Abstract
Background
Increased circulating levels of hemostatic factors as well as anemia have been associated with increased risk of cardiovascular disease (CVD). Known associations between hemostatic factors and sequence variants at genes encoding these factors explain only a small proportion of total phenotypic variation. We sought to confirm known putative loci and identify novel loci that may influence either trait in genome-wide association and linkage analyses using the Affymetrix GeneChip 100K single nucleotide polymorphism (SNP) set.
Methods
Plasma levels of circulating hemostatic factors (fibrinogen, factor VII, plasminogen activator inhibitor-1, von Willebrand factor, tissue plasminogen activator, D-dimer) and hematological phenotypes (platelet aggregation, viscosity, hemoglobin, red blood cell count, mean corpuscular volume, mean corpuscular hemoglobin concentration) were obtained in approximately 1000 Framingham Heart Study (FHS) participants from 310 families. Population-based association analyses using the generalized estimating equations (GEE), family-based association test (FBAT), and multipoint variance components linkage analyses were performed on the multivariable adjusted residuals of hemostatic and hematological phenotypes.
Results
In association analysis, the lowest GEE p-value for hemostatic factors was p = 4.5*10-16 for factor VII at SNP rs561241, a variant located near the F7 gene and in complete linkage disequilibrium (LD) (r2 = 1) with the Arg353Gln F7 SNP previously shown to account for 9% of total phenotypic variance. The lowest GEE p-value for hematological phenotypes was 7*10-8 at SNP rs2412522 on chromosome 4 for mean corpuscular hemoglobin concentration. We presented top 25 most significant GEE results with p-values in the range of 10-6 to 10-5 for hemostatic or hematological phenotypes. In relating 100K SNPs to known candidate genes, we identified two SNPs (rs1582055, rs4897475) in erythrocyte membrane protein band 4.1-like 2 (EPB41L2) associated with hematological phenotypes (GEE p < 10-3). In linkage analyses, the highest linkage LOD score for hemostatic factors was 3.3 for factor VII on chromosome 10 around 15 Mb, and for hematological phenotypes, LOD 3.4 for hemoglobin on chromosome 4 around 55 Mb. All GEE and FBAT association and variance components linkage results can be found at http://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?id=phs000007
Conclusion
Using genome-wide association methodology, we have successfully identified a SNP in complete LD with a sequence variant previously shown to be strongly associated with factor VII, providing proof of principle for this approach. Further study of additional strongly associated SNPs and linked regions may identify novel variants that influence the inter-individual variability in hemostatic factors and hematological phenotypes.
Similar content being viewed by others
Background
The relationship of hemostasis and thrombosis with atherothrombotic cardiovascular disease has been extensively studied in the past decades. Elevated circulating levels of hemostatic factors, such as fibrinogen [1–3], plasminogen activator inhibitor (PAI-1) [4, 5], von Willebrand factor (vWF) [6], tissue plasminogen activator (tPA) [4, 5, 7], factor VII (FVII) [8], and D-dimer [9, 10] are linked to the development of atherothrombosis and are risk markers for coronary heart disease (CHD), stroke and other cardiovascular disease (CVD) events.
In addition to coagulation proteins, the cellular and rheological components of circulating blood have been implicated in CHD, stroke and peripheral arterial disease, including hematological phenotypes such as hematocrit (HCT), hemoglobin (Hgb), red blood cell count (RBCC) and size, mean corpuscular volume (MCV) and mean corpuscular hemoglobin (MCH) [11, 12], as well as measures of platelet aggregation (induced by adenosine 5'-diphosphate (ADP), epinephrine (Epi) and collagen respectively) [12, 13], and viscosity [14, 15].
Cis-acting sequence variants in the following genes – fibrinogen-β (FGB), fibrinogen-α (FGA), fibrinogen-γ (FGG), FVII (F7), and PAI-1 (SERPINE1) – have been associated with corresponding levels of circulating hemostatic factor. By comprehensively characterizing common genetic variation at each of these loci, we have recently clarified that cis-acting variants, in sum, explain a modest proportion of phenotypic variation, ranging from 1% – 10% [16, 17]. For hematological variables such as hematocrit and hemoglobin, sequence variation in the major hemoglobin genes is well described to be associated with anemias, such as beta- and alpha-thalassemia, and sickle cell anemia [18–20].
