated. Biochemical studies of the individual members of the matrix metalloproteinase family are now underway, ultimately leading to a more detailed understanding of the function of their domain structures and to defining their specific role in cellular systems and the way that they are regulated. Coupled with a more comprehensive and detailed study of proteinase expression in different cells of joint tissues during the progress of arthritic diseases, it will be possible for the future development and application of highly specific proteinase inhibitors to be directed at specific key cellular events.
Similar content being viewed by others
Introduction
The past decade has seen major advances in the understanding of the pathogenesis of arthritic diseases and has markedly influenced pharmacological approaches. The development of inflammatory rheumatic diseases, including an initiation phase associated with genetic susceptibilities, the activation of synovial cells and the development of the pannus are well characterised. Subsequent cartilage and bone destruction leads to an irreversible pathology. Osteoarthritis (OA) is far less well understood, but also leads ultimately to joint tissue destruction. Hence, a detailed knowledge of the events underlying these degradative processes is a prerequisite for all arthritic diseases for the development of therapies targeting their prevention. Key to this is an understanding of the proteinases involved in terms of their regulation and specific function.
This review outlines the detailed studies of the background biochemistry of the matrix metalloproteinases (MMPs), which are major players in extracellular matrix (ECM) turnover in both physiology and pathology. Such studies are the basis of specific inhibitor development as potential therapies. The study of metalloproteinase expression in relation to the progress of different forms of arthritis is outlined. It is clear that the latter picture is particularly incomplete, with most studies focused on the MMPs identified some years ago but rarely focused on newer potential contributors to the degradative pathology of arthritic tissues. Future approaches will need to look at the overall patterns of proteinase function in joint tissues allowing the most precise targeting of new generations of highly specific inhibitors.
Historical background
Progressive degradation of the ECM that comprises joint tissues, including articular cartilage, bone and even intra-articular ligaments and tendons, is a major feature of the arthritic diseases, leading to permanent loss of function. Although proteinases of all mechanistic classes play a role in the degradation of connective tissue macromolecules, it has long been thought that the major activities involved in this process belong to the family of MMPs. These enzymes are secreted by both the resident cells of joint tissues as well as by invading cells, they are active around neutral values of pH, and they have the combined ability to degrade all the components of the ECM (Table 1). MMPs play significant roles in both developmental and repair processes, and it appears that aberrant regulation, which can occur at many levels (see next section), leads to their hyperactivity in diseases such as rheumatoid arthritis (RA) and OA.
There is now significant evidence for the overexpression of MMPs in tissues derived from patients with arthritic disease. Cultures of cells derived from rheumatoid synovia secreted a collagenolytic activity into the medium [1], and stromelysin-1 [2] and collagenase-1 were detected by immunolocalisation at sites of cartilage erosion in rheumatoid joints [3]. Both of these enzymes, as well as the tissue inhibitor of metalloproteinase (TIMP)-1, were immunolocalised in synovial samples from both RA and OA patients [4].
The finding of stromelysin-1 in all synovial samples from 10 patients with different clinical diagnoses and histories, in contrast to its absence from normal synovia [5], clearly implicated this enzyme in the arthritic process. It was also shown that collagenase-1, gelatinase A and matrilysin may have a role in the synovitis associated with RA, but that they are not a significant feature in osteoarthritic joints. Marked regional variations were found in the synthesis of these MMPs, however, indicating that these diseases are episodic and that the control of enzyme synthesis is focal. This indicates the need for further work to colocalise MMP synthesis with cytokine and matrix expression in synovia from diseased joints in order to explore further the mechanisms that control the synthesis and degradation of ECM components of articular cartilage. Stromelysin-1 and collagenase-1 have also been measured in the synovial fluids from rheumatoid and osteoarthritic knee joints [6–8].
Other studies of MMP expression in normal and diseased cartilage have documented the presence of stromelysin-1, with lower levels of collagenase-1 [9]. Gelatinase B is detected as both mRNA and protein in osteoarthritic cartilage, but not in normal tissue [10], and is detectable in the synovial fluid from rheumatoid joints. Collagenase-2 and collagenase-3 have more recently been identified in arthritic cartilage [11–14]. Konttinen et al. [15] analysed the expression of 16 MMPs at the mRNA level in trauma and RA, and they found some (e.g. collagenase-3 and the membrane-type matrix metalloproteinase [MT-MMP] MT2-MMP) exclusively present in the rheumatoid tissue.
The precise targets of MMP action in joint tissues are not always clear. In particular, there has been extensive debate concerning their role in the degradation of cartilage proteoglycans relative to that of the related family of a disintegrin, a metalloproteinase and thrombospondin 'aggrecanase' (ADAM-TS) [16, 17]. Kozaci et al. [18] used a model system of collagen degradation to conclude that stromelysin-1, collagenase-2 and collagenase-3 are unlikely to contribute to proteoglycan degradation, but that collagenases and gelatinase have major roles in type II collagen breakdown. MT1-MMP is expressed by rheumatoid synovial fibroblasts and is regulated by tumour necrosis factor alpha [19]. The identity of the collagenase responsible for cartilage collagen loss is also still a subject of detailed study. There is evidence that specific MMP-13 inhibitors can block interleukin-1-induced collagen loss, and an enhanced cleavage of type II collagen in osteoarthritic cartilage seems to be correlated with MMP-13 activity [20].
The MMP family
The MMP family consists of 25 zinc-dependent and calcium-dependent proteinases in mammalian systems (Table 1), and MMPs are now thought to be the major proteolytic enzymes that facilitate tissue remodelling in both physiological and pathological situations [21–23]. The MMPs do indeed have the combined ability to degrade the major components of the ECM [24].
MMPs can be classified into at least five main groups, according to their substrate specificity, primary structure and cellular localisation; namely, the collagenases, gelatinases, stromelysins, matrilysins and MT-MMPs [25]. There are some MMPs, however, such as macrophage elastase (MMP-12), stromelysin-3 (MMP-11), MMP-19, enamelysin (MMP-20), CA-MMP (MMP-23) and epilysin (MMP-28), that apparently do not fall into any of these categories. In addition, some enzymes, such as MT1-MMP (MMP-14), which displays collagenolytic activity and is membrane associated, may be classified into more than one group.
The collagenases (collagenase-1 [MMP-1], collagenase-2 [MMP-8] and collagenase-3 [MMP-13]) are able to cleave native triple-helical fibrillar collagens (i.e. type I, type II and type III) at a single bond, generating characteristic one-quarter and three-quarter fragments.
