Abstract
Purpose
Consanguinity increases the likelihood of the inheritance of homozygous pathogenic alleles which may predispose to rare autosomal recessive disorders. Here we discuss the role of consanguinity in informing inherited disease with a focus on rare diseases.
Methods
We reviewed the literature concerning the impact of consanguinity on human diseases and chose examples to illustrate the most important themes.
Results
Consanguinity rates vary hugely between different populations influencing the prevalence of rare autosomal recessive diseases. Some founder genetic variants leading to human disease are specific for a single country, or a specific ethnic or geographic group while others are shared more widely. Inherited diseases of known molecular genetic etiology are characterized by their genotype and phenotype but many exhibit marked heterogeneity which may be population dependent. Increased rates of consanguinity are associated with rare autosomal recessive inherited diseases and can lead to more than one human genetic disease in affected individuals leading to complex and overlapping phenotypes. Next-generation sequencing strategies allow new insights into these cases. In contrast, the impact of consanguinity on malignancies and common multifactorial diseases is less predictable and needs further exploration.
Conclusions
High rates of consanguinity remain prevalent in certain populations and lead to an increased burden of rare autosomal recessive inherited diseases. Strategies to reduce consanguinity are needed to reduce these disease consequences and will require global improvements in education, social, and economic conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
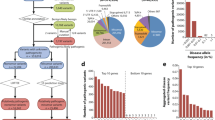
The term consanguinity literally means “shared blood.” Consanguineous marriage is defined as a marriage between individuals who are closely related and is associated with an increased risk of autosomal recessive genetic diseases in the offspring of these parents [4]. More than 1.2 billion of the current population in the world are reported to practice consanguineous marriage. Consanguinity is often observed in poorly educated populations [22], and improving education allows greater independence and enables more informed life decisions. Consanguineous marriages are known to be practiced for many generations in many communities all around the world [18]. The most common form of consanguinity is between first cousin marriages. In this scenario, spouses share 1/8th of their genes inherited from their ancestors and their progeny are typically homozygous for 1/16th of all loci [19]. The prevalence of consanguinity varies from country to country (Fig. 1) and is shown to be influenced by multiple different factors such as religion, ethnicity, demography, geography (rural or urban areas), education, and economic factors [19]. Consanguineous marriages account for less than 20% to more than 50% of all marriages in Arab countries (Table 1), which span the region from North Africa to the Middle East and western Asia [12].
World map showing consanguineous marriages. Consanguineous marriages are defined here as second-degree cousins or closer, and frequency is shown in percentage (%). Image adapted from https://commons.m.wikimedia.org/wiki/File:Global_prevalence_of_consanguinity.svg and licensed under the Creative Commons Share Alike 3.0 unported
Consanguinity, unsurprisingly, has been identified as a risk for congenital malformation and major developmental medical conditions. These malformations include diverse phenotypes such as polydactyly, spinocerebellar degeneration, neural tube defects, anencephaly, and encephalocele [38]. Here, we discuss the global distribution of consanguinity and the impact of consanguinity on a wide variety of different diseases using examples of acute lymphoblastic leukemia, breast cancer, obesity, and rare genetic diseases to illustrate key messages. We also show how modern genetic sequencing techniques can inform the genetics of consanguinity with the identification of novel disease alleles, hypomorphic alleles, and founder alleles.
Global distribution of consanguinity
Recent data indicate that approximately 10.4% of the total population in the world is reported to be married to biological relatives [12]. In North Africa, West Asia, and South India, marriages to biological relatives are culturally favored and constitute 20–50% of all marriages [47]. In Qatar, it is reported that the rate of consanguinity is approximately 54% [8]. In Saudi Arabia, the rate is between 29.7 and 56% [47, 48]. In Libya, the interfamilial marriage rate was 37.6% in the city of Bengazi [1], and a study in Mauritania estimated a consanguinity rate of 47.2% [21]. In Pakistan, the rate of consanguineous rate is reported to be over 60% of marriages [43].
In European countries, South America, and Australia, the interfamily marriage rate is in comparison low [25]. In North America and Australia, the interfamily marriage is approximately 1%; in Europe, it is approximately 1.5% but depends on local geography and social conditions [27].
Consanguinity and rates of childhood malformations
The rates of early childhood malformations have been correlated with rates of consanguinity [19]. In addition, consanguineous marriage is shown to have a higher level of reproductive loss, risk of abortion, and neonatal or postnatal death [38]. However, in consanguineous populations overall there may be selection against severe recessive diseases. Many recessive genetic diseases are not compatible with life and reproduction, leading to a counter-selection of these pathogenic variants in the populations with ancient practices of consanguinity.
