Abstract
Background
Rapidly accumulating genome sequence data from multiple species offer powerful opportunities for the detection of DNA sequence evolution. Phylogenetic tree construction and codon-based tests for natural selection are the prevailing tools used to detect functionally important evolutionary change in protein coding sequences. These analyses often require multiple DNA sequence alignments that maintain the correct reading frame for each collection of putative orthologous sequences. Since this feature is not available in most alignment tools, codon reading frames often must be checked manually before evolutionary analyses can commence.
Results
Here we report an online codon-preserved alignment tool (OCPAT) that generates multiple sequence alignments automatically from the coding sequences of any list of human gene IDs and their putative orthologs from genomes of other vertebrate tetrapods. OCPAT is programmed to extract putative orthologous genes from genomes and to align the orthologs with the reading frame maintained in all species. OCPAT also optimizes the alignment by trimming the most variable alignment regions at the 5' and 3' ends of each gene. The resulting output of alignments is returned in several formats, which facilitates further molecular evolutionary analyses by appropriate available software. Alignments are generally robust and reliable, retaining the correct reading frame. The tool can serve as the first step for comparative genomic analyses of protein-coding gene sequences including phylogenetic tree reconstruction and detection of natural selection. We aligned 20,658 human RefSeq mRNAs using OCPAT. Most alignments are missing sequence(s) from at least one species; however, functional annotation clustering of the ~1700 transcripts that were alignable to all species shows that genes involved in multi-subunit protein complexes are highly conserved.
Conclusion
The OCPAT program facilitates large-scale evolutionary and phylogenetic analyses of entire biological processes, pathways, and diseases.
Similar content being viewed by others
Background
Multi-species comparisons offer a powerful way to identify functionally important DNA elements that are associated with the evolution of human phenotypes (e.g., the expanded neocortex, language production, and bipedal gait) and diseases that occur mostly in humans (e.g., pre-eclampsia) [1]. Rapidly accumulating whole genome sequence data from vertebrate species provide unprecedented opportunities for evolutionary analyses of protein coding genes. A necessary step in such analyses is the construction of in-frame multiple sequence alignments. Commonly used alignment tools such as CLUSTAL and T-COFFEE [2, 3] do not retain reading frame information, thus the achievement of in-frame alignments usually requires manual curation, which is impractical at a genome-wide scale. Moreover, genomic tools such as threaded blockset aligners [4] derive the reading frame from a single species and allow the others to have frame-shifts, which can affect downstream calculations of DNA substitution rates that are based on codon models. Therefore, none of these tools is wholly appropriate for phylogenetic analyses based on protein-coding models of sequence evolution [5]. To infer non-synonymous and synonymous substitutions for a large gene set, tools that automate codon-preserved alignments are required.
To address these issues, we developed a tool to automate gene alignments on a genome-wide scale with the reading-frame preserved for each set of putatively orthologous coding sequences. The tool is called OCPAT (Online Codon-Preserved Alignment Tool).
Implementation
The OCPAT pipeline is composed of 1) a user interface [6], 2) a CGI program to handle queries, 3) a genomic database to store sequences, and 4) the main program to generate alignments. Output is stored on a server and users are notified through email of URLs containing their results.
The current version of OCPAT aligns genes from Homo sapiens (human) [7], Pan troglodytes (chimpanzee) [8], Macaca mulatta (Rhesus macaque) [9], Mus musculus (mouse) [10], Rattus norvegicus (rat) [11], Oryctolagus cuniculus (rabbit), Canis familiaris (dog) [12], Bos taurus (cow), Dasypus novemcinctus (armadillo), Loxodonta africana (elephant), Echinops telfairi (tenrec), Monodelphis domestica (opossum) [13], Ornithorhynchus anatinus (platypus), Gallus gallus (chicken) [14], and Xenopus tropicalis (frog). mRNA and/or cDNA files are downloaded from the RefSeq mRNA databases [15], the ENSEMBL cDNA databases [16], and the NR (Non-redundant) mRNAs [17]. mRNA/cDNA sequences are then sorted by species and formatted and indexed using the "formatdb" program [18]. The GenBank formatted human mRNA and protein sequences are downloaded from RefSeq as well [19]. For the analysis described, data were updated on November 2, 2006.
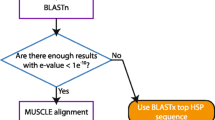
The procedure for obtaining and aligning sequences is executed using multiple available tools linked to one another through perl modules and scripts [see additional file 1]. OCPAT implements the following steps (Fig. 1):
Workflow of OCPAT. The pipeline implements the steps as shown in the figure. The human RefSeq mRNA serves as the initial query by default. Putative orthologs are defined by sequence similarity and gene symbol. Frameshift causing substitutions are masked, and OCPAT does not distinguish true frameshifts from sequencing errors. Unlike other alignment tools, a RefSeq golden peptide sequence record guides OCPAT alignments rather than the predicted amino acid sequence derived from the mRNA record.
