Abstract
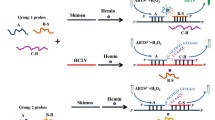
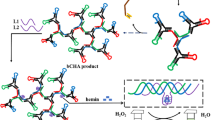
African swine fever virus (ASFV) causes hemorrhagic infectious disease in pigs with a fatality rate of nearly 100%. In this study, we developed a visual strand exchange amplification detection assay for ASFV. In the presence of ASFV, DNA amplification products containing multimeric G-quadruplex sequences were amplified by strand exchange amplification. These G-quadruplexes, assembled with hemin to form DNAzyme, displayed enhanced significant “turned-on” colorimetric signals to indicate detection results. The results showed that dimeric DNAzyme had the best visualization effect. Under the optimal reaction parameters, there was a linear relationship between the absorbance of the reaction solution at 417 nm and the logarithm of ASFV concentration ranged from 1 × 101 to 1 × 103 copies/μL, and the detection limit was 2.7 copies/μL. We hoped this visual assay could be helpful in the rapid and sensitive detection of ASFV, and the results of multimeric G-quadruplex/hemin DNAzyme could be helpful for the development of better visual detection assays.
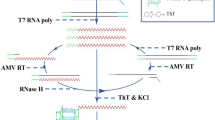
Graphical abstract







Similar content being viewed by others
References
S. Liu, Y. Luo, Y. Wang, S. Li, Z. Zhao, Y. Bi, J. Sun, R. Peng, H. Song, D. Zhu, Y. Sun, S. Li, L. Zhang, W. Wang, Y. Sun, J. Qi, J. Yan, Y. Shi, X. Zhang, G. Gao, Cell Host Microbe 26(6), 836–843 (2019). https://doi.org/10.1016/j.chom.2019.11.004
S. Cappai, S. Rolesu, F. Feliziani, P. Desini, V. Guberti, F. Loi, Vaccines (Basel) 8(4), 723 (2020). https://doi.org/10.3390/vaccines8040723
J.N. Hakizimana, L. Nyabongo, J.B. Ntirandekura, C. Yona, D. Ntakirutimana, O. Kamana, H. Nauwynck, G. Misinzo, Front. Vet. Sci. 7, 578474 (2020). https://doi.org/10.3389/fvets.2020.578474
L. Bosch-Camós, E. López, F. Rodriguez, Porcine. Health Manage. 6, 17 (2020). https://doi.org/10.1186/s40813-020-00154-2
Z. Pejsak, M. Truszczyński, K. Tarasiukl, Med. Weter. 75, 6186–2019 (2019). https://doi.org/10.21521/mw.6186
K. Wu, J. Liu, L. Wang, S. Fan, Z. Li, Y. Li, L. Yi, H. Ding, M. Zhao, J. Chen, Vaccines (Basel) 8(3), 531 (2020). https://doi.org/10.3390/vaccines8030531
B. Pepin, T. Williams, D. Polson, P. Gauger, S. Dee, Transbound Emerg. Dis. 68(4), 2239–2249 (2020). https://doi.org/10.1111/tbed.13876
S.S. Patil, K.P. Suresh, V. Vashist, A. Prajapati, B. Pattnaik, P. Roy, Vet. World 13(10), 2275–2285 (2020). https://doi.org/10.14202/vetworld.2020.2275-2285
D.H. Tran, H.T. Tran, U.P. Le, X.D. Vu, T.B.N. Trinh, H.D.K. Do, V.T. Than, L.M. Bui, V.V. Vu, T.L. Nguyen, H.T.T. Phung, V.P. Le, Transbound. Emerg. Dis. 68(4), 2595–2602 (2020). https://doi.org/10.1111/tbed.13879
M. Juszkiewicz, M. Walczak, N. Mazur-Panasiuk, G. Woźniakowski, Pathogens 9(11), 878 (2020). https://doi.org/10.3390/pathogens9110878
X. Wang, S. He, N. Zhao, X. Liu, Y. Cao, G. Zhang, G. Wang, C. Guo, BMC Microbiol. 20(1), 282 (2020). https://doi.org/10.1186/s12866-020-01966-6
Z. Liu, C. Yao, Y. Wang, C. Yang, Anal. Methods UK 10, 848–854 (2018). https://doi.org/10.1039/C7AY02908J
J. Xu, Y. Hu, J. Guo, Y. Yang, J. Qiu, X. Li, Z. Xin, Food Anal. Methods 12, 422–430 (2019). https://doi.org/10.1007/s12161-018-1373-0
Z. Liu, Y. Wang, L. Li, J. Li, Y. Yuan, Anal. Methods 11(15), 2082–2088 (2019). https://doi.org/10.1039/C8AY02798F
Y. Wang, X. Li, D. Xi, X. Wang, RSC Adv. 9(64), 37144–37147 (2019). https://doi.org/10.1039/C9RA05709A
