Abstract
Background
Intracellular membrane fusion processes are mediated by the spatial and temporal control of SNARE complex assembly that results in the formation of a four-helical bundle, composed of one vesicle SNARE and three target membrane SNARE polypeptide chains. Syntaxins are essential t-SNAREs and are characterized by an N-terminal Habc domain, a flexible linker region, a coiled-coil or SNARE motif and a membrane anchor. The N-terminal Habc domain fulfills important regulatory functions while the coiled-coil motif, present in all SNAREs, is sufficient for SNARE complex formation, which is thought to drive membrane fusion.
Results
Here we report the crystal structure of the Habc domain of neuronal syntaxin from the squid Loligo pealei, s-syntaxin. Squid Habc crystallizes as a dimer and the monomer structure consists of a three-helical bundle. One molecule is strikingly similar to mammalian syntaxin 1A while the second one shows a structural deviation from the common fold in that the C-terminal part of helix C unwinds and adopts an extended conformation.
Conclusion
Conservation of surface residues indicates that the cytosolic part of s-syntaxin can adopt an auto-inhibitory closed conformation that may bind squid neuronal Sec1, s-Sec1, in the same manner as observed in structure of the rat nSec1/syntaxin 1A complex. Furthermore, despite the overall structural similarity, the observed changes at the C-terminus of one molecule indicate structural plasticity in neuronal syntaxin. Implications of the structural conservation and the changes are discussed with respect to potential Habc domain binding partners such as Munc13, which facilitates the transition from the closed to the open conformation.
Similar content being viewed by others
Background
SNARE (soluble NSF attachment protein receptor) proteins are an evolutionary conserved family of membrane-anchored or membrane-associated proteins, which contain a conserved 60 amino acid motif of hydrophobic heptad repeats typical for coiled-coil interactions [1]. Complexes formed from cognate SNARE motifs located on opposing membranes physically link membranes prior to membrane fusion. Specifically, three SNARE chains from the target membrane and one chain from the vesicle align in a parallel four-helical bundle [2]. In neurotransmission, the vesicle SNARE synaptobrevin (also VAMP) forms a highly stable complex with the synaptic membrane proteins syntaxin 1A and SNAP-25, which contains two SNARE motifs [3, 4]. Deletion or ablation of any of these proteins resulted in a complete block in Ca2+-evoked neurotransmission [5–8]. Although trans-SNARE complex formation is necessary and sufficient for in vitro vesicle fusion [9, 10], studies with neurotoxins like Botulinum and Tetanus toxin, which block neurotransmission by proteolytically cleaving SNARE proteins that are not engaged in SNARE complex formation, indicate that SNARE complex formation occurs upstream of regulated neurotransmitter release in vivo [11].
While synaptobrevin and SNAP-25 comprise little more than SNARE motifs and a membrane anchor, syntaxin 1A contains an additional N-terminal domain, which was implicated in regulation of SNARE complex assembly [12–14]. The Habc domain from syntaxin, also classified as a Qa-SNARE [15, 16], folds into a three-helical bundle [17, 18]. Such a conserved, autonomously folded domain is characteristic of all Qa-SNAREs and is found in some Qb SNAREs such as Vti1b [19–22]. In the neuronal SNARE complex the Habc domain is connected to the SNARE motif by a highly flexible linker [23].
When not engaged in SNARE complex formation, the linker region and most of the SNARE motif in rat syntaxin 1A can fold back onto the Habc domain and form a "closed conformation", that interacts with nSec1 (also Munc18-1), an essential positive regulator of neurotransmission [24–26]. nSec1 belongs to the evolutionary conserved family of SM proteins (named after the yeast homologue Sec1p and the mammalian counterpart Munc18), which are involved in all eukaryotic membrane fusion processes [27]. Removal of the Habc domain or mutations resulting in an open conformation significantly accelerate SNARE complex formation in vitro [12–14]. This indicated that nSec1 acts as a negative regulator of membrane fusion. Further evidence however, suggests that nSec1 also plays a positive regulatory role in SNARE-mediated membrane fusion [28–30]. In addition, the presynaptic active zone proteins Munc13-1 and -2 that are essential for priming of synaptic vesicles [31–34] interact with the Habc domain of syntaxin 1 [35]. In order to further investigate the evolutionary conservation of the N-terminal domains of neuronal syntaxin between molluscs and mammals, we determined the crystal structure of the Habc domain derived from the neuronal syntaxin, s-syntaxin, of the squid Loligo pealei.