Consanguinity and incidence of cancer
Many studies from different research groups have indicated that rates of consanguinity have little or no effect on the incidence of cancers. Bener et al. [10] showed that although the rate of consanguinity in Qatar was high, it had no effect on the incidence of cancers overall [10]. However, at a tissue-specific level, an increase in risk for leukemia and lymphoma and colorectal and prostate cancer was shown while a reduction in breast, skin, thyroid, and female genital cancers was noted [10].
Breast cancer is the most common type of cancer in adult females. The majority of cases are sporadic, but 5–10% are reported to be inherited, with pathogenic variants in BRCA1 and BRCA2 accounting for the majority of these cases. In family genetic studies in Morocco, using next-generation sequencing (NGS) technologies, four heterozygous pathogenic variant genotypes were found: BRCA1 c.212insA and c.3453delT and BRCA2 c.1310_1313delAAGA and c.723insG [23]. The BRCA1 c.3453delT allele was novel and is likely to be a local founder allele, prompting better characterization of population-specific alleles. In studies from Arabian countries, consanguinity has been shown to be protective against breast cancer [9, 15, 28] and this may be in part due to the fact that BRCA1/BRCA2 deleterious variants are lethal in their homozygous state and are outbred from the population. There may also be an increased carrier rate of protective alleles, which may have an increased effect if present homozygously [11].
In contrast, much rarer oncogene pathogenic variants may be revealed in consanguineous populations. Ripperger et al. reported a case with constitutional mismatch repair deficiency caused by a novel MSH6 pathogenic variant leading to a T-cell lymphoma and colonic adenocarcinoma [37]. The constitutional mismatch repair deficiency syndrome (CMMRD), is an example of a rare recessive inherited cancer syndrome with a broad tumor spectrum including hematological malignancies, brain tumors, and colon cancer in childhood and adolescence. Baris et al. found a consanguineous Bedouin family, a homozygous MSH6 pathogenic variant (c.3603_3606delAGTG) [6].
Consanguinity and obesity
Obesity is known to be a risk factor for many different diseases including cardiovascular disease, insulin resistance, and type 2 diabetes mellitus. Polymorphisms in the ACE gene have been implicated in different metabolic disorders, including obesity. A recent study investigated genetic associations in the offspring of first cousins and found an association of the ACE II polymorphism with obesity in the Saudi population [4]. Also, Alharbi et al. noted that while screening for obesity in children from consanguineous parents they noted that adolescents and adults were more prone (three times more likely) to develop obesity [3]. The exact molecular mechanisms have not been explored but metabolic pathways that regulate obesity are influenced by genetic background [45], as well as environmental factors [42]. In outbred populations, only 2–5% of obesity is secondary to monogenic disorders. Interestingly, in a Pakistani inbred population pathogenic variants in monogenic genes LEP, LEPR, and MC4R were able to explain 30% of severe childhood obesity. The genetics of common obesity is more complex but studies have shown that genes associated with monogenic causes of obesity (LEPR, POMC, MC4R, BDNF, SH2B1, and PCSK1) [29, 31, 41, 49] are enriched for more common alleles in obese patients from the general population. Therefore, variants in the same genes are having different penetrance and consanguineous populations may be enriched for both rare and common genetic variants contributing to an overall increase in obesity [40].
