Abstract
In this paper, the Lie algebraic method is applied to solve biological population models described by time-inhomogeneous birth-death processes. Notwithstanding no obvious symmetry, the solution is expressed by matrix exponentials through suitably generated low-dimensional Lie algebras. This methodology may offer useful insights for other biological and ecological applications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
During the past few decades, Markovian stochastic systems have come to play a vital role in a host of branches of science and engineering applications. Biological population models, due to the random nature of diffusion of populations, are often captured by continuous-time Markovian models [1–7]. For example, the Moran process [8], which describes the probabilistic dynamics in a finite population in which two alleles A1 and A2 are competing for dominance, can be suitably described as a birth-death process. In general, stochastic effects on populations change over time, which give rise to time-inhomogeneous behavior, increasing the complexity of population dynamics.
An effective and often easy-to-use method for solving time-inhomogeneous Markov chains was proposed in [9] by using low-dimensional Lie algebras. This method (which we will review below) has broad applications in physical and chemical sciences (see, e.g., [10–13]), and certain symmetries of the systems are used as a guide to generate an appropriate Lie algebra. Recently, the Lie algebraic method was applied to solve biological population models [14, 15] where symmetry is insufficient.
In this paper, we implement the Lie algebraic method to a biological population model to find analytical solutions through matrix exponentials. The population model considered is not obviously symmetric and is described by time-inhomogeneous birth-death processes. Our result generalizes a previous model in [14].
Lie algebraic methodology
A Lie algebra [16] is a vector space V over some field F together with a bilinear operation [ ·,·]:V×V→V called the Lie bracket, which obeys [ X,X] =0 and the Jacobi identity
for all X,Y,Z∈V. For X∈V, define a linear operator adX by
for Y∈V. Thus, multiple Lie brackets can be expressed in a compact way, e.g., (adX)2Y= [ X,[ X,Y] ], etc. Let be the general linear group of order n over  , where n can be finite or infinite. For two matrices , define
, where n can be finite or infinite. For two matrices , define
as the commutator (or Lie bracket) of X and Y. Therefore, the classical Baker-Campbell-Hausdorff formula [17] can be rewritten as
The type of processes considered here are continuous-time Markov chains [18], taking values in the state space . The dynamical behavior of the Markov chain is governed by a matrix Q(t)=(q ij (t),i,j∈S), where q ij (t) is the rate of transition from state i to state j, for j≠i, and is the total rate at which we leave state i at time t. By employing the Kolmogorov forward equation, the probability distribution of the process at time t, p(t)=(p i (t),i∈S), is given by
where H(t)=Q(t)T (T means transpose), and p(t) is a column probability vector with element p i (t) representing the probability of finding the system in state i at time t. H(t) is time-dependent, meaning that the process is time-inhomogeneous.
The Lie algebraic methodology proposed in [9] first requires a decomposition of the matrix H(t) as
such that a i (t) are real-valued functions, and H i (i=1,⋯,m) are linearly independent constant matrices generating a Lie algebra by implementing a Lie bracket
for . It was shown that the solution of system (5) can be uncoupled into a product of exponentials [9]
where g i (t) are real-valued functions and g i (0)=0.
Feeding (6) and (8) into (5), we obtain
Since by using (4), we have
On the other hand, multiplying U(t)-1 on the right of (9) yields
Combining (10), (11), and (9), we obtain
since p(0) is arbitrary.
Notice that the matrices H i are linearly independent, and thus, the exact solution to (12) is reduced to that of a linear system between a i (t) and , involving η ijk , with initial values g i (0)=0. A remarkable advantage of this method lies in reducing computational complexity: the calculation of p(t) can be achieved in O(1) through (12) rather than O(t) through incremental direct integrations. Besides, the matrix exponential form (8) would be useful if the derivative of the solution with respect to a model parameter is required [13].
Application on a population model
The population model considered here is described by a time-inhomogeneous birth-death process N(t) taking values in . The system goes from state i to i+1 with birth rate b(t)≥0, while it goes from state i to i-1 with death rate f(i)d(t)≥0. For technical reasons, we will assume that
for some and f(0)=0. Let be the generating function of the sequence f(i); it is easy to check that
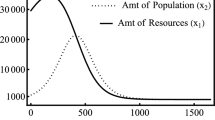
This population model could describe the survival of juvenile animals dying at a rate that depends on climate and some regularly varied resource when introduced to an inhospitable region by seasonal breeding happening at another site [2, 14].