Results and discussion
s-Syntaxin characterization
s-Syntaxin (residues 2 to 175, additional N-terminal sequence: MRGSPR) comprising the N-terminal Habc domain and the linker region was produced in E. coli and originally tested whether it is sufficient for interaction with s-Sec1, different from the analogous closed nSec1/syntaxin 1A conformation [25]. However, no interaction between s-Sec1 and s-syntaxin(2–175) could be detected by conventional methods including affinity chromatography pull-down, co-purification and native gel electrophoresis (data not shown). Given the fact that nSec1 plays a positive regulatory role during membrane fusion [27], we speculated that a potential low-affinity binding might be sufficient for its action as part of a postulated multi-protein complex at the site of fusion. We therefore tried to co-crystallize s-Sec1 and s-syntaxin in order to trap such potential low-affinity interactions, which however resulted only in crystals containing s-syntaxin. Recently, it has been shown that other SM proteins such as Sly1p and Vps45p interact with an N-terminal helical peptide of their cognate syntaxins mediated by contacts with domain I of the respective SM protein different from the nSec1 closed conformation syntaxin 1A interaction [36–39]. Although s-syntaxin contains this potential peptide region, which has been predicted to have a propensity for α-helix formation [22], no binding in vitro and no co-crystallization could be detected. We can, however, not exclude the possibility that the extra sequences at the N-terminus of s-syntaxin (MRGSPR) might have interfered with binding to s-Sec1.
Structural overview
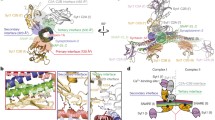
The crystals of s-syntaxin belong to space group P43212 and diffracted X-rays to 3.3 Å resolution using synchrotron radiation. The asymmetric unit contains two molecules of s-syntaxin (molecules A and B, Figure 1A), which both fold into three-helical bundle structures with an approximate length of 45 Å. By convention, the helices are named Ha, Hb and Hc from N- to C-terminus. The N-termini, which include the proposed helical segment, the C-terminal linker to the SNARE motif and all but one of the loop regions connecting the helices were disordered in both molecules. The final models start at residues 31 or 32 and their C-terminal ends are defined by residues 151 (molecule A) and 156 (molecule B). Cα-Atoms of residues 32 to 142 from the two molecules can be superposed with an r.m.s. (root mean square) deviation of 0.555 Å. However, the C-terminal ends diverge substantially (Figure 1B). In molecule A, the third helix Hc is almost four helical turns shorter and the corresponding part, residues 143 to 151, which is highly conserved among syntaxins (Figure 3) forms an open coil structure which binds to a hydrophobic cleft opening between helices Ha and Hc of molecule B (Figure 1A). Although, the C-terminal coil region in molecule A (residues 143 to 151) and the corresponding helical region in molecule B have elevated B-factors compared to the overall model (A, 67.0 Å2, and B, 66.1 Å2, versus all atoms, 42.3 Å2), the electron density in this region is well interpretable (Figure 2). The extended conformation is stabilized by an intermolecular disulfide bond, Cys148A-Cys148B, and three hydrogen bonds (Figure 2). It should be noted, that if both syntaxin molecules assumed a conformation like in molecule B severe steric clashes would result. In molecule B, Cys148 lies rather protected at the interface of helices Hb and Hc. This indicates that the C-terminal end of the Habc domain can adopt different conformations, which may be important when interacting with different regulatory partners thus adding to the structural plasticity observed for neuronal syntaxins [40]. In accordance, a recent time-resolved FRET (fluorescence resonance energy transfer) study indicates that rat neuronal syntaxin 1A undergoes fast conformational conversion with only a minority population in the closed conformation [41]. Structural flexibility at or close to the linker region of syntaxin might promote rapid interconversion of conformations. Therefore, this additional syntaxin conformation might be of physiological relevance as it is not obvious how local unfolding would have been favored by the mild crystallization conditions (see Methods).
Asymmetric unit content in crystals of neuronal syntaxin from L. pealei (A) Ribbon diagram of the dimer in the asymmetric unit. N- and C-terminal ends are indicated. A brace indicates the C-terminal Habc region (in molecule B), which can have either helical or extended conformation. (B) Overlay of the Cα backbone of molecules A and B. Note that the C-terminus of molecule A deviates from the helical fold of Hc from molecule B as 4 helical turns are unwound.
Stereo diagram of the electron density at the interface of the s-syntaxin dimer. 2Fo-Fc electron density at 1.0 σ of the contact region between molecule B and the C-terminus of molecule A is shown as a meshwork in cyan. Carbon atoms in molecules A and B are shown in beige and grey, respectively. Heteroatoms are indicated in red (oxygen), blue (nitrogen) and yellow (sulphur). Dashed lines indicate polar interactions stabilizing such a conformation. Residues 32 to 69 in molecule B have been omitted for clarity.