Systematic searches for novel genes beyond the known genetic determinants influencing these phenotypes have been carried out using genome-wide linkage analyses with microsatellite markers: Chromosome regions that may harbor novel loci influencing fibrinogen, PAI-1 [21, 22], hematocrit, Hgb, RBCC, MCV and MCH [23, 24], have been identified. However, linkage scans with microsatellite markers generally had low power to detect loci with small effects, and lacked precision in localizing the loci; thus, few novel loci have been identified.
The recent completion of a genome-wide scan using the Affymetrix GeneChip Human Mapping 100K single nucleotide polymorphism (SNP) set on participants in the Framingham Heart Study offered the opportunity to conduct a genome-wide association study (GWAS) and linkage scan for variants that influence hemostatic factors and hematological phenotypes.
Methods
Study participants and genotyping methods
The Framingham Heart Study design and the genotyping of the Affymetrix GeneChip Human Mapping 100K SNP set on Framingham Heart Study participants are detailed in the overview of this project [25]. To avoid potential bias due to genotyping artifacts, we limited the association analyses to 70987 SNPs on autosomes with minor allele frequency (MAF) ≥ 10%, genotyping call rate ≥ 80%, and Hardy-Weinberg equilibrium test p-value ≥ 0.001.
Measurements of hemostatic factors and hematological phenotypes
Venous blood samples of Framingham Heart Study Offspring Cohort taken at the first and second examination cycles (1971–1975, and 1979–1983) were used to measure Hgb, RBCC, MCV and MCH, and samples taken at the fifth examination cycle (1991–1995) were used to measure all the hemostatic factors, platelet aggregation, D-dimer, and viscosity. Fibrinogen was additionally measured at the sixth (1995–1998) and seventh (1998–2001) examination cycles, and PAI-I antigen levels at the sixth exam. Details of the assessment of hemostatic factor levels have been described previously [17, 26]. Plasma fibrinogen levels were measured using the Clauss method [27]. Plasma PAI-I antigen, tPA antigen, von Willebrand factor and FVII antigen were assessed using enzyme-linked immunosorbent assays.
The determination of hematological phenotypes has been detailed previously. Platelet aggregation was performed according to the method of Born [28]. The reagents used were epinephrine, ADP and collagen. The percent extent of aggregation in duplicate to epinephrine and ADP was determined in varying concentrations (0.01 to 15 mmol/L). For each subject, the aggregation response (yes/no) was also tested to a fixed concentration of arachidonic acid (5 mg/mL). The collagen lag time was measured in response to 1.9 mmol/L collagen. Participants who were taking aspirin were excluded from the analyses for platelet aggregation phenotypes as well as PAI-1 and tPA.
HCT was measured by the Wintrobe method [29]. Blood was collected and spun at 5000 rpm for 20 minutes in a balanced oxalate tube. The percent of total blood volume that was due to red blood cells was determined visually against a calibrated scale. MCV is the average volume of an individual's red blood cells determined as the ratio of HCT to RBCC. MCH is the average amount of hemoglobin of an individual's red cell determined as the ratio of Hgb to RBCC.
Statistical methods
Standardized multivariable adjusted residuals of the hemostatic and hematological phenotypes were computed and used in all the linkage and association analyses. Covariates used in the adjustments were determined based upon what has been reported in the literature as potential risk factors for hemostatic factors or hematological phenotypes. Hardy-Weinberg equilibrium was examined using an exact chi-square test statistic [30]. Association between each SNP and each hemostatic or hematological phenotype was examined using a population based association method via generalized estimating equations (GEE) [31] and family-based association test (FBAT) [32], assuming an additive genetic model. Variance components linkage analyses were conducted using a subset of SNPs with pairwise r2 < 0.5. Details of both association and linkage methods are described in the overview of this project [25].
In secondary analyses, we combined the GEE association tests results across multiple phenotypes that may share the common pathway to reduce the type I error rates, and possibly detect SNPs of smaller effect sizes. We ranked SNPs by the number of GEE test p-values less than 0.01, and then by the geometric mean of the GEE test p-values. We also examined the β coefficient from the GEE regression that is the change in the phenotype in one standardized deviation unit with an increment of a copy of the alphabetically second allele (for example, allele G for a SNP with alleles A and G). This analysis was conducted for a phenotype assessed using multiple measurement methods such as the platelet aggregation with ADP-, collagen-, and Epi-induced platelet aggregation; or for a phenotype with serial measurements such as fibrinogen level measured at examination cycles 5, 6 and 7.