Consanguinity and rare genetic diseases
Rare diseases are by definition those that affect a minor proportion of the population. A prevalence of <0.05% is considered to be a rare disease by the European Union, while in the USA, a disorder affecting fewer than 200,000 people is considered rare (roughly 0.086%). Rare autosomal recessive disorders are known to be increased with consanguineous parents [33]. Cockayne syndrome (CS) is a rare autosomal recessive genetic disease caused by pathogenic variants in ERCC6 or ERCC8. CS is characterized by psychomotor retardation, cerebral atrophy, microcephaly, mental retardation, sensorineural hearing loss, premature aging, kyphosis, ankyloses, and optic atrophy [51]. In a Tunisian patient from a consanguineous family, a novel homozygous variant in ERCC6 (c.3156dup; p.Arg1053Thr*8) was identified [51]. Similarly, in a consanguineous family from Jordan with a severe CS phenotype a novel frameshift ERCC6 variant (c.2911_2915del5Ins9; p.Lys971Tyrfs*14)) was found. Such findings of a rare disease diagnosis with novel pathogenic homozygous alleles in inbred populations are frequent. Studying rare diseases in families allows numerous other opportunities for genetic discoveries. Modern NGS, such as targeted panel sequencing, whole exome sequencing, and whole genome sequencing offers huge potential for molecular genetic diagnostics in these families [44]. What is interesting is that homozygous alleles predicted to be benign, such as synonymous changes within coding regions can be given disease pathogenicity if the variant is rare, segregates with disease phenotype, and is investigated at the transcriptomic level. An interesting example of such a finding is the identification of a synonymous NPHP3 allele in a consanguineous Omani family with a ciliopathy syndrome phenotype (hepatorenal fibrocystic kidney and liver disease) [32]. The NPHP3 variant (c.2805C>T; p.Gly935Gly) was initially filtered out as it was predicted to be non-pathogenic. However, the allele was exceedingly rare, within a large region of homozygosity by descent, and predicted to be pathogenic by in silico splicing tools. The allele was segregated with the disease phenotype and the identical allele was found in 4 other cases with similar phenotypes. Finally, abnormal splicing secondary to this allele was shown using RT-PCR [32]. As NGS sequencing moves from exomes to genomes there will be opportunities to identify and determine the pathogenicity of rare deep intronic alleles that may be driving rare disease phenotype. An example of this is the identification of a deep intronic allele in PKHD1 (c.8798–459C>A) leading to an antenatal presentation of ARPKD in two fetuses in a consanguineous Chinese family [14]. Consanguineous populations allow these genetic studies to be driven forward.
Founder alleles may also be identified by studying specific genetic disorders within specific inbred populations. An example is the identification of a GBA c.1246G>A; p.Gly377Ser homozygous missense variant in patients with Gaucher’s disease from Northeastern Brazil. The original population from Portugal was of Sephardic Jewish extraction and settled in this location in the 1700s. The combination of this founder allele and high rates of consanguinity contributed to a high prevalence of Gaucher’s disease in this population [13]. Other such examples are seen in other inbred populations, including the identification of a founder allele in AGL, leading to Glycogen storage disease type IIIa in Inuit populations [39].
Awareness of rare disease alleles within specific populations is now growing and premarital genetic screening for such alleles is becoming more frequent. Ashkenazi Jews have an exceptionally high carrier frequency for a range of genetic disorders named “Jewish Genetic Disorders” which include the lysosomal storage disorder Tay Sachs disease [5]. Carrier screening for these disorders is now recommended and allows informed decisions about marriage and reproduction to be taken. A recent report performed carrier screening of forty disease-causing variants in individuals from Syrian and Iranian Jewish ancestry and compared these to Ashkenazi Jewish carrier frequency rates [52]. Over 8% of the study population were carriers for at least one pathogenic variant, supporting the importance of premarital genetic screening in order to reduce the incidence of autosomal recessive disease. Such screening programs need to be adopted by other at-risk populations. In a Saudi Arabian population, the carrier frequency of variants in 35 genes associated with the most prevalent disorders was recently performed [2]. As an example, an allele, in MPL (c.317C>T; p.Pro106Leu) which causes thrombocytopenia was seen in 2.46% of the population compared to 0.01% in the gnomAD database. There are clearly issues regarding economic ethical and social implications for such screening programs and compliance of genetic screening programs can be low [46].
Within consanguineous families, the occurrence of more than one rare genetic disorder is also more frequently seen [24]. There are numerous reports of more than one rare homozygous disease-causing variant giving a combination of disease phenotypes. For example, a Chinese patient had both Wilson disease and retinitis pigmentosa and homozygous pathogenic variants were identified in ATP7B and CNGA1 accounting for the two phenotypes respectively [50]. Where concurrent inherited genetic disorders lead to overlapping phenotypes there can be diagnostic confusion. Perrault syndrome is a disorder characterized by primary ovarian insufficiency in females and sensorineural deafness in males and females. Whole exome sequencing in a consanguineous family with six deaf individuals, with the proband also having primary ovarian insufficiency, identified a pathogenic variant in CLDN14 which explained the sensorineural deafness phenotype and a SGO2 homozygous pathogenic variant explaining the concurrent ovarian insufficient [20]. Both deafness and primary ovarian insufficiency are genetically heterogeneous, and the variants were likely acting independently to produce a blended phenotype suggestive of Perrault syndrome. Caution therefore needs to be taken when researchers widen phenotypic spectra of monogenic disorders in consanguineous individuals without excluding a second homozygous disease-causing allele contributing to the disease phenotype.