Two gelatinases (gelatinase A [MMP-2] and gelatinase B [MMP-9]) have been identified. Gelatinase A is expressed by a broad spectrum of mesenchymal cells, whereas gelatinase B is associated with macrophages and peripheral blood mononuclear cells, as well as activated connective tissue cells and tumours. The gelatinases have a broad substrate specificity and may contribute, together with collagenases, to the degradation of fibrillar collagens, basement membrane components and stromal ECM molecules (e.g. fibronectin).
Stromelysins (stromelysin-1 [MMP-3] and stromelysin-2 [MMP-10]) have one of the broadest substrate spectra of the MMPs and can degrade most ECM components, such as gelatin, fibronectin, laminin and aggrecan, but not triple-helical collagens. They are expressed by synovial fibroblasts from a rheumatoid joint but not by normal synovial cells. Stromelysin-3 (MMP-11), however, has weak proteolytic activity and is activated intracellularly by a furin-like convertase. Both these properties distinguish it from the other two members of the stromelysin group.
Although matrilysins (matrilysin-1 [MMP-7] and matrilysin-2 [MMP-26]) lack the hinge region and the COOH-terminal, hemopexin-like domain common to almost all other members of the MMP subfamily, they are potent proteinases. Matrilysin-1 is expressed by a range of benign and malignant tumours, whereas matrilysin-2 was only recently cloned and detected in uterus, placenta and endometrial tumours.
The group of six cell-associated MT-MMPs (MT1-MMP [MMP-14], MT2-MMP [MMP-15], MT3-MMP [MMP-16], MT4-MMP [MMP-17], MT5-MMP [MMP-24] and MT6-MMP [MMP-25]) seems to be activated intracellularly by a furin-type proprotein convertase. With the exception of MT4-MMP and MT6-MMP, which are membrane bound via a glycosylphosphatidylinositol molecule and not via a transmembrane domain, these enzymes also contain a COOH-terminal cytoplasmic tail.
MT1-MMP was shown to be a major activator of proMMP-2 on the cell surface. This activation is thought to be facilitated via a ternary complex of MT1-MMP, TIMP-2 and proMMP-2, enabling cells (e.g. tumour cells) to invade stromal tissue or to cross the basement membrane of blood vessels during metastasis. With the exception of MT6-MMP, which is predominantly expressed in leukocytes, the other MT-MMPs are found in many cell types, although they have not as yet been rigorously studied in relation to arthritic tissues.
Structure and function of MMP domains
MMPs are multidomain proteins consisting of a signal peptide, a propeptide, and the catalytic and COOH-terminal domains, as shown in Figure 1. MT-MMPs and gelatinases have additional features such as a COOH-terminal transmembrane region followed by a short cytotail, and three fibronectin type II repeats in the catalytic domain, respectively.
Basic domain structures of the matrix metalloproteinases (MMPs). MMPs consist of: a propeptide (grey), which maintains the enzymes in a latent state; a catalytic domain (blue) with the active site and the catalytic zinc (Zn) (red); and, with the exception of the matrilysins, a COOH-terminal domain (C) (yellow) with homology to the serum protein hemopexin. The latter two domains are connected by a linker peptide. Gelatinases have an insert of three fibronectin type II repeats (turquoise) in the catalytic domain, which is involved in substrate recognition. Membrane-type MMPs contain a transmembrane domain (black) and a cytoplasmic tail (green) at the COOH terminus, which anchors these enzymes in the cell membrane.
All MMPs are translated with a NH2-terminal hydrophobic sequence of 18–30 residues that is responsible for trafficking of the enzyme through the endoplasmic reticulum and the Golgi apparatus, and for its subsequent secretion into the extracellular space. The signal peptide is cleaved off during the secretion process. Interestingly, the signal peptide in CA-MMP (MMP-23) contains a transmembrane motif that anchors the enzyme to the membrane. Activation and secretion is achieved by cleavage in a proprotein convertase recognition sequence located between the propeptide and the catalytic domain.
The propeptide domain, N-terminal to the catalytic domain, consists of about 80 residues arranged in three α-helices, and it is responsible for enzyme latency. This domain contains the sequence motif PRCGVPD (also called the 'cysteine switch' motif), which is highly conserved among MMPs. In the latent enzyme, the propeptide domain is located directly opposite the active site cleft and coordinates the catalytic zinc ion with the thiol of the cysteine residue, thus preventing binding of a water molecule required for peptide hydrolysis [26]. Apart from the direct coordination of the zinc ion, several β-structure-like interchain hydrogen bonds are formed between the propeptide and the catalytic site, similar to the bonds established with substrates and inhibitors. It should be noted, however, that the binding orientation of the propeptide is the opposite to that of the substrate
All MMPs (except the MT-MMPs, stromelysin-3 and epilysin, which are activated intracellularly by a furin-type proprotein convertase) probably become activated in a stepwise manner outside the cell by proteolytic cleavage. Activating proteases, like MMPs, serine or cysteine proteinases, attack the so-called 'bait region', an exposed and flexible stretch of around 30–40 residues, which probably results in destabilisation of the cysteine switch-zinc interactions [26]. These molecular rearrangements promote the full activation of the enzyme by exposing the final cleavage site between propeptide and catalytic domain, generally around residues 80–90, to proteolytic cleavage. The activation of proMMPs (e.g. by other proteinases and MMPs) will be discussed in more detail later.
The MMP catalytic domain consists of approximately 160–170 residues and contains the active centre with the catalytic zinc ion, responsible for substrate hydrolysis and inhibitor interactions. As already described, three conserved histidine residues in the HExxHxxGxxH motif, located 50–55 residues from the COOH-terminal end of the domain, ligate the catalytic zinc ion. X-ray analysis of various MMP catalytic domains, some of which complexed with an inhibitor (i.e. collagenase-1, stromelysin-1, matrilysin-1, collagenase-2 and MT1-MMP), revealed high structural similarity among enzymes.
Essentially, the MMP catalytic domain is made up of a twisted β-sheet covering two long α-helices (helix A and helix B), and of a separate helix at the COOH-terminal end of the domain. The sheet itself consists of four parallel β-strands (strand I, strand II, strand III and strand V) and one antiparallel β-strand (strand IV) (Fig. 2). In addition to the catalytic zinc, a second zinc ion (the so-called 'structural zinc') and one to three calcium ions (depending on the MMP) are also bound in this domain and seem necessary for its function. Two of the three histidine residues coordinating the active site zinc are located within helix B, whereas the third histidine is located in the loop between helices B and C and lies opposite the other two histidine residues (Fig. 2).