Structure based sequence alignment of neuronal syntaxins Sequences from five animal phyla (SwissProt/ TrEMBL accession codes, L. pealei O46345, H. medicinalis O01390, D. melanogaster Q24547, C. elegans O16000, R. norvegicus P32851) were used. Secondary structure elements are indicated for neuronal syntaxin from L. pealei, above the sequence, based on the crystal structure of the Habc domain and the squid SNARE complex (pdb code 1L4A) [62]. Secondary structure element assignment for the rat homologue is based on the X-ray structure of syntaxin 1A in complex with nSec1 (pdb code 1DN1) [25] and is shown below the sequence, thus highlighting the different conformations that can be adopted by the SNARE motif. Identical residues are indicated by red background. Homologous residues are shown in red. Green triangles indicate syntaxin residues that interact with nSec1 in the closed conformation [25]. The region with divergent conformations at the C-terminus of the squid Habc domain is marked with a blue dashed line.
Molecule B and the core of molecule A are however strikingly similar to mammalian syntaxin 1A. The secondary structure of the three-helical bundle domain of rat neuronal syntaxin was initially determined by NMR (pdb code 1BR0 [17]) and confirmed by X-ray crystallography (pdb code 1EZ3 [18]). In addition, the linker region and the SNARE motif of neuronal syntaxin fold back onto the three-helical bundle in the closed conformation, as observed in the crystal structure of the complex with nSec1 (pdb code 1DN1 [25]), which leaves the N-terminal domain virtually unaltered.
The Cα atoms of squid (molecule B) and rat neuronal syntaxin (X-ray structure, chains A to C [18]) can be superposed with r.m.s. deviations between 0.711 and 0.761 Å for residues 32 to 149 (not shown). These values are very close to the deviation between the two copies in the asymmetric unit of squid syntaxin itself and therefore the structures of rat and squid neuronal syntaxin can be considered virtually identical despite the considerable evolutionary distance. The deviation from the all-helical fold at the C-terminal region of the Habc domain in molecule A, however, is novel albeit its functional significance has yet to be proven.
Sequence conservation and functional implications
Within animals, neuronal syntaxin is remarkably well conserved. The homologues of rat, H. medicinalis, C. elegans and D. melanogaster share 47.5 % sequence identity (80.3 % homology) with the sequence of the squid L. pealei (Figure 3). In comparison to the SNARE motif, which is highly conserved (residues 183 to 255, 63 % identity, 86.3 % homology), the three-helical bundle domains share somewhat lower sequence identity (residues 32 to 154, 50.8 % identity, 75.6 % homology). The surface of the Habc domain is highly conserved however (Figures 4B and 4E), underlining their functional conservation. While the N-terminal region preceding the three-helical bundle is generally the least conserved region in neuronal syntaxin, the ultimate N-terminus, which comprises the SM protein binding signature region, residues 1 to 18, in the syntaxin homologues Sed5p and Tlg2p exhibits remarkable conservation [36–38]. This is even more pronounced among the invertebrate sequences. However, in vitro binding of this region of neuronal syntaxin to nSec1 could not be shown experimentally for both the rat and the squid proteins [39, 42] (unpublished observations).
Surface conservation of squid neuronal syntaxin Two projections of squid neuronal syntaxin related by an 180° rotation are shown. In the middle panels B and E, shades of green indicate the degree of amino acid conservation within distantly related neuronal syntaxins, white, no conservation, and dark green, absolute conservation. The surface conservation score was calculated with ESPript based on the sequence alignment in Figure 3[60]. Conserved surface residues in the putative Munc13 binding region are labeled. On the right (panels C and F), rat syntaxin 1A residues involved in intramolecular closed conformation and nSec1 interactions are shown for comparison in turquoise and orange, respectively. Ribbon representations of syntaxin using rainbow color-coding are shown on the left for orientation (panels A and D).
Surface regions of high amino acid sequence conservation within neuronal syntaxins match very well with the sites of intramolecular interactions in the closed conformation. Indeed all residues involved in the formation of the closed conformation as well as those involved in contacting nSec1 in the complex structure [25] are strictly conserved between squid and rat syntaxin (Figures 3, 4). Therefore, we predict that squid neuronal syntaxin can assume the same closed conformation as its rat homologue and that it will bind s-Sec1 in the same way [16], although the degree of conservation between squid and rat neuronal SM proteins is somewhat lower (66 % vs. 83 % identity).