We attempted to identify association of 100K SNPs in or within 60 kilo base pairs (kbp) of selected candidate genes previously reported to be associated with hemostatic factors or hematological phenotypes. For hemostatic factors and platelet aggregation phenotypes, we included the following candidate genes in the search: F7, fibrinogen gene cluster (FGB, FGA, FGG), SERPINE1, plasminogen activator-tissue (PLAT), vWF and integrin beta 3 (ITGB3). For hematological phenotypes excluding platelet aggregation, we included erythropoietin receptor (EPOR), erythropoietin (EPO), erythrocyte membrane protein band 4.1-like 2 (EPB41L2), Kruppel-like factor 1(KLF1), heme binding protein 2 (HEBP2), the hemoglobin gene clusters on chromosome 11: hemoglobin-β chain complex (HBB), hemoglobin-δ (HBD), hemoglobin-γ A (HBG1), hemoglobin-γ G (HBG2), hemoglobin-ε 1 (HBE1), and the hemoglobin gene clusters on chromosome 16: hemoglobin-α 1 (HBA1), hemoglobin-α 2 (HBA2), hemoglobin-μ (HBM).
Results
Table 1 displays the hemostatic and hematological phenotypes analyzed in this study, as well as the number of individuals, examination cycles, and covariates used in multivariable models. The sample size ranged from 702 to 1073. Traits measured at multiple examinations were analyzed using multivariable adjusted residuals from each examination measure, and also the average of all the multivariable adjusted residuals from individual examination cycles.
Among individuals who were included in the genotyping and had at least one hemostatic factor or platelet aggregation phenotype measured at examination cycle five, 52% were women, mean age was 52 years, and 6% had prevalent CVD. Among individuals who were included in the genotyping and had at least one hematological phenotype measured at examination cycle one or two, 52% were women, with a mean age over the two examinations of 36 years, and 2% had prevalent CVD.
Association between SNPs and hemostatic and hematological phenotypes
We report the 25 SNPs with lowest GEE association test p-values in Table 2 for hemostatic factors, and in Table 3 for hematological phenotypes. The lowest GEE p-value (4.5*10-16) for hemostatic factors was obtained from the test of association between circulating levels of FVII and rs561241; this SNP resides near the F7 gene on chromosome 13 and is in complete linkage disequilibrium (LD) (r2 = 1) with the Arg353Gln F7 SNP (rs6046) we previously reported to account for 9% of total phenotypic variance [16]. The lowest GEE p-value (6.9*10-8) for hematological phenotypes was obtained in the test of association between MCH and rs1397048 on chromosome 11 near the olfactory receptors, olfactory receptor, family 5, subfamily AP, member 2 (OR5AP2), olfactory receptor, family 5, subfamily AR, member 1 (OR5AR1), olfactory receptor, family 9, subfamily G, member 1(OR9G1) and olfactory receptor, family 9, subfamily G, member 4 (OR9G4). The 25 SNPs with lowest FBAT association test p-values are presented in Additional file 1, Table A1 and Table A2, respectively.
Linkage results
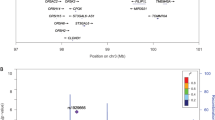
Maximum multipoint LOD scores greater than 2 and the 1.5-LOD support intervals around the maximum LOD scores are presented in Table 4. The highest LOD score for hemostatic factors was 3.3 for factor VII at approximately 15 Mb on chromosome 10. The highest LOD for hematological phenotypes was 3.4 for Hgb at approximately 55 Mb on chromosome 4.
Combining association tests across multiple phenotypes
The top 10 SNPs with most number of p-values < 0.01 and lowest mean p-values are reported in Tables 5 and 6 for platelet aggregation phenotypes and fibrinogen levels respectively. The top ranked SNP for platelet aggregation was rs10500631 on chromosome 11 located near an olfactory gene cluster. The p-values of the GEE association test for ADP-, collagen- and epinephrine-induced platelet aggregation levels with this SNP were all less than 0.01, with average p-value 0.007 over the three tests. The range of the regression coefficients was 0.19–0.24, indicating the effect size was consistently estimated across the three phenotypes.