Structure of the catalytic domain of the matrix metalloproteinase MMP-3. The ribbon diagram was created using WebLab Viewer software based on a crystal structure analysis by Gomis-Rüth et al. [61]. Strands (sI–sV) and helices (hA–hC) are labelled in black; the catalytic zinc (centre), coordinated by three histidine residues (pink), and the structural zinc (Zn) (top) are labelled in black and white, respectively; and the NH2 terminus (N) and COOH terminus (C) are labelled in red. The active site cleft and the characteristic methionine (Met) turn are indicated with arrows.
The typical MMP active site cleft, carved into the surface of the ellipsoidal domain, is relatively flat on the left-hand side (nonprimed) and contains the catalytic zinc in its centre, but is deep on the right-hand side (primed) [27]. Studies using peptide substrates revealed that a substrate is fixed to its cognate MMP through seven intermain-chain hydrogen bonds and that the scissile peptide bond carbonyl group is directed towards the catalytic zinc ion to be strongly polarised. For the mechanism of substrate hydrolysis by MMPs, it was proposed that the glutamic acid residue within the HExxHxxGxxH motif then polarises a water molecule for a nucleophilic attack on the carbonyl carbon atom of the substrate scissile amino bond.
The sequence requirements for a certain substrate to be cleaved depend on the depth and structure of the catalytic site of the MMP. For example, dominant hydrophobic interactions are made through P1' and P3 residue side chains of the substrate and the respective pockets in the MMP. P1, P2, P3, etc. and P1', P2', P3', etc. indicate peptide substrate residues in the NH2-terminal and COOH-terminal directions from the scissile bond, respectively, whereas S1, S2, S3, etc. and S1', S2', S3', etc. indicate the opposing subsites of the MMP [27].
In general, proline and leucine residues at the P3 and P2 positions of the substrate, respectively, are the preferred amino acids for MMPs. A large residue in the P1' position of the substrate is also preferential for binding to the MMP active site. This residue interacts with the so-called 'specificity pocket' (S1' pocket), a deep cavity within the MMP active site. The S1' pocket is a varied structural feature in the active site of MMPs, and it has a major influence on substrate and inhibitor specificity. Stromelysin-1 and MT1-MMP both have very deep hydrophobic S1' pockets and prefer residues with aliphatic or aromatic side chains in the P1' position of the substrate. In fact, synthetic peptides containing unusual amino acids with extremely long side chains in the P1' position were hydrolysed by MT1-MMP and stromelysin-1 with higher efficiency than peptides containing natural amino acids in the same position.
Collagenases
Studies on the catalytic domains of the collagenases have shown that they contribute to the specificity for the cleavage of triple-helical collagen. We investigated exon 5 of collagenase-1 in this respect by exchanging it with the exon 5 of the noncollagenolytic proteinase MMP-3. Exon 5 harbours major features of the active site including the three histidine ligands of the functional zinc, the S1' binding site and components of the other substrate binding sites, S2', S3' and S1–S3. The overall exon 5 sequence shows a high degree of similarity between collagenase-1 and stromelysin-1. The major variations include the greater hydrophilicity of collagenase-1 and its slightly smaller size. Hence the exchange of exon 5 causes no predictable major structural changes.
The exon 5 mutant of collagenase-1 did demonstrate a change in peptide substrate and small inhibitor specificity as well as binding to N-TIMP-1, the three active N-terminal loops becoming more stromelysin-1-like. The exon 5 mutant also lost most of its ability to cleave type I collagen as well as a reduction in gelatin turnover, indicating that the active site cleft is itself a determinant of specific collagenolytic capacity [28].
A unique feature of the catalytic domain of gelatinases A and B is the presence of three fibronectin type II repeats just upstream of the zinc binding motif, contributing around 20 kDa to the size of the enzyme. These repeats are located between strand V and helix B of the MMP catalytic domain, consist of two double-stranded antiparallel β-sheets and two large irregular loops each, and are important for binding of the substrate in the active site. It was demonstrated that recombinantly expressed repeats bind to denatured type I collagen, denatured type IV and type V collagens, elastin and native type I collagen. While deletion of the three repeats in MMP-2 resulted in decreased gelatinolytic activity and abrogated binding to collagen, TIMP binding was not affected.
Between the catalytic domain and the COOH-terminal hemopexin-like domain lies a variable stretch of 2–72 amino acids, which is termed the 'linker' or 'hinge region'. This region is very extended in the case of collagenases, being rich in proline residues and possibly being responsible for activity versus fibrillar collagens. A hypothetical mechanistic contribution of these linker peptides has been postulated, whereby the proline-rich peptides adopt a helical conformation that might 'unwind' the helix. We analysed the hinge region of collagenase-2 by alanine mutagenesis and found that this region had a pronounced effect on the stability and collagenolytic activity of the enzyme. Notably, some proline residues (shown underlined) were modified to alanine in the sequence Gly-Leu-Ser-Ser-Asn-Pro-Ile-Gln-Pro-Thr-Gly-Pro-Ser-Thr-Pro-Lys-Pro, to show that they were critical to the collagenolytic mechanism [29].
The MMP COOH-terminal domain has strong sequence similarity with the serum protein hemopexin and consists of approximately 200 amino acids. It has the shape of an ellipsoidal disk, as demonstrated by X-ray analysis of the crystal structures of this domain in porcine collagenase-1, human gelatinase A and human collagenase-3. The domain consists of four β-sheets (blades I–IV) with very similar structure, each consisting of four antiparallel β-strands, which are arranged almost symmetrically around a central core. The overall appearance is that of a four-bladed propeller, whereby the innermost strands of all four blades are arranged almost parallel to each other, building a central channel that contains Ca2+ and Cl- ions. The stability of this structure is ensured by a disulfide bond between blades I and IV, which is conserved in all MMPs.
The role of the C-terminal hemopexin-like domain varies widely according to the MMP, one of the most notable being the ability to interact with fibrillar collagens. The hemopexin-like domain of stromelysin-1 was shown to bind to type I and type II collagen, although neither can be cleaved by stromelysin-1. In contrast, all collagenolytic enzymes appear to require the hemopexin-like domain for hydrolysis of triple-helical collagens. In our laboratory, we have prepared C-terminal domain deletion mutants of collagenases, gelatinase A and MT1-MMP to show that, in each case, the domain is absolutely required for 'specific collagenolysis' (i.e. cleavage of type I collagen helix 3/4 from the N-terminus) to occur [3–8].