The syntaxin binding partner Munc13 (murine Unc-13) was found to be essential for neurotransmitter release in organisms ranging from C. elegans to mammals [31, 32, 43], a conservation which also postulates a squid homologue of Munc13. It has been proposed that Munc13 facilitates the transition from the closed syntaxin conformation in complex with nSec1 to the open state participating in SNARE complex assembly [43, 44]. This is in accordance with biochemical data showing that Munc13 interacts with the three-helical bundle domain of neuronal syntaxin in a yeast two-hybrid screen [35]. Specifically, the binding region was located between residues 53 and 79 in rat neuronal syntaxin, which includes part of Ha, the linker and part of Hb (Figure 3). The proposed binding site partly overlaps with the binding of the SNARE motif in the closed conformation, which might render nSec1 and Munc13 interactions exclusive.
Interestingly, the surface corresponding to this region also contains highly conserved residues, which are involved in neither the intramolecular closed conformation nor the intermolecular nSec1 interactions (arrow, Figure 4). In addition, the extended conformation of the C-terminal end of Habc locates next to this putative Munc13 interaction site. Therefore it is possible that the structural changes might be either necessary for Munc13 binding or might be a consequence of Munc13 binding or of some other potential ligand in vivo.
Another possible interaction partner of syntaxin that is capable of dissociating Munc18 from syntaxin-1 is tomosyn [45], which might imply interaction with the Habc domain. On the other hand, a C-terminal SNARE motif in tomosyn competes for SNARE complex assembly [46].
Function of the three-helical bundle domain of neuronal syntaxin
Three-helical bundle Habc domains appear to be a general feature of syntaxin-type SNARE proteins, e.g. the paralogous yeast syntaxins Sso1p, Sed5p, Tlg2p and Vam3p have all been shown to contain homologous domains [20, 21, 36, 37]. Furthermore, Q-SNARES such as syntaxin 6, syntaxin 7 and Vti1p, which substitute SNAP-25 in the endosomal SNARE complex, contain structurally similar three-helical bundle domains [19, 22].
The Habc domain of neuronal syntaxin interferes with SNARE complex formation by promoting the closed conformation and serves as a scaffold for the interaction with nSec1. However, only the syntaxin homologue Sso1p was shown to form a stable closed conformation [21]. Sec1p, the corresponding SM protein and the closest homologue to nSec1 in yeast, binds exclusively to the assembled SNARE complex [47]. Furthermore, short N-terminal signature sequences in the yeast syntaxin homologues Sed5p and Tlg2p and their mammalian counterparts are sufficient for recruitment of the respective SM proteins Sly1p and Vps45p [36–38].
The structural conservation of the autonomously folded N-terminal domain of Qa SNAREs suggest that the Habc domains might recruit not only SM proteins but also other regulatory proteins involved in vesicle trafficking. Accordingly, constitutively open full-length but not N-terminally truncated yeast Sso1p can complement a ΔSSO1 ΔSSO2 yeast strain [21, 48]. In addition, the formation of a closed conformation as observed in the neuronal system seems to be rather an exception among diverse syntaxins and might be necessary to ensure a tighter control of the syntaxin conformation and thus SNARE complex assembly than in other vesicle transport and fusion systems.
Conclusions
Structural and evolutionary conservation of the Habc domain of squid neuronal syntaxin indicates the fundamental role of neuronal syntaxin for neurotransmission. Almost identical conformation and strong conservation of surface residues suggest that the interplay of neuronal syntaxin with its interaction partners nSec1 and Munc13 is conserved from mollusks to mammals. Therefore, squid neuronal syntaxin undergoes the same conformational cycle as its rat homologue. Interestingly, the structure also indicates some newly identified structural changes at the C-terminus of helix Hc, which might play an important role in the control of the conformational flexibility of syntaxins and thus in membrane fusion at the synapse.
Methods
Cloning, expression and purification
A fragment of Loligo pealei syntaxin comprising residues 2 to 175 was cloned into a modified pQE30 vector (Qiagen) missing the region coding for a N-terminal His6-tag (additional N-terminal sequence, MRGSPR). The protein was expressed in E. coli Bl21 pUBS cells.