For fibrinogen, the top ranked SNP was rs4861952 on chromosome 4, which was also listed in the Table 2 as one of the 25 most significantly associated SNPs with hemostatic factors. This SNP was consistently associated with fibrinogen levels across three examination cycles with effect size ranging from -0.28 to -0.17.
Association of SNPs in known candidate genes
100K SNPs residing in or near known candidate genes for hemostatic factors are presented in Table 7. Among the candidate genes for hemostatic factors, no 100K SNP was in or within 60 kb of PLAT. Only SNPs in or near the rest of the candidate genes (F7, FGG, FGA, FGB, ITGB3, SERPINE1 and vWF) are presented. Among all these associations, three reached nominal significance (p-value < 0.05): rs561241 for factor VII, and rs6950982 and rs6956010 for PAI-1.
Among the candidate genes for hematological traits, no 100K SNP was in or within 60 Kb of EPOR, EPO, KLF1, HBA1, HBA2, HBM. For the rest of the candidate genes, associations between hematological phenotypes and 100K SNPs in/near EPB41L2, the beta hemoglobin gene cluster on chromosome 11 (HBB, HBD, HBG1, HBG2, HBE1), and HEBP2 are presented in Table 8. The most significant associations were SNP rs1582055 near EPB41L2 with hematocrit (p = 7.7 × 10-5), Hgb (p = 2.9 × 10-4), and RBCC (p = 3.9 × 10-4); SNP rs4897475 with hematocrit (p = 1.6 × 10-4) and Hgb (p = 6.0 × 10-4).
Discussion
We conducted a GWAS and a genome-wide linkage analyses for hemostatic factors and hematological phenotypes measured in Framingham Heart Study Offspring participants. We identified a highly significant association between factor VII level and SNP rs561241 in complete LD with the F7 SNP rs6046 (Arg353Gln) previously demonstrated to explain about 9% of total phenotypic variation. This association is significant after Bonferroni correction for multiple testing (we used a conservative α = 5 × 10-8), and confirms the strong association at this locus that has previously been reported by us and others. This SNP was also significant (p-value = 3.4 × 10-4) at a nominal α level 0.05 for FBAT and linkage test (LOD = 1.8, p-value = 0.002), but not after Bonferroni correction. That may be explained by the well known fact that FBAT and linkage test are less powerful than population-based association tests.
FBAT lacks power to detect variants that explain small proportion of variance for this study. It is difficult to distinguish true positives from false ones among FBAT results because it was evident that few 100K SNPs explain a large proportion of variance for hemostatic factors or hematological phenotypes. Given that there is no evidence for major population substructure in FHS [33] and there is greater power from use of GEE testing, we emphasize our population-based GEE analysis results in this report. Linkage analyses have the same problem of low power to detect small effects. However, a linkage peak can be caused by loci in linkage but not in LD with the SNPs, or by several loci of small effects in the region. Thus linkage peaks deserve additional attention. For example, we identified a linkage peak on chromosome 10 for multivariate adjusted factor VII. The SNP underneath the peak is rs2400107. However, the GEE association p-value was 0.52. This could occur because rs2400107 was linked but not in LD with the disease locus (loci) under the peak, or because this linkage peak was caused by several loci of small effects, or this peak was a false positive. Therefore, a more careful examination of the association results of SNPs under the linkage peak along with potentially additional genotyping may be needed to confirm the linkage results.
Among the SNPs with top GEE p-values in single phenotype or multiple phenotypes analyses, only a few resided near genes that were known for a likely role in hemostasis and thrombosis and hematological biology. For hemostatic factors, the cis-acting SNP rs561241 near F7 gene was associated with factor. For hematological phenotypes, we identified rs6811964 near PDGFC, platelet derived growth factor-C. It has been shown that PDGFC highly expressed in vascular smooth muscle cells, renal mesangial cells and platelets, and was likely involved in platelet biology [34]. This SNP was found associated with Epi-induced platelet aggregation (P = 10-5, Table 3), with ADP-induced platelet aggregation at nominal significance (P = 0.02), and with collagen-induced platelet aggregation at borderline nominal significance (P = 0.08). Other associations were found with SNPs in genes not clearly related to the phenotypes, or with SNPs that are not in known genes. These associations, together with other findings from this GWAS, must be viewed as hypotheses that warrant further testing in other cohorts.