Mutants of the collagenases comprised of the prodomain and catalytic domain are peptidolytic on removal of the propeptide, but cannot cleave native fibrillar collagen. Cleavage of other collagens (e.g. type IV collagen) by collagenase-3 and other matrix macromolecules does not seem to require the C-terminal domain. Furthermore, 'exchange' of C-terminal domains between MMPs (e.g. N-terminal collagenase-1-C-terminal MMP3, N-terminal collagenase-3-C-terminal MMP-19) generates proteinases that either do not exhibit specific collagen cleaving potential or, in the case of collagenase-3, gelatinase A and gelatinase A-stromelysin-1, exhibit much reduced activity [28, 30, 31] (Knäuper et al., unpublished data).
Gelatinase A is a remarkable 'collagenase', although it acts very slowly on native collagens relative to denatured forms. We found that deletion of the collagen-gelatin binding type II fibronectin-like domain caused a loss of the ability of MMP-2 to cleave gelatin efficiently or to bind to collagen or gelatin [32, 33]. However, cleavage of native collagen still absolutely required the presence of the C-terminal domain rather than the fibronectin-like domain. Indeed, the mechanism of MMP-2 cleavage of collagen proceeds in two phases: the first resembling that of the interstitial collagenases, and the second being gelatinolysis [31].
The COOH-terminal domain is thus necessary for activity of the collagenases versus fibrillar collagens, but it does not appear to be important for substrate specificity of most other MMPs. McQuibban et al. recently reported a physiological substrate of gelatinase A, monocyte chemoattractant protein-3, however, which was discovered using the gelatinase A hemopexin-like domain as bait in the yeast two-hybrid system [34].
We have constructed and expressed recombinant forms of gelatinase B in which the C-terminal domain has been deleted to compare the activation, catalytic and TIMP binding properties with the full-length enzyme. In vitro, the truncated MMP-9 and wild-type MMP-9 behaved identically when activated by organomercurial or stromelysin-1. When we assessed activation in a cell-based system using either plasminogen or stromelysin-1, both forms of MMP-9 activated at similar rates. Furthermore, the active form of C-truncated gelatinase B showed similar kinetics to wild-type gelatinase B for the turnover of peptide substrates and gelatin. Indeed, the only differences in the behaviour of truncated MMP-9 and full-length MMP-9 was found to be in the rate of binding to TIMP-1.
Full-length TIMP-1 inhibited gelatinase B in a biphasic manner, with a rapid first phase yielding 50–70% inhibition, apparently dependent on the particular enzyme preparation, followed by a slow phase. In comparison, the C-truncated gelatinase B binding to TIMP-1 was difficult to analyse as the data were suggestive of an isomerisation of the enzyme-inhibitor complex [35]. It was apparent that binding of TIMP-1 was severely abrogated by the lack of the C-terminal domain of gelatinase B, reducing the value of the association constant by one to three orders of magnitude [35]. In comparison, association of both forms of gelatinase B with N-TIMP-1 in which the three C-terminal loops have been deleted gave binding constant values similar to those of the slow phase of wild-type TIMP-1-gelatinase B binding. TIMP-2, however, does not show any discrimination in its binding to different forms of gelatinase B. No biological explanation for the 'special' association between TIMP-1 and gelatinase B has been established.
Studies on the domains of gelatinase A have been quite extensive. The effect of deletion of the fibronectin type II-like domain was, to some extent, discussed earlier under collagenases. Apart from a role in the binding of the enzyme to collagen and the turnover of gelatin, this domain does not appear to be required for the turnover of peptide substrates or for the binding of low molecular weight active site inhibitors or TIMPs [32].
The effect of deletion of the C-terminal domain of progelatinase A and the loss of ability to cleave native collagen was also discussed earlier. It has also been shown that the C-domain of gelatinase A is essential for its binding and activation at the surface of many cell types. It has been established that this involves the ability of the C-domain of gelatinase A to interact quite tightly with the inhibitor TIMP-2 as a complex with the membrane-associated MT1-MMP [36–38]. Kinetic studies of the domain interactions between TIMP-2 and gelatinase A show very fast association rates between the full-length proteins that are markedly abrogated by truncation of the TIMP-2 to the N-terminal three loops or of the gelatinase A to remove the C-terminal domain. TIMP-2 binding to the C-domain of gelatinase A is very tight and can occur in the absence of catalytic domain interactions in progelatinase A. The binding has two identifiable components: a charged interaction (salt sensitive) involving the C-terminal tail of the C-terminal three loops of TIMP-2, and a hydrophobic interaction between the remainder of the TIMP-2 and the C-domain of gelatinase A [39].
Our analyses of domain motif function in the membrane-associated MT1-MMP have shown that the C-terminal domain is required for collagenolysis, as already described [40]. Kinetic studies with the TIMPs, however, have shown that the C-domain plays a negligible role in TIMP interactions. TIMP-1 has very little ability to inhibit MT1-MMP for reasons that are not yet entirely clear. However, TIMP-2, TIMP-3 and TIMP-4 are effective inhibitors, either as full-length forms or as the N-terminal three loops, against full-length or C-truncated MT1-MMP. Hence, the complexes of TIMPs with MT1-MMP are all theoretically able to bind the C-domain of MMP2 simultaneously since the necessary binding sites should remain available.
One difference between the MT-MMPs and other MMP family members is the insertion of eight amino acids between strands βII and βIII in the catalytic domain (163-PYAYIREG-170 in MT1-MMP). To investigate the role of this region of MT1-MMP in its function, we have made a number of mutations and deletions in this region. The motif appeared to have no role in the cleavage of peptide substrates or extracellular proteins such as fibrinogen or in the binding of TIMP-2. However, the motif is involved in interactions with progelatinase A, facilitating both the solution-phase MT1-MMP activation mechanism as well as that at the cell surface [41].
The collagenase-3 C-terminal domain deletion abrogates fibrillar collagen cleavage as already discussed, but has no effect on its ability to interact with peptide substrates or inhibitors. Association with TIMP-1, TIMP-2 or TIMP-3 is marginally decreased, suggesting that C-domain interactions are not of great significance [42]. This is of interest in the light of the observation that the activation of collagenase-3 mediated by cell-associated MT1-MMP requires the presence of the C-domain of collagenase-3 [42]. The mechanism underlying this observation remains to be elucidated.
Interactions of the MMPs with their specific natural inhibitors, the TIMPs, predominantly involve the enzyme catalytic domains and the N-terminal three disulfide-bonded loops of the TIMPs. Motifs within these two interacting domains determine a degree of specificity between individual MMPs and TIMPs but these are not well characterised. The C-domain interactions of certain MMPs with the three C-terminal disulfide-bonded loops of the TIMPs, however, do seem to play a major role in the acceleration of association rates in ways that may be biologically relevant.