-
1.
Submission: Submits a list of human RefSeq IDs (e.g., NM_002425, NM_181814) in single-column format. Users can choose which (or all) of 14 additional taxa will be included in alignments.
-
2.
Ortholog extraction: Retrieves mRNAs or CDs from the other species by BLAST [20] search of respective cDNA or mRNA databases using the human CDs as the query. In the case of multiple sequences from one organism showing high similarity to the human CDs (i.e. less than 5% difference between paralogs in the nonhuman organism), the annotations (e.g. gene symbol) of those genes are used to choose the putative ortholog. OCPAT defines putative orthologs solely by sequence similarity. OCPAT also measures sequence concordance, which is a measure of the relative proportions of subject to query alignment length. Sequence concordance is calculated where Concordance = 2 * matched sequence length/(query sequence length + aligned subject sequence length). Aligned sequences shorter than half the length of the queried human sequence are eliminated from further consideration.
-
3.
Error correction: Pre-aligns sequences using CLUSTALW [3]. Searches these alignments for possible places where only one or two sequences has a frame shift introduced by nucleotide insertions or deletions while all the other sequences are perfectly aligned to each other. Indels causing gaps in these multiple sequence alignments are filled with "N"s so the subsequent translation does not cause frame shifting.
-
4.
Determination of reading frame: Obtains a human peptide for the queried RefSeq directly from the human.rna.gbff file. Translates putative orthologous gene sequences into peptides and determines the reading frame of each gene by aligning the translated sequence to the human peptide using the bl2seq program [21]. An "X" is used for amino acids derived from codons containing ambiguous nucleotides. Each sequence is then trimmed so codons correctly begin with the first nucleotide position. Peptides for the orthologous gene sequences are aligned using the CLUSTALW program [3]. Aligned peptides are then "translated" back to their corresponding cDNA sequences by sequence mapping in which the "reverse translation" is directed by correlated CDS and peptide positions (e.g., the Nth amino acid in the peptide maps back to the 3N-2, 3N-1, 3N nucleotides in the CDS), thus avoiding the problem of codon degeneracy. This translation, alignment and "reverse translation" procedure generates alignments that preserve the codon reading frames.
-
5.
Core alignment: Evaluates the cDNA alignment for the core alignment region, in which the suboptimal alignments at the beginning and end of genes (often due to poor predictions or sequence errors) are removed. A sliding window of three consecutive amino acids, beginning from the 5' end, is moved across the multiple sequence alignment. The "identical count" is determined by calculating the number of identical amino acids at each position in a three-amino-acid window. For a multiple sequence alignment of N sequences, the maximum "identical count" per window is 3N. When the "identical count" reaches 2.2N, the first amino acid in the sliding window is marked as the start point of the core alignment. This represents slightly over 70% identity in the alignments. The same sliding window strategy trims the 3' end of the core alignment. Large, single species insertions are also removed from the core alignments. The remaining "core alignments" always begin with the first nucleotide within a codon and end at the third nucleotide within a codon
-
6.
Output: Produces NEXUS-, PHYLIP-, and CLUSTAL- formatted files, which can be utilized by a variety of phylogenetic programs including PAUP*, MacClade, PAML, and Mr.Bayes [5, 22–24]. Additional output files include standard error files and a summary file (ocpat.align.sum).
Results
Using OCPAT we generated 20,658 multiple sequence alignments derived from human mRNA RefSeq IDs. Among these alignments 10,258 included 10 or more species. The pairwise numbers of alignable putative orthologous sequences is shown in Table 1, and a recent version of alignment files for putative orthologous sequences are available at [25]. All putative orthologs are considered provisional, and certainly there are some non-orthologous sequences included in individual gene alignments due to genome assembly errors, lineage specific gene duplications, and ascertainment errors. As expected, we obtained a greater number of putative human orthologs from species more closely related to human (e.g., chimpanzee, macaque) than from more distantly related species (e.g., chicken, frog). We also found that mammal species whose genomes were sequenced at 2-fold coverage had fewer recovered orthologs than did mammals with higher quality sequences. Despite these limitations, there are 1,698 human RefSeqs for which we were able to obtain putative orthologs from all taxa queried (N = 13; the platypus Genebuild was not available as of Nov. 2, 2006). Phylogenetic analyses have been conducted on these genes using parsimony, distance, likelihood, and Bayesian methods [26].