A further pitfall of the investigation of a rare disease in consanguineous families is the transmission of two alleles (in heterozygous or homozygous state) within the same gene within the same family in such a way that it mimics autosomal dominant inheritance patterns. A consanguineous family of 13 individuals with variable features of Alport syndrome (including hematuria, proteinuria, and kidney failure) appeared to have a male-to-male transmission of disease pattern suggesting autosomal dominant inheritance. Genetic analysis however showed a mixture of homozygous and compound heterozygous alleles producing this pseudo-dominant transmission pattern [30]. This case is a good example of how assuming identity by descent in consanguineous families can be misleading.
NGS approaches in consanguineous families are a useful way of identifying homozygous hypomorphic alleles. In autosomal recessive diseases, these alleles typically give milder phenotypes when homozygous and more severe disease phenotypes when in trans with a heterozygous deleterious allele. Such variants have been reported in TMEM67 leading to more limited liver and kidney phenotypes rather than the embryonic lethal Meckel syndrome [34]. Homozygous hypomorph alleles in autosomal dominant diseases can be identified also, which provide exceptional cases for studying disease pathogenicity, such as the identification of a homozygous UMOD allele in a Pakistani family. These families allow a direct comparison of heterozygote and homozygote allele carriers in order to unravel gene dosage effects [16].
Whole exome and whole genome sequencing to investigate consanguineous families
With the advent of affordable NGS approaches, the use of whole exome and whole genome sequencing for the investigation of rare diseases and cancers is rapidly becoming the first line. Individual families or large cohorts of families with shared phenotypes are subject to exome or genome sequencing and results yield high diagnostic rates, add to the number of disease-causing alleles, and inform global genetics projects. Some care does need to be given before assigning pathogenicity to genetic variants in rare diseases. An example of this was the identification of a homozygous variant in CCDC28B as a potential novel genetic cause of Joubert syndrome [36]. However, further analysis of this variant showed that it was not ultra-rare and had been seen in its homozygous state in control samples from a wide range of ethnicities [7]. A similar example was the initially reported findings of a homozygous c.428delG variant in KIAA0586 in patients with Joubert syndrome. However, careful segregation and RNA studies, alongside population frequency data demonstrated that this allele on its own was not pathogenic [35].
In countries with limited resources, singleton whole exome sequencing has been advocated as a first-tier diagnostic test to limit costs [26]. This approach may be suitable to detect known alleles in known genes, but without segregation of alleles some caution, given the examples above, needs to be given to using this approach for gene discovery in consanguineous families, where homozygous alleles may be numerous and rare but not necessarily pathogenic.
Consanguinity and education
There are numerous reports correlating poor levels of education and consanguinity [17]. Women with low levels of education are more likely to be in a consanguineous marriage. Improving the education of women will allow more informed decisions based on the huge evidence base of the adverse health effects on their children resulting from a consanguineous marriage. However, there is also evidence that deep-rooted social and cultural beliefs and personal preferences outweigh improvements in education [17]. The modern advent of genetic screening and awareness of certain risk alleles within specific inbred populations allows opportunities for positive health care interventions such as premarital screening to reduce risks of inherited diseases.
Conclusions
The effects of consanguinity on health and disease are being increasingly recognized. We have used examples from cancer diseases, obesity, and rare inherited diseases to define how the effects of consanguinity need to be carefully considered. NGS approaches in consanguineous families with both common and rare disease has allowed many new gene disease discoveries. Such technologies make it much more accessible to investigate consanguineous families for inherited diseases and predisposition to other disorders such as cancers. It is important to increase knowledge and public awareness regarding the risks of consanguinity and worldwide education programs may help with this. Patients, families, and their physicians should actively engage into research on the relationship between consanguinity and disease through a multidisciplinary approach.
Availability of data and materials
Data sharing is not applicable to this article as no datasets were generated or analyszed during the current study.
References
Abudejaja AH, Khan MA, Singh R, Toweir AA, Narayanappa M, Gupta BS, Umer S. Experience of a family clinic at Benghazi, Libya, and sociomedical aspects of its catchment population. Fam Pract. 1987;4(1):19–26.
Aleissa M, Aloraini T, Alsubaie LF, Hassoun M, Abdulrahman G, Swaid A, Eyaid WA, Mutairi FA, Ababneh F, Alfadhel M, Alfares A. Common disease-associated gene variants in a Saudi Arabian population. Ann Saudi Med. 2022;42(1):29–35.