Although we only summarized results for multivariable adjusted phenotype, we have also conducted linkage and association analyses for age-sex adjusted phenotypes. It is possible that the effects of some loci may be mediated through the covariates included in multivariable adjustment, and thus only associated with age and sex adjusted phenotypes. Among the 52 SNPs that were associated with age and sex adjusted hemostatic factors or hematological phenotypes with a GEE p-value equal or less than 10-5, 28 SNPs had a GEE p-value greater than 10-5 with multivariable adjusted phenotypes. However, no age and sex adjusted GEE p-value for the 28 SNPs reached genome-wide significance (p-value < 5 × 10-8), and no new highly plausible candidate genes resided within 60 Kb of these SNPs. The full disclosure results of all analyses, including the age-sex adjusted analyses, can be viewed at http://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?id=phs000007.
There are some limitations to this study. The participants are Caucasian and thus the results may not be generalizable to other racial groups. The study sample size was relatively small, and as such, we may have insufficient power to detect small effects. To avoid worsening the multiple testing problem, we performed only sex-pooled and not sex-specific analyses. There may be some SNPs that are associated with some phenotypes only in female or male undetected in the current study. The advantages of this study are that we had family data, which enabled us to also apply family-based association tests that are robust to population admixture, and linkage analyses that can detect loci not in LD but in linkage with any 100K SNP. The study subjects were recruited without regarding to their phenotypic values, which makes the analyses of multiple phenotypes possible without the need to correct ascertainment bias.
Finally, compared with studies focused only on SNPs within candidate genes, GWAS approaches are unbiased and as such they have the advantage of detecting novel genes or confirming genes that are not well-known to have an influence on a phenotype. However, since the current GWAS uses only a subset of all the SNPs in HapMap [35], it may miss some genes due to lack of coverage. For the same reason, GWAS data usually are not enough to study a candidate gene comprehensively. To understand the roles played by each SNP in a candidate gene, additional genotyping, and single-SNP and haplotype analyses are needed. A large GWAS involving more than 550,000 SNPs in more than 9000 participants of FHS will be available for analysis later in 2007, providing increased power for detection of smaller effects for the hemostatic and hematological phenotypes.
Conclusion
In summary, we have tested for association and linkage using the Affymetrix 100K SNPs and a set of hemostatic factor and hematological phenotypes. We have confirmed a previously reported association, providing proof of principle (a "positive control") for the GWAS approach. Our results provide a set of hypotheses that warrant testing in additional studies.
Abbreviations
- ADP:
-
adenosine 5'-diphosphate
- bp:
-
base pair(s)
- CHD:
-
coronary heart disease
- CVD:
-
cardiovascular disease
- CHR:
-
chromosome
- EPB41L2:
-
erythrocyte membrane protein band 4.1-like 2
- EPO:
-
erythropoietin
- EPOR:
-
erythropoietin receptor
- FBAT:
-
family-based association test
- FGB:
-
fibrinogen-β
- FGA:
-
fibrinogen-α
- FGG:
-
fibrinogen-γ
- FHS:
-
Framingham Heart Study
- FVII:
-
factor VII
- GAW:
-
Genetic Analysis Workshop
- GEE:
-
generalized estimating equations
- GWAS:
-
genome-wide association study
- HBA1:
-
hemoglobin-α 1
- HBA2:
-
hemoglobin-α 2
- HBB:
-
hemoglobin-β chain complex
- HBD:
-
hemoglobin-δ
- HBE1:
-
hemoglobin-ε 1
- HBG1:
-
hemoglobin-γ A
- HBG2:
-
hemoglobin-γ G
- HBM:
-
hemoglobin-μ
- HCT:
-
hematocrit
- HEBP2:
-
heme binding protein 2
- Hgb:
-
hemoglobin
- ITGB3:
-
integrin beta 3 (platelet glycoprotein IIIa, antigen CD61)
- kb:
-
kilo base pairs (=1000 bp)
- KLF1:
-
Kruppel-like factor 1
- LD:
-
linkage disequilibrium
- MAF:
-
minor allele frequency
- MCV:
-
mean corpuscular volume
- MCH:
-
mean corpuscular hemoglobin
- Mb:
-
mega base pairs (=1000,000 bp)
- OR5AP2:
-
olfactory receptor family 5, subfamily AP, member 2
- OR5AR1:
-
olfactory receptor family 5, subfamily AR, member 1
- OR9G1:
-
olfactory receptor family 9, subfamily G, member 1
- OR9G4:
-
olfactory receptor family 9, subfamily G, member 4
- PAI-1:
-
plasminogen activator inhibitor
- PDGFC:
-
platelet derived growth factor-C
- RBCC:
-
red blood cell count
- SERPINE1:
-
serpin peptidase inhibitor clade E, member 1
- SNP:
-
single nucleotide polymorphism
- tPA:
-
tissue plasminogen activator
- vWF:
-
von Willebrand factor
- WBC:
-
white blood cell.