To explore the biological significance of the genes found in all species we conducted a functional annotation clustering analysis using the default settings of the DAVID package [27]. The results of this analysis indicated a statistically significant over-representation of genes that encode proteins found in multi-subunit complexes (n = 263 RefSeqs; p = 5.0E-39). Other overrepresented annotations in functional clusters include one comprised of ribosomal proteins and one containing proteins in the histone core. We consider the genes with putative orthologs for all species to be a good indicator of conservation (i.e., more identifiable orthologs indicates more functional constraint on the protein). Taken together, these results suggest that protein-protein interactions in multi-subunit complexes are under considerable evolutionary constraint. Therefore, mutations in these proteins are possibly more likely to be harmful when they occur.
Discussion
In silico gene prediction algorithms often fail at the 5' and 3' ends of a gene. Consequently, the 5'end and the 3'end of the predicted ORFs are error prone. This can lead to low-quality alignments in the 5' ends and the 3' ends of a given gene. OCPAT trims the low-quality alignment regions at the ends. The remaining high-quality core alignments in the middle of the gene may be less "noisy" than whole alignments. We also found that many of the genes predicted for opossum, chicken, and frog have large insertions when compared to genes from placental mammals. Therefore, if only one species has a big insertion while all the others do not, OCPAT removes the insertion. This treatment is most effective when there are other sequences that partially overlap the insertion. By removing these large insertions, the smaller overlapping regions are not "lost" as alignment gaps in subsequent phylogenetic analyses. If, after the initial run, the user finds the inclusion of one species disrupts the alignment due to factors such as poor gene prediction and short length, the user can re-run OCPAT for that gene and eliminate any disrupting sequences.
Conclusion
In summary, we provide a simple tool for aligning genes with the protein coding frames preserved. Alignments are formatted so they can be applied to evolutionary analyses using appropriate software. The tool is effective for creating alignments on a genome-wide scale. Future versions of OCPAT will use the genome sequences of additional species.
Availability and requirements
Project name: OCPAT
Project home page: http://homopan.med.wayne.edu/pise/ocpat.html.
Operating system(s): Mac OS X or Solaris 9/10; web server version is platform independent
Programming language: Perl
Other requirements: for the command line; NCBI BLAST utility, CLUSTAL; genome data
License: GNU General Public License
Any restrictions to use by non-academics: None
Abbreviations
- OCPAT:
-
Online Codon Preserved Alignment Tool.
References
Goodman M, Grossman LI, Wildman DE: Moving primate genomics beyond the chimpanzee genome. Trends Genet. 2005, 21 (9): 511-517. 10.1016/j.tig.2005.06.012.
Notredame C, Higgins DG, Heringa J: T-Coffee: A novel method for fast and accurate multiple sequence alignment. Journal of molecular biology. 2000, 302 (1): 205-217. 10.1006/jmbi.2000.4042.
Thompson JD, Higgins DG, Gibson TJ: CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic acids research. 1994, 22 (22): 4673-4680. 10.1093/nar/22.22.4673.
Blanchette M, Kent WJ, Riemer C, Elnitski L, Smit AF, Roskin KM, Baertsch R, Rosenbloom K, Clawson H, Green ED, et al.: Aligning multiple genomic sequences with the threaded blockset aligner. Genome research. 2004, 14 (4): 708-715. 10.1101/gr.1933104.
Yang Z: PAML: a program package for phylogenetic analysis by maximum likelihood. Comput Appl Biosci. 1997, 13 (5): 555-556.
Letondal C: A Web interface generator for molecular biology programs in Unix. Bioinformatics. 2001, 17 (1): 73-82. 10.1093/bioinformatics/17.1.73.
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, Devon K, Dewar K, Doyle M, FitzHugh W, et al.: Initial sequencing and analysis of the human genome. Nature. 2001, 409 (6822): 860-921. 10.1038/35057062.
The Chimpanzee Sequencing and Analysis Consortium: Initial sequence of the chimpanzee genome and comparison with the human genome. Nature. 2005, 437 (7055): 69-87. 10.1038/nature04072.
Gibbs RA, Rogers J, Katze MG, Bumgarner R, Weinstock GM, Mardis ER, Remington KA, Strausberg RL, Venter JC, Wilson RK, et al.: Evolutionary and biomedical insights from the rhesus macaque genome. Science. 2007, 316 (5822): 222-234. 10.1126/science.1139247.
Waterston RH, Lindblad-Toh K, Birney E, Rogers J, Abril JF, Agarwal P, Agarwala R, Ainscough R, Alexandersson M, An P, et al.: Initial sequencing and comparative analysis of the mouse genome. Nature. 2002, 420 (6915): 520-562. 10.1038/nature01262.