We have focused our studies on the interactions of TIMPs with MT1-MMP (see earlier) because they are involved in the mechanism for MT1-MMP activation of progelatinase A, apparently bridging the two proteinases to form a tri-molecular complex. We found that one specific interaction between TIMP-2 and MT1-MMP could be demonstrated between the hairpin turn of the A and B β strands of TIMP-2 and MT1-MMP. Detailed site-directed mutagenesis of the AB loop showed the residue Tyr36 is almost wholly responsible for the binding of this region to MT1-MMP [43].
We also recently extended our observations to compare TIMP-2 with TIMP4. TIMP-2 and TIMP-4 have a number of similar characteristics with respect to sequence and kinetics of MMP-2 and MT1-MMP interaction, but the latter does not support MT1-MMP activation of progelatinase A. A series of TIMP-4 mutants were produced, including chimeric proteins with TIMP-2 and forms of the C-terminal charged tail that contained sequences from TIMP-2. In biochemical and cell-based studies, we found that TIMP-4 chimeras containing the C-domain of TIMP-2, and even TIMP-4 C-tail mutants containing elements of the TIMP-2 C-tail sequence (Arg186-Pro195), were able to support the formation of gelatinase A-MT1-MMP trimolecular complexes and to support gelatinase A activation (Knäuper et al., unpublished data).
In conclusion, MMPs have a domain structure to facilitate their interaction with activators, substrates and inhibitors. Specific binding to cell membrane or ECM components may also be determined by these domains. These are of importance to focus proteolytic activities at specific pericellular sites. For each MMP, a detailed analysis of the relationship between the function of its domains and their specific structure will give valuable new data to aid the design of future novel and specific antiproteinase molecules as potential therapeutics. The mutagenesis and kinetic studies that we have performed have indicated the extent of the task and, with the advent of all the relevant crystal structures, further directed studies may proceed.
MMP cleavage of ECM components
The MMP family contains the only vertebrate proteinases that can specifically degrade triple-helical collagens type I, type II and type III, as described earlier. Cleavage characteristically and specifically occurs at a single locus in all three collagen chains, at a point approximately three-quarters from the N-terminus of the molecule. At physiological temperature, the cleaved triple helix then unwinds, becoming susceptible to attack from other, less specific, proteinases.
The precise MMPs responsible for cartilage collagen cleavage in the arthritides is still open for debate. There are obvious differences in the turnover of collagen between OA and RA with respect to the location of early changes. This may suggest that a different MMP is predominantly responsible for collagen cleavage and that the involvement of different metalloproteinases is temporally distinct in each disease. Changes within the subchondral bone appear to precede cartilage changes in deep layers of cartilage, with subsequent fibrillation of the articular surface in OA. In contrast, early changes in RA appear in the surface layers with the majority of cartilage collagen destruction occurring at the cartilage-pannus junction, although changes within the cartilage matrix may precede and allow penetration by synovial cells.
Collagenase-1 and collagenase-3 are both made by chondrocytes (as are collagenase-2, gelatinase A and MT1-MMP), while collagenase-1 is also produced by the synovial fibroblasts. Hence, some workers consider collagenase-1 may be the foremost collagenase in RA, where the synovium proliferates to form a pannus that can invade cartilage, while collagenase-3 may predominate in OA where the chondrocyte drives the cartilage destruction [13, 20, 44]. Collagenase-2, previously thought to be restricted to neutrophils, has recently been described in chondrocytes [11] and in synovial cells [45], but a role in cartilage collagen destruction is still unproven [15]. MT1-MMP and gelatinase A (along with all the collagenases) have also been localised to the invading rheumatoid pannus, suggesting a possible role for either (or both) enzymes in disease [46].
MMPs and therapeutic strategies
MMP activity is controlled at several levels, of which gene expression is arguably the most important and is modulated by a variety of growth factors and cytokines [47]. A number of individual polymorphisms of MMP genes have recently been described with some disease associations [48], and associations with arthritic diseases may emerge in future research. After secretion, the activation described earlier (i.e. the proteolytic removal of the propeptide from the latent enzyme) represents one critical step in controlling MMP activity in the ECM. Once activated, regulation in the ECM is effected by natural inhibitors, such as the TIMPs and α2-macroglobulin [49]. The balance between MMP levels and TIMP levels is thought to be of great importance in determining levels of proteolysis.
Potential strategies for the modulation of MMPs include the development of effectors at the level of gene induction (cytokine inhibitors, receptor antagonists, signal transduction), production (transcription, secretion), zymogen activation (inhibit activating enzyme or prevent conversion of proactive to active enzyme) or enzyme activity (production of TIMPs or inhibition by small molecules). The potential of signal transduction inhibitors for the treatment of arthritis was recently discussed [50] and future directions in the treatment of OA at the level of MMPs has been summarised by Malemud and Goldberg [51].
The direct inhibition by synthetic low molecular weight entities that bind to the zinc and other features of the catalytic site of the enzyme has been most vigorously pursued [17, 52]. Early developments led to inhibitors of relatively poor specificity. However, the availability of crystal structures for a number of MMPs has allowed the design and production of inhibitors with increased specificity.
The issue of which MMP (or which other metalloproteinase, such as ADAM-TS4/5 with aggrecanase activity) to inhibit still remains. Van Meurs et al. found that active MMPs, particularly stromelysin-1, played a pivotal role in cartilage destruction in both the immune-mediated complex model of arthritis and in antigen-induced arthritis models. Absence of stromelysin-1 did not affect aggrecan depletion in arthritis but did prevent the appearance of collagen cleavage products and cartilage erosions [53, 54], suggesting a role for stromelysin-1 in the activation of procollagenases. Elliot and Cawston [55] and Clark et al. [22] discuss the issue of whether it is preferable to inhibit the degradation of the proteoglycan or the collagen component of cartilage as a therapeutic strategy for arthritis, but the question of the precise target enzymes remains. There is some support for the concept that the use of a broad-spectrum MMP inhibitor that also inhibits the ADAM proteinases (e.g. TACE, ADAM 17) would be most appropriate in arthritis therapy [55, 56].
MMP inhibitors have frequently exhibited toxicity in clinical trials, however, with the development of musculo-skeletal problems such as arthralgia, myalgia and tendinitis. These were predominantly in the upper limbs and were reversible. Interestingly, it has been shown that MT1-MMP-deficient mice not only develop dwarfism and osteopaenia, but also arthritic symptoms and soft tissue fibrosis, demonstrating the important function of this MMP in normal tissue remodelling and suggesting that it would be advantageous to selectively inhibit MMPs active within disease contexts [57].