References
Kannel WB, Wolf PA, Castelli WP, D'Agostino RB: Fibrinogen and risk of cardiovascular disease. The Framingham Study. JAMA. 1987, 258: 1183-1186. 10.1001/jama.258.9.1183.
Danesh J, Collins R, Appleby P, Peto R: Association of Fibrinogen, C-reactive Protein, Albumin, or Leukocyte Count With Coronary Heart Disease: Meta-analyses of Prospective Studies. JAMA. 1998, 279: 1477-1482. 10.1001/jama.279.18.1477.
Wilhelmsen L, Svardsudd K, Korsan-Bengtsen K, Larsson B, Welin L, Tibblin G: Fibrinogen as a risk factor for stroke and myocardial infarction. N Engl J Med. 1984, 311: 501-505.
Johansson L, Jansson JH, Boman K, Nilsson TK, Stegmayr B, Hallmans G: Tissue Plasminogen Activator, Plasminogen Activator Inhibitor-1, and Tissue Plasminogen Activator/Plasminogen Activator Inhibitor-1 Complex as Risk Factors for the Development of a First Stroke. Stroke. 2000, 31: 26-32.
Thogersen AM, Jansson JH, Boman K, Nilsson TK, Weinehall L, Huhtasaari F, Hallmans G: High Plasminogen Activator Inhibitor and Tissue Plasminogen Activator Levels in Plasma Precede a First Acute Myocardial Infarction in Both Men and Women : Evidence for the Fibrinolytic System as an Independent Primary Risk Factor. Circulation. 1998, 98: 2241-2247.
Jansson JH, Nilsson TK, Johnson O: von Willebrand factor in plasma: a novel risk factor for recurrent myocardial infarction and death. Br Heart J. 1991, 66: 351-355.
Lowe GDO, Danesh J, Lewington S, Walker M, Lennon L, Thomson A, Rumley A, Whincup PH: Tissue plasminogen activator antigen and coronary heart disease: Prospective study and meta-analysis. Eur Heart J. 2004, 25: 252-259. 10.1016/j.ehj.2003.11.004.
Meade TW, Mellows S, Brozovic M, Miller GJ, Chakrabarti RR, North WR, Haines AP, Stirling Y, Imeson JD, Thompson SG: Haemostatic function and ischaemic heart disease: principal results of the Northwick Park Heart Study. Lancet. 1986, 2: 533-537. 10.1016/S0140-6736(86)90111-X.
Ono N, Koyama T, Suehiro A, Oku K, Fujikake K, Kakishita E: Influence of atrial fibrillation on coagulo-fibrinolytic markers in patients with cerebral infarction. Int Angiol. 1992, 11: 298-303.
Speiser W, Speiser P, Minar E, Korninger C, Niessner H, Huber K, Schernthaner G, Ehringer H, Lechner K: Activation of coagulation and fibrinolysis in patients with arteriosclerosis: relation to localization of vessel disease and risk factors. Thromb Res. 1990, 59: 77-88. 10.1016/0049-3848(90)90273-F.
Kannel WB, Gordon T, Wolf PA, McNamara P: Hemoglobin and the risk of cerebral infarction: the Framingham Study. Stroke. 1972, 3: 409-420.