C. Wu, G. Gao, K. Zhai, L. Xu, D. Zhang, Food Chem. 331, 127208 (2020). https://doi.org/10.1016/j.foodchem.2020.127208
R. Zou, Y. Ma, C. Li, F. Zhang, C. Chen, C. Cai, Microchem. J. 156, 104764 (2020). https://doi.org/10.1016/j.microc.2020.104764
D. Ji, H. Meng, J. Ge, L. Zhang, H. Wang, D. Bai, J. Li, L. Qu, Z. Li, Microchim. Acta 184, 1–8 (2017). https://doi.org/10.1007/s00604-017-2343-8
R.M. Kong, L. Ma, X. Han, C. Ma, F. Qu, L. Xia, Spectrochim. Acta A Mol. Biomol. Spectrosc. 228, 117855 (2020). https://doi.org/10.1016/j.saa.2019.117855
F. Zhao, H. Zhang, J. Zheng, Sensor Actuat. B Chem. 327, 128898 (2021). https://doi.org/10.1016/j.snb.2020.128898
R. Adeoye, D. Osalaye, T.K. Ralebitso-Senior, A. Boddis, A. Reid, A. Fatokun, A. Powell, S. Malomo, F. Olorunniji, Catalysts 9, 613 (2019). https://doi.org/10.3390/catal9070613
C. Shi, F. Shang, M. Zhou, P. Zhang, Y. Wang, C. Ma, Chem. Commun. 52, 11551–11554 (2016). https://doi.org/10.1039/C6CC05906F
C. Yang, Y. Li, J. Deng, M. Li, C. Ma, C. Shi, Anal. Bioanal. Chem. 412(30), 8391–8399 (2020). https://doi.org/10.1007/s00216-020-02977-y
J. Chen, Y. Zhang, M. Cheng, J.L. Mergny, Q. Lin, J. Zhou, H. Ju, Mikrochim. Acta 186(12), 786 (2019). https://doi.org/10.1007/s00604-019-3950-3
G.J. Hafner, I.C. Yang, L.C. Wolter, M.R. Stafford, P.M. Giffard, Biotechniques 30(4), 852–867 (2001). https://doi.org/10.2144/01304rr03
Y. Wang, J. Dai, Y. Liu, J. Yang, Q. Hou, Y. Ou, Y. Ding, B. Ma, H. Chen, M. Li, Y. Sun, H. Zheng, K. Zhang, A.K. Wubshet, A.D. Zaberezhny, T.I. Aliper, K. Tarasiuk, Z. Pejsak, Z. Liu, Y. Zhang, J. Zhang, Front. Microbiol. 12(516), 609821 (2021). https://doi.org/10.3389/fmicb.2021.609821
R. Li, Q. Liu, Y. Jin, B. Li, Microchim. Acta 187, 139 (2020). https://doi.org/10.1007/s00604-019-4093-2
L. Ganges, H.R. Crooke, J.A. Bohórquez, A. Postel, Y. Sakoda, P. Becher, N. Ruggli, Virus Res. 289, 198151 (2020). https://doi.org/10.1016/j.virusres.2020.198151
J. Chaudhari, H.L.X. Vu, Viruses 12(11), 198151 (2020). https://doi.org/10.3390/v12111245
E. Domingo, E. Baranowski, C. Escarmís, F. Sobrino, Comp. Immunol. Microbiol. Infect. Dis. 25(5–6), 297–308 (2002). https://doi.org/10.1016/s0147-9571(02)00027-9
G. Wong, J. Lu, W. Zhang, G.F. Gao, Emerg. Microbes Infect. 8(1), 150–154 (2019). https://doi.org/10.1080/22221751.2018.1563459
J. Fernández-Pinero, C. Gallardo, M. Elizalde, A. Robles, C. Gómez, R. Bishop, L. Heath, E. Couacy-Hymann, F.O. Fasina, V. Pelayo, A. Soler, M. Arias, Transbound Emerg. Dis. 60(1), 48–58 (2013). https://doi.org/10.1111/j.1865-1682.2012.01317.x
Y. Gao, X.-Y. Meng, H. Zhang, Y. Luo, Y. Sun, Y. Li, M. Abid, H.-J. Qiu, Sensor Actuat. B Chem. 274, 304–309 (2018). https://doi.org/10.1016/j.snb.2018.07.164
G. Wozniakowski, M. Fraczyk, A. Kowalczyk, M. Pomorska-Mol, K. Niemczuk, Z. Pejsak, Sci. Rep UK 7, 42903 (2017). https://doi.org/10.1038/srep42903
Z. Wang, W. Yu, R. Xie, S. Yang, A. Chen, Anal. Bioanal. Chem. 413(18), 4665–4672 (2021). https://doi.org/10.1007/s00216-021-03408-2
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 31972158).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wu, X., Chen, Q., Huang, Y. et al. Signal-enhanced visual strand exchange amplification detection of African swine fever virus by the introduction of multimeric G-quadruplex/hemin DNAzyme. ANAL. SCI. 38, 675–682 (2022). https://doi.org/10.1007/s44211-022-00087-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s44211-022-00087-6