There are currently no synthetic MMP inhibitors in clinical trials for arthritis due to the failure of early studies, for reasons such as those already outlined. A tetracycline derivative, doxycycline, in subantimicrobial doses (Periostat; CollaGenex Pharmaceuticals Inc., Newtown, PA, USA) is currently the only MMP inhibitor approved by the US Food and Drug Administration and is used as an adjunct therapy in adult periodontitis. The use of tetracyclines for the treatment of arthritic diseases is limited, although doxycycline has been shown to improve some disease parameters as well as reducing the levels of collagenase activity in some patients with RA [58, 59].
Concluding remarks
There are now 25 known MMPs, as detailed in this review, but many have not been substantially characterised in relation to arthritic diseases. It will be important for such studies to be carried out in relation to specific disease states. Although it is likely that the MMPs identified are major players, others could have subtle but critical activities in relation to the tissue cells and their interaction with their environment. A proteomics approach [60] should prove useful, if these techniques become sensitive enough, to document the range of proteinases expressed, followed by more detailed studies using quantitative RT-PCR in relation to well-documented disease status. Specific abrogation of individual MMPs in model systems may then give clues vital to the focusing of the design of antiMMP agents as potential therapies.
Glossary of terms
ADAM-TS = a disintegrin, a metalloproteinase and thrombospondin 'aggrecanase'; ECM = extracellular matrix; MMP = matrix metalloproteinase; MT-MMP = membrane-type matrix metalloproteinase; TIMP = tissue inhibitor of metalloproteinases.
References
Dayer JM, Krane SM, Russell RG, Robinson DR: Production of collagenase and prostaglandins by isolated adherent rheumatoid synovial cells. Proc Natl Acad Sci USA. 1976, 73: 945-949.
Sirum KL, Brinckerhoff CE: Cloning of the genes for human stromelysin and stromelysin 2: differential expression in rheumatoid synovial fibroblasts. Biochemistry. 1989, 28: 8691-8698.
Woolley DE, Evanson JM: Collagenase and its natural inhibitors in relation to the rheumatoid joint. Connect Tissue Res. 1977, 5: 31-35.
Hembry RM, Bagga MR, Reynolds JJ, Hamblen DL: Immuno-localisation studies on six matrix metalloproteinases and their inhibitors, TIMP-1 and TIMP-2, in synovia from patients with osteo- and rheumatoid arthritis. Ann Rheum Dis. 1995, 54: 25-32.
McCachren SS: Expression of metalloproteinases and metalloproteinase inhibitor in human arthritic synovium. Arthritis Rheum. 1991, 34: 1085-1092.
Walakovits LA, Moore VL, Bhardwaj N, Gallick GS, Lark MW: Detection of stromelysin and collagenase in synovial fluid from patients with rheumatoid arthritis and posttraumatic knee injury. Arthritis Rheum. 1992, 35: 35-42.
Lohmander LS, Hoerrner LA, Lark MW: Metalloproteinases, tissue inhibitor, and proteoglycan fragments in knee synovial fluid in human osteoarthritis. Arthritis Rheum. 1993, 36: 181-189.
Ishiguro N, Ito T, Oguchi T, Kojima T, Iwata H, Ionescu M, Poole AR: Relationships of matrix metalloproteinases and their inhibitors to cartilage proteoglycan and collagen turnover and inflammation as revealed by analyses of synovial fluids from patients with rheumatoid arthritis. Arthritis Rheum. 2001, 44: 2503-2511. 10.1002/1529-0131(200111)44:11<2503::AID-ART430>3.0.CO;2-P.
Wolfe GC, MacNaul KL, Buechel FF, McDonnell J, Hoerrner LA, Lark MW, Moore VL, Hutchinson NI: Differential in vivo expression of collagenase messenger RNA in synovium and cartilage. Arthritis Rheum. 1993, 36: 1540-1547.
Mohtai M, Smith RL, Schurman DJ, Tsuji Y, Torti FM, Hutchinson NI, Stetler-Stevenson WG, Goldberg GI: Expression of 92-kD type IV collagenase/gelatinase (gelatinase B) in osteoarthritic cartilage and its induction in normal human articular cartilage by interleukin 1. J Clin Invest. 1993, 92: 179-185.
Cole AA, Chubinskaya S, Schumacher B, Huch K, CS-Szabo G, Yao J, Mikecz K, Hasty KA, Kuettner KE: Chondrocyte matrix metalloproteinase-8 – Human articular chondrocytes express neutrophil collagenase. J Biol Chem. 1996, 271: 11023-11026. 10.1074/jbc.271.18.11023.
Moldovan F, Pelletier JP, Hambor J, Cloutier JM, Martel-Pelletier J: Collagenase-3 (matrix metalloprotease 13) is preferentially localized in the deep layer of human arthritic cartilage in situ – In vitro mimicking effect by transforming growth factor?. Arthritis Rheum. 1997, 40: 1653-1661.
Tetlow LC, Woolley DE: Comparative immunolocalization studies of collagenase 1 and collagenase 3 production in the rheumatoid lesion, and by human chondrocytes and synoviocytes in vitro. Br J Rheumatol. 1998, 37: 64-70. 10.1093/rheumatology/37.1.64.
Tetlow LC, Adlam DJ, Woolley DE: Matrix metalloproteinase and proinflammatory cytokine production by chondrocytes of human osteoarthritic cartilage: associations with degenerative changes. Arthritis Rheum. 2001, 44: 585-594. 10.1002/1529-0131(200103)44:3<585::AID-ANR107>3.3.CO;2-3.
Konttinen YT, Ainola M, Valleala H, Ma J, Ida H, Mandelin J, Kinne RW, Santavirta S, Sorsa T, Lopez-Otin C, Takagi M: Analysis of 16 different matrix metalloproteinases (MMP-1 to MMP-20) in the synovial membrane: different profiles in trauma and rheumatoid arthritis. Ann Rheum Dis. 1999, 58: 691-697.
Fosang AJ, Last K, Knauper V, Neame PJ, Murphy G, Hardingham TE, Tschesche H, Hamilton JA: Fibroblast and neutrophil collagenases cleave at two sites in the cartilage aggrecan interglobular domain. Biochem J. 1993, 295: 273-276.