Let p i (t)=P(N(t)=i) for t≥0. The Kolmogorov equation governing this process can be written as
where H(t)=Q(t)T=
In terms of the Kronecker delta, we expand H(t) as
where (R) ij =δi,j+1, (I) ij =δi,j, (L) ij =f(j-1)δi,j-1, and (M) ij =f(j-1)δi,j. It is necessary to include another matrix J with (J) ij =(f(i)-f(j-1))δi,j to have an algebra that is closed under the action of the Lie bracket. In Table 1, we show the complete set of Lie brackets.
We will look for a solution of the form
By using (12) and the action of the exponential operator shown in Table 2, we can derive
Equating terms in (19) in front of the same base matrices yields
where , and g2(t) is determined by the initial value problem
To derive g2, set in (21), and then we obtain
This is a Riccati equation, which can be solved in some situations by reduction techniques (see, e.g., [19]).
When c=0 and f(1)=1, the recursive relation (13) gives f(n)=n for all . This is the example studied in [14, Section 3.1]. The solution of (19) can be obtained as
where . It is easy to check that (18) together with (23) agrees with the solution derived in [14].
Conclusions
In the past few years, stochastic modeling has received much attention, and it is widely used to study different types of dynamical systems subject to abrupt changes in their structure, such as failure-prone manufacturing systems, neural networks, power systems, economics systems, etc. In this paper, we employed the Lie algebraic methodology to find the analytical solutions of biological population models described by time-inhomogeneous birth-death processes. The Lie algebra was shown to be a very powerful and efficient approach in finding analytical solutions for numerous physical systems but has not been widely used in the context of biological populations due to insufficient symmetries. It is hoped that the technique described in this paper will find applications in broad classes of biological and ecological models.
References
Alonso D, McKane A: Extinction dynamics in mainland-island metapopulations: an n-patch stochastic model. Bull. Math. Biol 2002, 64: 913–958. 10.1006/bulm.2002.0307
Brauer F, Castillo-Chávez C: Mathematical Models in, Population Biology and Epidemiology. New York: Springer; 2010.
Cooper B, Lipsitch M: The analysis of hospital infection data using hidden Markov models. Biostatistics 2004, 5: 223–237. 10.1093/biostatistics/5.2.223
Keeling MJ, Ross JV: Efficient methods for studying stochastic disease and population dynamics. Theor. Popul. Biol 2009, 75: 133–141. 10.1016/j.tpb.2009.01.003
Ross JV: A stochastic metapopulation model accounting for habitat dynamics. J. Math. Biol. 2006, 52: 788–806. 10.1007/s00285-006-0372-8
Shang Y: Likelihood estimation for stochastic epidemics with heterogeneous mixing populations. World Acad. Sci. Eng. Technol. 2011, 55: 929–933.
Shang Y: The limit behavior of a stochastic logistic model with individual time-dependent rates. J. Math 2013, 2013: 502635.
Moranx P: The Statistical, Processes of Evolutionary Theory. Oxford: Clarendon Press; 1962.
Wei J, Norman E: Lie algebra solution of linear differential equations. J. Math. Phys. 1963, 4: 575–581. 10.1063/1.1703993
Alvermann A, Fehske H: High-order commutator-free exponential time-propagation of driven quantum systems. J. Comput. Phys. 2011, 230: 5930–5956. 10.1016/j.jcp.2011.04.006
Blanes S, Casas F, Oteo JA, Ros J: The magnus expansion and some of its applications. Phys. Rep 2009, 470: 151–238. 10.1016/j.physrep.2008.11.001
Fernández F, Castro EA: Algebraic Methods in Quantum Chemistry and Physics. CRC, Boca Raton 1996.
Wilcox RM: Exponential operators and parameter differentiation in quantum physics. J. Math. Phys. 1967, 8: 962–982. 10.1063/1.1705306
House T: Lie algebra solution of population models based on time-inhomogeneous Markov chains. J. Appl. Probab 2012, 49: 472–481. 10.1239/jap/1339878799
Shang Y: A lie algebra approach to susceptible-infected-susceptible epidemics. Electron. J. Differ. Equ. 2012 2012, 233.
Humphreys JE: Introduction to Lie Algebras and Representation Theory. New York: Springer; 1972.
Campbell JE: On a law of combination of operators (second paper). Proc. Lond. Math. Soc. 1897, 28: 381–390.
Norris JR: Markov Chains. New York: Cambridge University Press; 1997.
Polyanin AD, Zaitsev VF: Handbook of Exact Solutions for Ordinary Differential Equations, 2nd edition. Boca Raton: CRC; 2003.
Acknowledgements
The author would like to thank the learned referees for their valuable suggestions.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The author declares that he has no competing interests.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Shang, Y. Lie algebra method for solving biological population model. J Theor Appl Phys 7, 67 (2013). https://doi.org/10.1186/2251-7235-7-67
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/2251-7235-7-67