After 3 h of induction at 37°C, the cells were lysed by ultrasonication in 50 mM HEPES KOH pH 7.4. The clarified supernatant was applied to a Sepharose Q column (Amersham Biotech) at 4°C and eluted with a linear gradient to 500 mM KCl. Subsequently, 4 M ammonium sulphate was added until the pooled fractions had a final concentration of 0.8 M. The solution was applied to a Phenyl Sepharose 6 FF column (Amersham Biotech), and the unbound fraction collected. Size exclusion chromatography on Superdex200 (Amersham Biotech) equilibrated with 20 mM HEPES KOH pH 7.4 and 100 mM KCl served as final purification step.
Crystallisation
Large octahedral crystals were obtained by the hanging drop method after mixing equal volumes of 26 mg/ml s-syntaxin with reservoir buffer containing either 30 % PEG-400, 0.1 M Tris HCl pH 8.5 and 0.2 M Na-citrate or 18 – 20 % PEG-550MME, 0.1 M Tris HCl pH 8.5 and 0.2 M Na-citrate. For cryo-protection, the crystals were incubated with 30 % PEG-400, 10 % glycerol, 0.1 M Tris HCl pH 8.5 and 0.2 M Na-citrate over night. The crystals of space group P 43212 with lattice dimensions of a, b = 73.056 Å and c = 224.434 Å diffracted to 3.3 Å resolution at ESRF beamline ID29.
Structure solution and refinement
Diffraction data were processed and scaled with Denzo and Scalepack [49]. The structure was solved by molecular replacement using the program AMoRe with rat syntaxin (chain A in pdb entry 1ez3 [18]) as a search model [50]. The asymmetric unit contains two molecules. A Matthews coefficient of 3.74 Å3 Da-1 and a solvent content of 66.9 % were calculated [51]. The model was refined using simulated annealing, gradient minimization and individual B-factor refinement as implemented in CNS alternated by rebuilding cycles using the program O [52, 53]. In a final step, the tensor elements describing the anisotropic displacement of the individual helices of syntaxin were refined using the TLS option in Refmac5 followed by 5 cycles of restrained minimisation [54, 55]. The final R-factor is 0.309 with a Rfree of 0.375.
The final model contains only 227 out of 360 residues including residues 31 to 66, 74 to 107 and 113 to 151 of chain A and residues 32 to 105 and 113 to 156 of chain B. All of the residues are in allowed regions of the Ramachandran plot as defined in Procheck [56]. Due to missing electron density for side chains, 66 solvent exposed residues were modeled as alanine, which, together with the disordered regions (> 1/3 of the protein crystal content) contribute to the somewhat high R factors.
The model coordinates were deposited at the pdb database under accession code 1s94.
Figures were generated with the programs Molscript [57], Bobscript [58], Raster-3D [59], ESPript [60] and Grasp [61].
References
Jahn R, Südhof TC: Membrane fusion and exocytosis. Annu Rev Biochem 1999, 68: 863–911. 10.1146/annurev.biochem.68.1.863
Fukuda R, McNew JA, Weber T, Parlati F, Engel T, Nickel W, Rothman JE, Söllner TH: Functional architecture of an intracellular membrane t-SNARE. Nature 2000, 407: 198–202. 10.1038/35025084
Fasshauer D, Eliason WK, Brunger AT, Jahn R: Identification of a minimal core of the synaptic SNARE complex sufficient for reversible assembly and disassembly. Biochemistry 1998, 37: 10354–10362. 10.1021/bi980542h
Sutton RB, Fasshauer D, Jahn R, Brunger AT: Crystal structure of a SNARE complex involved in synaptic exocytosis at 2.4 A resolution. Nature 1998, 395: 347–353. 10.1038/26412
Fergestad T, Wu MN, Schulze KL, Lloyd TE, Bellen HJ, Broadie K: Targeted mutations in the syntaxin H3 domain specifically disrupt SNARE complex function in synaptic transmission. J Neurosci 2001, 21: 9142–9150.
Schoch S, Deak F, Königstorfer A, Mozhayeva M, Sara Y, Südhof TC, Kavalali ET: SNARE function analyzed in synaptobrevin/VAMP knockout mice. Science 2001, 294: 1117–1122. 10.1126/science.1064335
Washbourne P, Thompson PM, Carta M, Costa ET, Mathews JR, Lopez-Bendito G, Molnar Z, Becher MW, Valenzuela CF, Partridge LD, Wilson MC: Genetic ablation of the t-SNARE SNAP-25 distinguishes mechanisms of neuroexocytosis. Nat Neurosci 2002, 5: 19–26.