Bottomley KM, Johnson WH, Walter DS: Matrix metalloproteinase inhibitors in arthritis. J Enzyme Inhib. 1998, 13: 79-101.
Kozaci LD, Brown CJ, Adcocks C, Galloway A, Hollander AP, Buttle DJ: Stromelysin 1, neutrophil collagenase, and collagenase 3 do not play major roles in a model of chondrocyte mediated cartilage breakdown. J Clin Pathol Mol Pathol. 1998, 51: 282-286.
Migita K, Eguchi K, Kawabe Y, Ichinose Y, Tsukada T, Aoyagi T, Nakamura H, Nagataki S: TNF-alpha-mediated expression of membrane-type matrix metalloproteinase in rheumatoid synovial fibroblasts. Immunology. 1996, 89: 553-557. 10.1046/j.1365-2567.1996.d01-789.x.
Billinghurst RC, Dahlberg L, Ionescu M, Reiner A, Bourne R, Rorabeck C, Mitchell P, Hambor J, Diekmann O, Tschesche H, Chen J, Van Wart H, Poole AR: Enhanced cleavage of type II collagen by collagenases in osteoarthritic articular cartilage. J Clin Invest. 1997, 99: 1534-1545.
Nagase H, Woessner JF: Matrix metalloproteinases. J Biol Chem. 1999, 274: 21491-21494. 10.1074/jbc.274.31.21491.
Clark IM, Rowan AD, Cawston TE: Matrix metalloproteinase inhibitors in the treatment of arthritis. Curr Opin Anti-Inflamm Immunomodulat Invest Drugs. 2000, 2: 16-25.
Vu TH, Werb Z: Matrix metalloproteinases: effectors of development and normal physiology. Genes Dev. 2000, 14: 2123-2133. 10.1101/gad.815400.
Birkedal-Hansen H: Proteolytic remodeling of extracellular matrix. COCB. 1995, 7: 728-735. 10.1016/0955-0674(95)80116-2.
MEROPS database. [http://www.Merops.ac.uk/Merops/index.htm]
Nagase H: Activation mechanisms of matrix metalloproteinases. Biol Chem. 1997, 378: 151-160.
Bode W, Huber R: Structural basis of the endoproteinase-protein inhibitor interaction. Biochim Biophys Acta. 2000, 1477: 241-252. 10.1016/S0167-4838(99)00276-9.
Knauper V, Patterson ML, Gomis-Ruth FX, Smith B, Lyons A, Docherty AJ, Murphy G: The role of exon 5 in fibroblast collagenase (MMP-1) substrate specificity and inhibitor selectivity. Eur J Biochem. 2001, 268: 1888-1896.
Knäuper V, Docherty AJP, Smith B, Tschesche H, Murphy G: Analysis of the contribution of the hinge region of human neutrophil collagenase (HNC, MMP-8) to stability and collagenolytic activity by alanine scanning mutagenesis. FEBS Lett. 1997, 405: 60-64. 10.1016/S0014-5793(97)00158-0.
Murphy G, Allan JA, Willenbrock F, Cockett MI, O'Connell JP, Docherty AJP: The role of the C-terminal domain in collagenase and stromelysin specificity. J Biol Chem. 1992, 267: 9612-9618.
Patterson ML, Atkinson SJ, Knauper V, Murphy G: Specific collagenolysis by gelatinase A, MMP-2, is determined by the hemopexin domain and not the fibronectin-like domain. FEBS Lett. 2001, 503: 158-162. 10.1016/S0014-5793(01)02723-5.
Murphy G, Nguyen Q, Cockett MI, Atkinson SJ, Allan JA, Knight CG, Willenbrock F, Docherty AJP: Assessment of the role of the fibronectin-like domain of gelatinase A by analysis of a deletion mutant. J Biol Chem. 1994, 269: 6632-6636.
Allan JA, Docherty AJP, Barker PJ, Huskisson NS, Reynolds JJ, Murphy G: Binding of gelatinases A and B to type-I collagen and other matrix components. Biochem J. 1995, 309: 299-306.
McQuibban GA, Gong JH, Tam EM, McCulloch CA, Clark-Lewis I, Overall CM: Inflammation dampened by gelatinase A cleavage of monocyte chemoattractant protein-3. Science. 2000, 289: 1202-1206. 10.1126/science.289.5482.1202.
O'Connell JP, Willenbrock F, Docherty AJP, Eaton D, Murphy G: Analysis of the role of the COOH-terminal domain in the activation, proteolytic activity, and tissue inhibitor of metalloproteinase interactions of gelatinase B. J Biol Chem. 1994, 269: 14967-14973.
Atkinson SJ, Crabbe T, Cowell S, Ward RV, Butler MJ, Sato H, Seiki M, Reynolds JJ, Murphy G: Intermolecular autolytic cleavage can contribute to the activation of progelatinase A by cell membranes. J Biol Chem. 1995, 270: 30479-30485. 10.1074/jbc.270.30.17797.
Will H, Atkinson SJ, Butler GS, Smith B, Murphy G: The soluble catalytic domain of membrane type 1 matrix metalloproteinase cleaves the propeptide of progelatinase A and initiates auto-proteolytic activation – Regulation by TIMP-2 and TIMP-3. J Biol Chem. 1996, 271: 17119-17123. 10.1074/jbc.271.29.17124.
Butler GS, Butler MJ, Atkinson SJ, Will H, Tamura T, Van Westrum SS, Crabbe T, Clements J, d'Ortho M-P, Murphy G: The TIMP2 membrane type 1 metalloproteinase 'receptor' regulates the concentration and efficient activation of progelatinase A. J Biol Chem. 1998, 273: 871-880. 10.1074/jbc.273.2.871.
Willenbrock F, Crabbe T, Slocombe PM, Sutton CW, Docherty AJP, Cockett MI, O'Shea M, Brocklehurst K, Phillips IR, Murphy G: The activity of the tissue inhibitors of metalloproteinases is regulated by C-terminal domain interactions: a kinetic analysis of the inhibition of gelatinase A. Biochemistry. 1993, 32: 4330-4337.
d'Ortho M-P, Will H, Atkinson S, Butler GS, Messent A, Gavrilovic J, Smith B, Timpl R, Zardi L, Murphy G: Membrane-type matrix metalloproteinases 1 and 2 exhibit broad-spectrum proteolytic capacities comparable to many matrix metalloproteinases. Eur J Biochem. 1997, 250: 751-757.
English WR, Holtz B, Vogt G, Knauper V, Murphy G: Characterization of the role of the 'MT-loop': an eight-amino acid insertion specific to progelatinase A (MMP2) activating membrane-type matrix metalloproteinases. J Biol Chem. 2001, 276: 42018-42026. 10.1074/jbc.M107783200.