O'Donnell CJ, Larson MG, Feng D, Sutherland PA, Lindpaintner K, Myers RH, D'Agostino RA, Levy D, Tofler GH: Genetic and Environmental Contributions to Platelet Aggregation : The Framingham Heart Study. Circulation. 2001, 103: 3051-3056.
Fuster V, Badimon L, Badimon JJ, Chesebro JH: The pathogenesis of coronary artery disease and the acute coronary syndromes (1). N Engl J Med. 1992, 326: 242-250.
Szapary L, Horvath B, Marton Z, Alexy T, Demeter N, Szots M, Klabuzai A, Kesmarky G, Juricskay I, Gaal V, et al: Hemorheological disturbances in patients with chronic cerebrovascular diseases. Clin Hemorheol Microcirc. 2004, 31: 1-9.
Fisher M, Meiselman HJ: Hemorheological factors in cerebral ischemia. Stroke. 1991, 22: 1164-1169.
Kathiresan S, Yang Q, Larson MG, Camargo AL, Tofler GH, Hirschhorn JN, Gabriel SB, O'Donnell CJ: Common Genetic Variation in Five Thrombosis Genes and Relations to Plasma Hemostatic Protein Level and Cardiovascular Disease Risk. Arterioscler Thromb Vasc Biol. 2006, 26: 1405-1412. 10.1161/01.ATV.0000222011.13026.25.
Kathiresan S, Gabriel SB, Yang Q, Lochner AL, Larson MG, Levy D, Tofler GH, Hirschhorn JN, O'Donnell CJ: Comprehensive Survey of Common Genetic Variation at the Plasminogen Activator Inhibitor-1 Locus and Relations to Circulating Plasminogen Activator Inhibitor-1 Levels. Circulation. 2005, 112: 1728-1735. 10.1161/CIRCULATIONAHA.105.547836.
Steinberg MH, Embury SH: Alpha-thalassemia in blacks: genetic and clinical aspects and interactions with the sickle hemoglobin gene. Blood. 1986, 68: 985-990.
Weatherall DJ, Clegg JB: The Thalassaemia Syndromes. 2001, Oxford: Blackwell Science
Adorno EV, Zanette A, Lyra I, Souza CC, Santos LF, Menezes JF, Dupuit MF, Almeida MN, Reis MG, Goncalves MS: The beta-globin gene cluster haplotypes in sickle cell anemia patients from Northeast Brazil: a clinical and molecular view. Hemoglobin. 2004, 28: 267-271. 10.1081/HEM-120040310.
Yang Q, Tofler GH, Cupples LA, Larson MG, Feng D, Lindpaintner K, Levy D, D'Agostino RB, O'Donnell CJ: A genome-wide search for genes affecting circulating fibrinogen levels in the Framingham Heart Study. Thrombosis Research. 2003, 110: 57-64. 10.1016/S0049-3848(03)00288-3.
Bladbjerg EM, de Maat MPM, Christensen K, Bathum L, Jespersen J, Hjelmborg J: Genetic influence on thrombotic risk markers in the elderly – a Danish twin study. Journal of Thrombosis and Haemostasis. 2006, 4: 599-607. 10.1111/j.1538-7836.2005.01778.x.
Lin JP, O'Donnell CJ, Levy D, Cupples LA: Evidence for a gene influencing haematocrit on chromosome 6q23–24: genomewide scan in the Framingham Heart Study. J Med Genet. 2005, 42: 75-79. 10.1136/jmg.2004.021097.
Lin JP, O'Donnell CJ, Li J, Fox CS, Yang Q, Cupples LA: Evidence for linkage of red blood cell size and count : genome-wide scans in the Framingham Heart Study. American Journal of Hematology. 2006
Cupples LA, Arruda HT, Benjamin EJ, D'Agostino RB, Demissie S, DeStefano AL, Dupuis J, Falls K, Fox CS, Gottlieb DJ, Govindaraju DR, Guo CY, Heard-Costa NL, Hwang SJ, Kathiresan S, Kiel DP, Laramie JM, Larson MG, Levy D, Liu CY, Lunetta KL, Mailman MD, Manning AK, Meigs JB, Murabito JM, Newton-Cheh C, O'Connor GT, O'Donnell CJ, Pandey MA, Seshadri S, Vasan RS, Wang ZY, Wilk JB, Wolf PA, Yang Q, Atwood LD: The Framingham Heart Study 100K SNP genome-wide association study resource: Overview of 17 phenotype working group reports. BMC Med Genet. 2007, 8 (Suppl 1): S1-
Rosito GA, D'Agostino RB, Massaro J, Lipinska I, Mittleman MA, Sutherland P, Wilson PW, Levy D, Muller JE, Tofler GH: Association between obesity and a prothrombotic state: the Framingham Offspring Study. Thromb Haemost. 2004, 91: 683-689.