Sorensen JB, Nagy G, Varoqueaux F, Nehring RB, Brose N, Wilson MC, Neher E: Differential control of the releasable vesicle pools by SNAP-25 splice variants and SNAP-23. Cell 2003, 114: 75–86. 10.1016/S0092-8674(03)00477-X
Parlati F, Weber T, McNew JA, Westermann B, Söllner TH, Rothman JE: Rapid and efficient fusion of phospholipid vesicles by the alpha-helical core of a SNARE complex in the absence of an N-terminal regulatory domain. Proc Natl Acad Sci U S A 1999, 96: 12565–12570. 10.1073/pnas.96.22.12565
Weber T, Zemelman BV, McNew JA, Westermann B, Gmachl M, Parlati F, Söllner TH, Rothman JE: SNAREpins: minimal machinery for membrane fusion. Cell 1998, 92: 759–772. 10.1016/S0092-8674(00)81404-X
Rettig J, Neher E: Emerging roles of presynaptic proteins in Ca++-triggered exocytosis. Science 2002, 298: 781–785. 10.1126/science.1075375
Dulubova I, Sugita S, Hill S, Hosaka M, Fernandez I, Südhof TC, Rizo J: A conformational switch in syntaxin during exocytosis: role of munc18. Embo J 1999, 18: 4372–4382. 10.1093/emboj/18.16.4372
Fiebig KM, Rice LM, Pollock E, Brunger AT: Folding intermediates of SNARE complex assembly. Nat Struct Biol 1999, 6: 117–123. 10.1038/5803
Nicholson KL, Munson M, Miller RB, Filip TJ, Fairman R, Hughson FM: Regulation of SNARE complex assembly by an N-terminal domain of the t-SNARE Sso1p. Nat Struct Biol 1998, 5: 793–802. 10.1038/1834
Fasshauer D, Sutton RB, Brunger AT, Jahn R: Conserved structural features of the synaptic fusion complex: SNARE proteins reclassified as Q- and R-SNAREs. Proc Natl Acad Sci U S A 1998, 95: 15781–15786. 10.1073/pnas.95.26.15781
Bock JB, Matern HT, Peden AA, Scheller RH: A genomic perspective on membrane compartment organization. Nature 2001, 409: 839–841. 10.1038/35057024
Fernandez I, Ubach J, Dulubova I, Zhang X, Südhof TC, Rizo J: Three-dimensional structure of an evolutionarily conserved N-terminal domain of syntaxin 1A. Cell 1998, 94: 841–849. 10.1016/S0092-8674(00)81742-0
Lerman JC, Robblee J, Fairman R, Hughson FM: Structural analysis of the neuronal SNARE protein syntaxin-1A. Biochemistry 2000, 39: 8470–8479. 10.1021/bi0003994
Antonin W, Dulubova I, Arac D, Pabst S, Plitzner J, Rizo J, Jahn R: The N-terminal domains of syntaxin 7 and vti1b form three-helix bundles that differ in their ability to regulate SNARE complex assembly. J Biol Chem 2002, 277: 36449–36456. 10.1074/jbc.M204369200
Dulubova I, Yamaguchi T, Wang Y, Südhof TC, Rizo J: Vam3p structure reveals conserved and divergent properties of syntaxins. Nat Struct Biol 2001, 8: 258–264. 10.1038/85012
Munson M, Chen X, Cocina AE, Schultz SM, Hughson FM: Interactions within the yeast t-SNARE Sso1p that control SNARE complex assembly. Nat Struct Biol 2000, 7: 894–902. 10.1038/79659
Misura KM, Bock JB, Gonzalez L. C., Jr., Scheller RH, Weis WI: Three-dimensional structure of the amino-terminal domain of syntaxin 6, a SNAP-25 C homolog. Proc Natl Acad Sci U S A 2002, 99: 9184–9189. 10.1073/pnas.132274599
Margittai M, Fasshauer D, Jahn R, Langen R: The Habc domain and the SNARE core complex are connected by a highly flexible linker. Biochemistry 2003, 42: 4009–4014. 10.1021/bi027437z
Verhage M, Maia AS, Plomp JJ, Brussaard AB, Heeroma JH, Vermeer H, Toonen RF, Hammer RE, van den Berg TK, Missler M, Geuze HJ, Südhof TC: Synaptic assembly of the brain in the absence of neurotransmitter secretion. Science 2000, 287: 864–869. 10.1126/science.287.5454.864
Misura KM, Scheller RH, Weis WI: Three-dimensional structure of the neuronal-Sec1-syntaxin 1a complex. Nature 2000, 404: 355–362. 10.1038/35006120
Yang B, Steegmaier M, Gonzalez L. C., Jr., Scheller RH: nSec1 binds a closed conformation of syntaxin1A. J Cell Biol 2000, 148: 247–252. 10.1083/jcb.148.2.247
Jahn R, Lang T, Südhof TC: Membrane fusion. Cell 2003, 112: 519–533. 10.1016/S0092-8674(03)00112-0
Dresbach T, Burns ME, O'Connor V, DeBello WM, Betz H, Augustine GJ: A neuronal Sec1 homolog regulates neurotransmitter release at the squid giant synapse. J Neurosci 1998, 18: 2923–2932.