Gibbs RA, Weinstock GM, Metzker ML, Muzny DM, Sodergren EJ, Scherer S, Scott G, Steffen D, Worley KC, Burch PE, et al.: Genome sequence of the Brown Norway rat yields insights into mammalian evolution. Nature. 2004, 428 (6982): 493-521. 10.1038/nature02426.
Lindblad-Toh K, Wade CM, Mikkelsen TS, Karlsson EK, Jaffe DB, Kamal M, Clamp M, Chang JL, Kulbokas EJ, Zody MC, et al.: Genome sequence, comparative analysis and haplotype structure of the domestic dog. Nature. 2005, 438 (7069): 803-819. 10.1038/nature04338.
Mikkelsen TS, Wakefield MJ, Aken B, Amemiya CT, Chang JL, Duke S, Garber M, Gentles AJ, Goodstadt L, Heger A, et al.: Genome of the marsupial Monodelphis domestica reveals innovation in non-coding sequences. Nature. 2007, 447 (7141): 167-177. 10.1038/nature05805.
International Chicken Genome Sequencing Consortium: Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution. Nature. 2004, 432 (7018): 695-716. 10.1038/nature03154.
RefSeq mRNA databases. [ftp://ftp.ncbi.nih.gov/refseq/]
Ensembl. [ftp://ftp.ensembl.org/pub/release-41/]
FASTA nr.gz. [ftp://ftp.ncbi.nih.gov/blast/db/FASTA/]
BLAST Executables. [ftp://ftp.ncbi.nih.gov/blast/executables/release/2.2.13/]
Human RefSeq database. [ftp://ftp.ncbi.nih.gov/refseq/H_sapiens/mRNA_Prot/]
Ye J, McGinnis S, Madden TL: BLAST: improvements for better sequence analysis. Nucleic acids research. 2006, W6-9. 10.1093/nar/gkl164. 34 Web Server
Tatusova TA, Madden TL: BLAST 2 Sequences, a new tool for comparing protein and nucleotide sequences. FEMS microbiology letters. 1999, 174 (2): 247-250. 10.1111/j.1574-6968.1999.tb13575.x.
Maddison DR, Maddison WP: MacClade 4: Analysis of Phylogeny and Character Evolution. 2000, Sunderland, MA: Sinauer
Ronquist F, Huelsenbeck JP: MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics. 2003, 19 (12): 1572-1574. 10.1093/bioinformatics/btg180.
Swofford DL: PAUP*. Phylogenetic analysis using parsimony (*and other methods. 2002, Sunderland, MA: Sinauer
OCPAT All. [http://homopan.wayne.edu/OCPAT_withPlatypus/]
Wildman DE, Uddin M, Opazo JC, Liu G, Lefort V, Guindon S, Gascuel O, Grossman LI, Romero R, Goodman M: Genomics, biogeography, and the diversification of placental mammals. Proc Natl Acad Sci USA. 2007, 104 (36): 14395-14400. 10.1073/pnas.0704342104.
Dennis G, Sherman BT, Hosack DA, Yang J, Gao W, Lane HC, Lempicki RA: DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome biology. 2003, 4 (5): P3-10.1186/gb-2003-4-5-p3.
Acknowledgements
We thank Dr. Juan C. Opazo (University of Nebraska) for his helpful discussions and suggestions. This work was supported in part by the Intramural Research Division of the National Institute of Child Health and Human Development National Institutes of Health, Department of Health and Human Services. The authors would like to acknowledge the sources of unpublished genome sequence data including: Baylor College of Medicine Human Genome Sequencing Center (cow) http://www.hgsc.bcm.tmc.edu/projects/; the Broad Institute (rabbit, elephant, tenrec, armadillo); and the U.S. DOE Joint Genome Institute (JGI) (frog).
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The author(s) declare that they have no competing interests.
Authors' contributions
All authors have read and approved the final manuscript. DEW defined the problem and designed the project. GL and MI wrote the code and implemented OCPAT. MU, GL, MI, and DEW tested and debugged the programs. All authors participated in the manuscript preparation.
Electronic supplementary material
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
Open Access This article is published under license to BioMed Central Ltd. This is an Open Access article is distributed under the terms of the Creative Commons Attribution License ( https://creativecommons.org/licenses/by/2.0 ), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Liu, G., Uddin, M., Islam, M. et al. OCPAT: an online codon-preserved alignment tool for evolutionary genomic analysis of protein coding sequences. Source Code Biol Med 2, 5 (2007). https://doi.org/10.1186/1751-0473-2-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1751-0473-2-5