Clauss A: [Rapid physiological coagulation method in determination of fibrinogen]. Acta Haematol. 1957, 17: 237-246.
Born GVR: Aggregation of Blood Platelets by Adenosine Diphosphate and its Reversal. Nature. 1962, 194: 927-929. 10.1038/194927b0.
The principles and technic of blood examination. 1961, Lea & Febiger
Emigh TH: On the Number of Observed Classes from A Multinomial Distribution. Biometrics. 1983, 39: 485-491. 10.2307/2531019.
Liang KY, Zeger SL: Longitudinal Data-Analysis Using Generalized Linear-Models. Biometrika. 1986, 73: 13-22. 10.1093/biomet/73.1.13.
Rabinowitz D, Laird N: A unified approach to adjusting association tests for population admixture with arbitrary pedigree structure and arbitrary missing marker information. Hum Hered. 2000, 50: 211-223. 10.1159/000022918.
Wilk JB, Manning AK, Dupuis J, Cupples LA, Larson MG, Newton-Cheh C, Demissie S, DeStefano AL, Hwang SJ, Liu C, et al: No evidence of major population substructure in the Framingham Heart Study [abstract]. Genet Epidemiol. 2007, 29: 286-
Fang L, Yan Y, Komuves LG, Yonkovich S, Sullivan CM, Stringer B, Galbraith S, Lokker NA, Hwang SS, Nurden P, et al: PDGF C Is A Selective {alpha} Platelet-Derived Growth Factor Receptor Agonist That Is Highly Expressed in Platelet {alpha} Granules and Vascular Smooth Muscle. Arterioscler Thromb Vasc Biol. 2004, 24: 787-792. 10.1161/01.ATV.0000120785.82268.8b.
The International HapMap Project. Nature. 2003, 426: 789-796. 10.1038/nature02168.
Acknowledgements
This work is supported by National Institute of Health/National Heart, Lung & Blood Institute (NHLBI) Contract N01-HC-25195. A portion of the research was conducted using the BU Linux Cluster for Genetic Analysis (LinGA) funded by the NIH NCRR (National Center for Research Resources) Shared Instrumentation grant (1S10RR163736-01A1). We express out gratitude to the Framinghan Heart Study participants, and helpful input from the collaborators: Emelia Benjamin and Martin G. Larson.
This article has been published as part of BMC Medical Genetics Volume 8 Supplement 1, 2007: The Framingham Heart Study 100,000 single nucleotide polymorphisms resource. The full contents of the supplement are available online at http://www.biomedcentral.com/1471-2350/8?issue=S1.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors' contributions
QY took a leading role in study design, results summarization and interpretation, and manuscript drafting. SK contributed to the study design, summarization and interpretation of the results, and the manuscript revision. JPL contributed to the study of hematological phenotypes. GHT contributed to the measurement and study of the hemostatic traits. COD contributed to study design, results summarization and manuscript drafting and revision. All authors read and approved the final manuscript.
Electronic supplementary material
12881_2007_288_MOESM1_ESM.doc
Additional file 1: The 25 SNPs with lowest FBAT association test p-values are presented in Additional data file 1, Table A1 and Table A2, respectively. (DOC 104 KB)
Rights and permissions
Open Access This article is published under license to BioMed Central Ltd. This is an Open Access article is distributed under the terms of the Creative Commons Attribution License ( https://creativecommons.org/licenses/by/2.0 ), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Yang, Q., Kathiresan, S., Lin, JP. et al. Genome-wide association and linkage analyses of hemostatic factors and hematological phenotypes in the Framingham Heart Study. BMC Med Genet 8 (Suppl 1), S12 (2007). https://doi.org/10.1186/1471-2350-8-S1-S12
Published:
DOI: https://doi.org/10.1186/1471-2350-8-S1-S12