Wu MN, Fergestad T, Lloyd TE, He Y, Broadie K, Bellen HJ: Syntaxin 1A interacts with multiple exocytic proteins to regulate neurotransmitter release in vivo. Neuron 1999, 23: 593–605. 10.1016/S0896-6273(01)80039-8
Voets T, Toonen RF, Brian EC, de Wit H, Moser T, Rettig J, Südhof TC, Neher E, Verhage M: Munc18–1 promotes large dense-core vesicle docking. Neuron 2001, 31: 581–591. 10.1016/S0896-6273(01)00391-9
Aravamudan B, Fergestad T, Davis WS, Rodesch CK, Broadie K: Drosophila UNC-13 is essential for synaptic transmission. Nat Neurosci 1999, 2: 965–971. 10.1038/14764
Augustin I, Rosenmund C, Südhof TC, Brose N: Munc13–1 is essential for fusion competence of glutamatergic synaptic vesicles. Nature 1999, 400: 457–461. 10.1038/22768
Richmond JE, Davis WS, Jorgensen EM: UNC-13 is required for synaptic vesicle fusion in C. elegans. Nat Neurosci 1999, 2: 959–964. 10.1038/12160
Varoqueaux F, Sigler A, Rhee JS, Brose N, Enk C, Reim K, Rosenmund C: Total arrest of spontaneous and evoked synaptic transmission but normal synaptogenesis in the absence of Munc13-mediated vesicle priming. Proc Natl Acad Sci U S A 2002, 99: 9037–9042. 10.1073/pnas.122623799
Betz A, Okamoto M, Benseler F, Brose N: Direct interaction of the rat unc-13 homologue Munc13–1 with the N terminus of syntaxin. J Biol Chem 1997, 272: 2520–2526. 10.1074/jbc.272.4.2520
Yamaguchi T, Dulubova I, Min SW, Chen X, Rizo J, Südhof TC: Sly1 binds to Golgi and ER syntaxins via a conserved N-terminal peptide motif. Dev Cell 2002, 2: 295–305. 10.1016/S1534-5807(02)00125-9
Dulubova I, Yamaguchi T, Gao Y, Min SW, Huryeva I, Südhof TC, Rizo J: How Tlg2p/syntaxin 16 'snares' Vps45. Embo J 2002, 21: 3620–3631. 10.1093/emboj/cdf381
Bracher A, Weissenhorn W: Structural basis for the Golgi membrane recruitment of Sly1p by Sed5p. Embo J 2002, 21: 6114–6124. 10.1093/emboj/cdf608
Dulubova I, Yamaguchi T, Arac D, Li H, Huryeva I, Min SW, Rizo J, Südhof TC: Convergence and divergence in the mechanism of SNARE binding by Sec1/Munc18-like proteins. Proc Natl Acad Sci U S A 2003, 100: 32–37. 10.1073/pnas.232701299
Misura KM, May AP, Weis WI: Protein-protein interactions in intracellular membrane fusion. Curr Opin Struct Biol 2000, 10: 662–671. 10.1016/S0959-440X(00)00151-2
Margittai M, Widengren J, Schweinberger E, Schröder GF, Felekyan S, Haustein E, König M, Fasshauer D, Grubmuller H, Jahn R, Seidel CA: Single-molecule fluorescence resonance energy transfer reveals a dynamic equilibrium between closed and open conformations of syntaxin 1. Proc Natl Acad Sci U S A 2003, 100: 15516–15521. 10.1073/pnas.2331232100
Calakos N, Bennett MK, Peterson KE, Scheller RH: Protein-protein interactions contributing to the specificity of intracellular vesicular trafficking. Science 1994, 263: 1146–1149.
Richmond JE, Weimer RM, Jorgensen EM: An open form of syntaxin bypasses the requirement for UNC-13 in vesicle priming. Nature 2001, 412: 338–341. 10.1038/35085583
Sassa T, Harada S, Ogawa H, Rand JB, Maruyama IN, Hosono R: Regulation of the UNC-18-Caenorhabditis elegans syntaxin complex by UNC-13. J Neurosci 1999, 19: 4772–4777.