Knäuper V, Murphy G: Membrane-type matrix metalloproteinases and cell surface-associated activation cascades for matrix metalloproteinases. In Matrix Metalloproteinases. Edited by: Edited by Parks WC, Mecham RP. 1998, San Diego, CA: Academic Press;, 199-218.
Williamson RA, Hutton M, Vogt G, Rapti M, Knauper V, Carr MD, Murphy G: Tyrosine 36 plays a critical role in the interaction of the AB loop of tissue inhibitor of metalloproteinases-2 with matrix metalloproteinase-14. J Biol Chem. 2001, 276: 32966-32970. 10.1074/jbc.M101843200.
Lindy O, Konttinen YT, Sorsa T, Ding YL, Santavirta S, Ceponis A, López-Otín C: Matrix metalloproteinase 13 (collagenase 3) in human rheumatoid synovium. Arthritis Rheum. 1997, 40: 1391-1399.
Hanemaaijer R, Sorsa T, Konttinen YT, Ding Y, Sutinen M, Visser H, van Hinsbergh VW, Helaakoski T, Kainulainen T, Ronka H, Tschesche H, Salo T: Matrix metalloproteinase-8 is expressed in rheumatoid synovial fibroblasts and endothelial cells. Regulation by tumor necrosis factor-alpha and doxycycline. J Biol Chem. 1997, 272: 31504-31509. 10.1074/jbc.272.50.31504.
Konttinen YT, Ceponis A, Takagi M, Ainola M, Sorsa T, Sutinen ME, Salo T, Ma J, Santavirta S, Seiki M: New collagenolytic enzymes cascade identified at the pannus-hard tissue junction in rheumatoid arthritis: Destruction from above. Matrix Biol. 1998, 17: 585-601. 10.1016/S0945-053X(98)90110-X.
Borden P, Heller RA: Transcriptional control of matrix metalloproteinases and the tissue inhibitors of matrix metalloproteinases. Crit Rev Eukaryot Gene Expr. 1997, 7: 159-178.
Ye S, Henney AM: Detecting polymorphisms in MMP genes. Methods Mol Biol. 2001, 151: 367-375.
Nagase H, Brew K: Engineering of tissue inhibitor of metalloproteinases mutants as potential therapeutics. Arthritis Res. 2002, 4 (suppl 3): S51-S61. 10.1186/ar573.
Vincenti MP, Brinckerhoff CE: Early response genes induced in chondrocytes stimulated with the inflammatory cytokine interleukin-1beta. Arthritis Res. 2001, 3: 381-388. 10.1186/ar331.
Malemud CJ, Goldberg VM: Future directions for research and treatment of osteoarthritis. Front Biosci. 1999, 4: D762-D771.
Skotnicki JS, Zask A, Nelson FC, Albright JD, Levin JI: Design and synthetic considerations of matrix metalloproteinase inhibitors. Ann NY Acad Sci. 1999, 878: 61-72.
van Meurs J, van Lent P, Stoop R, Holthuysen A, Singer I, Bayne E, Mudgett J, Poole R, Billinghurst C, van der Kraan P, Buma P, van den Berg W: Cleavage of aggrecan at the ASN341-PHE342 site coincides with the initiation of collagen damage in murine antigen-induced arthritis. A pivotal role for stromelysin 1 in matrix metalloproteinase activity. Arthritis Rheum. 1999, 42: 2074-2084. 10.1002/1529-0131(199910)42:10<2074::AID-ANR7>3.0.CO;2-5.
van Meurs J, van Lent P, Holthuysen A, Lambrou D, Bayne E, Singer I, van den Berg W: Active matrix metalloproteinases are present in cartilage during immune complex-mediated arthritis: a pivotal role for Stromelysin-1 in cartilage destruction. J Immunol. 1999, 163: 5633-5639.
Elliott S, Cawston T: The clinical potential of matrix metalloproteinase inhibitors in the rheumatic disorders. Drugs Aging. 2001, 18: 87-99.
Conway JG, Andrews RC, Beaudet B, Bickett DM, Boncek V, Brodie TA, Clark RL, Crumrine C, Leenitzer MA, McDougald DL, Han B, Hedeen K, Lin P, Milla M, Moss M, Pink H, Rabinowitz MH, Tippin T, Scates P, Selph J, Stimpson SA, Warner J, Becherer JD: Inhibition of tumor necrosis factor-α (TNF-α) production and arthritis in the rat by GW a dual inhibitor of TNF-α-converting enzyme and matrix metalloproteinases. J Pharm Exp Ther. 3333, 298: 900-908.
Holmbeck K, Bianco P, Caterina J, Yamada S, Kromer M, Kuznetsov SA, Mankani M, Robey PG, Poole AR, Pidoux I, Ward JM, Birkedal-Hansen H: MT1-MMP-deficient mice develop dwarfism, osteopenia, arthritis and connective tissue disease due to inadequate collagen turnover. PubMed Cell. 1999, 1: 81-92.
Nordstrom D, Lindy O, Lauhio A, Sorsa T, Santavirta S, Konttinen YT: Anti-collagenolytic mechanism of action of doxycycline treatment in rheumatoid arthritis. Rheumatol Int. 1998, 17: 175-180. 10.1007/s002960050030.
O'Dell JR: Is there a role for antibiotics in the treatment of patients with rheumatoid arthritis?. Drugs. 1999, 57: 279-282.
Kumar S, Connor JR, Dodds RA, Halsey W, Van Horn M, Mao J, Sathe G, Mui P, Agarwal P, Badger AM, Lee JC, Gowen M, Lark MW: Identification and initial characterisation of 5000 expressed sequence tags (ESTs) each from adult normal and osteoarthritic cartilage cDNA libraries. Ostoearthritis Cartilage. 2001, 9: 641-653. 10.1053/joca.2001.0421.
Gomis-Rüth FX, Maskos K, Betz M, Bergner A, Huber R, Suzuki K, Yoshida N, Nagase H, Brew K, Bourenko GP, Bartunik H, Bode W: Mechanism of inhibition of the human matrix metalloproteinase stromelysin-1 by TIMP-1. Nature. 1997, 389: 77-81. 10.1038/37995.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Murphy, G., Knäuper, V., Atkinson, S. et al. Matrix metalloproteinases in arthritic disease. Arthritis Res Ther 4 (Suppl 3), S39 (2002). https://doi.org/10.1186/ar572
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1186/ar572