Fujita Y, Shirataki H, Sakisaka T, Asakura T, Ohya T, Kotani H, Yokoyama S, Nishioka H, Matsuura Y, Mizoguchi A, Scheller RH, Takai Y: Tomosyn: a syntaxin-1-binding protein that forms a novel complex in the neurotransmitter release process. Neuron 1998, 20: 905–915.
Hatsuzawa K, Lang T, Fasshauer D, Bruns D, Jahn R: The R-SNARE motif of tomosyn forms SNARE core complexes with syntaxin 1 and SNAP-25 and down-regulates exocytosis. J Biol Chem 2003, 278: 31159–31166. 10.1074/jbc.M305500200
Carr CM, Grote E, Munson M, Hughson FM, Novick PJ: Sec1p binds to SNARE complexes and concentrates at sites of secretion. J Cell Biol 1999, 146: 333–344. 10.1083/jcb.146.2.333
Munson M, Hughson FM: Conformational regulation of SNARE assembly and disassembly in vivo. J Biol Chem 2002, 277: 9375–9381. 10.1074/jbc.M111729200
Otwinowski Z, Minor W: Processing of x-ray data collected in oscillation mode. Methods Enzymol 1997, 276: 307–326.
Navaza J: AMoRe: an Automated Package for Molecular Replacement. Acta Crystallogr A 1994, 50: 157–163. 10.1107/S0108767393007597
Collaborative Computational Project Number 4: The CCP4 suite: programs for protein crystallography. Acta Crystallogr D 1994, 50: 760–763. 10.1107/S0907444994003112
Brunger AT, Adams PD, Clore GM, DeLano WL, Gros P, Grosse-Kunstleve RW, Jiang JS, Kuszewski J, Nilges M, Pannu NS, Read RJ, Rice LM, Simonson T, Warren GL: Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr D 1998, 54: 905–921. 10.1107/S0907444998003254
Jones TA, Zou JY, Cowan SW, Kjelgaard M: Improved methods for building protein models in electron density maps and location of errors in these models. Acta Crystallogr A 1991, 47: 110–119. 10.1107/S0108767390010224
Murshudov GN, Vagin AA, Dodson EJ: Refinement of Macromolecular Structures by the Maximum-Likelihood Method. Acta Crystallogr D 1997, 53: 240–255. 10.1107/S0907444996012255
Winn MD, Isupov MN, Murshudov GN: Use of TLS parameters to model anisotropic displacements in macromolecular refinement. Acta Crystallogr D 2001, 57: 122–133. 10.1107/S0907444900014736
Laskowski RA, MacArthur MW, Moss DS, Thornton JM: PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Cryst 1993, 26: 283–290. 10.1107/S0021889892009944
Kraulis P: MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J Appl Cryst 1991, 24: 946–950. 10.1107/S0021889891004399
Esnouf RM: An extensively modified version of MolScript that includes greatly enhanced coloring capabilities. J Mol Graph Model 1997, 15: 132–4, 112–3. 10.1016/S1093-3263(97)00021-1
Merritt EA, Bacon DJ: Raster3D photorealistic graphics. Methods Enzymol 1997, 277: 505–524.
Gouet P, Courcelle E, Stuart DI, Metoz F: ESPript: multiple sequence alignments in PostScript. Bioinformatics 1999, 15: 305–308. 10.1093/bioinformatics/15.4.305
Nicholls A, Sharp KA, Honig B: Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 1991, 11: 281–296.
Bracher A, Kadlec J, Betz H, Weissenhorn W: X-ray structure of a neuronal complexin-SNARE complex from squid. J Biol Chem 2002, 277: 26517–26523. 10.1074/jbc.M203460200
Acknowledgements
We thank Drs. Thomas Dresbach and Heinrich Betz for providing the initial clone for squid neuronal syntaxin and JSBG staff at ESRF (Grenoble, France) beamline ID29 for help with data collection. A.B. was supported by a Marie Curie Fellowship from the European Union.
Author information
Authors and Affiliations
Corresponding author
Additional information
Authors' contributions
A.B. carried out the cloning, purification, crystallization, data collection, structure solution and refinement of s-syntaxin(2–175). W.W. conceived of the study, and participated in its design, coordination and writing of the manuscript. All authors read and approved the final manuscript.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Bracher, A., Weissenhorn, W. Crystal structure of the Habc domain of neuronal syntaxin from the squid Loligo pealei reveals conformational plasticity at its C-terminus. BMC Struct Biol 4, 6 (2004). https://doi.org/10.1186/1472-6807-4-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1472-6807-4-6