Alharbi KK, Al-Sheikh YA, Alsaadi MM, Mani B, Udayaraja GK, Kohailan M, Ali Khan I. Screening for obesity in the offspring of first-cousin consanguineous couples: a phase-I study in Saudi Arabia. Saudi J Biol Sci. 2020;27(1):242–6.
Alshammary AF, Khan IA. Screening of obese offspring of first-cousin consanguineous subjects for the angiotensin-converting enzyme gene with a 287-bp Alu sequence. J Obes Metab Syndr. 2021;30(1):63–71.
Arjunan A, Litwack K, Collins N, Charrow J. Carrier screening in the era of expanding genetic technology. Genet Med. 2016;18(12):1214–7.
Baris HN, Barnes-Kedar I, Toledano H, Halpern M, Hershkovitz D, Lossos A, Lerer I, Peretz T, Kariv R, Cohen S, Half EE, Magal N, Drasinover V, Wimmer K, Goldberg Y, Bercovich D, Levi Z. Constitutional mismatch repair deficiency in Israel: high proportion of founder mutations in MMR genes and consanguinity. Pediatr Blood Cancer. 2016;63(3):418–27.
Barroso-Gil M, Powell L, Sayer JA. Re: Clinical and molecular diagnosis of Joubert syndrome and related disorders. Pediatr Neurol. 2020;112:10.
Bener A, Alali KA. Consanguineous marriage in a newly developed country: the Qatari population. J Biosoc Sci. 2006;38(2):239–46.
Bener A, Ayoubi HR, Ali AI, Al-Kubaisi A, Al-Sulaiti H. Does consanguinity lead to decreased incidence of breast cancer? Cancer Epidemiol. 2010;34(4):413–8.
Bener A, El Ayoubi HR, Chouchane L, Ali AI, Al-Kubaisi A, Al-Sulaiti H, Teebi AS. Impact of consanguinity on cancer in a highly endogamous population. Asian Pac J Cancer Prev. 2009;10(1):35–40.
Bhinder MA, Sadia H, Mahmood N, Qasim M, Hussain Z, Rashid MM, Zahoor MY, Bhatti R, Shehzad W, Waryah AM, Jahan S. Consanguinity: a blessing or menace at population level? Ann Hum Genet. 2019;83(4):214–9.
Bittles AH, Black ML. Evolution in health and medicine Sackler colloquium: consanguinity, human evolution, and complex diseases. Proc Natl Acad Sci U S A. 2010;107 Suppl 1(Suppl 1):1779–86.
Chaves RG, Pereira Lda V, de Araújo FT, Rozenberg R, Carvalho MD, Coelho JC, Michelin-Tirelli K, Chaves Mde F, Cavalcanti GB Jr. Consanguinity and founder effect for Gaucher disease mutation G377S in a population from Tabuleiro do Norte, Northeastern Brazil. Clin Genet. 2015;88(4):391–5.
Chen J, Ma N, Zhao X, Li W, Zhang Q, Yuan S, Tan YQ, Lu G, Lin G, Du J. A rare deep intronic mutation of PKHD1 gene, c.8798-459 C > A, causes autosomal recessive polycystic kidney disease by pseudoexon activation. J Hum Genet. 2019;64(3):207–14.
Denic S, Bener A, Sabri S, Khatib F, Milenkovic J. Parental consanguinity and risk of breast cancer: a population-based case-control study. Med Sci Monit. 2005;11(9):Cr415–9.
Edwards N, Olinger E, Adam J, Kelly M, Schiano G, Ramsbottom SA, Sandford R, Devuyst O, Sayer JA. A novel homozygous UMOD mutation reveals gene dosage effects on uromodulin processing and urinary excretion. Nephrol Dial Transplant. 2017;32(12):1994–9.
El Goundali K, Chebabe M, Zahra Laamiri F, Hilali A. The determinants of consanguineous marriages among the Arab population: a systematic review. Iran J Public Health. 2022;51(2):253–65.
Fareed M, Afzal M. Evidence of inbreeding depression on height, weight, and body mass index: a population-based child cohort study. Am J Hum Biol. 2014;26(6):784–95.
Fareed M, Afzal M. Genetics of consanguinity and inbreeding in health and disease. Ann Hum Biol. 2017;44(2):99–107.
Faridi R, Rehman AU, Morell RJ, Friedman PL, Demain L, Zahra S, Khan AA, Tohlob D, Assir MZ, Beaman G, Khan SN, Newman WG, Riazuddin S, Friedman TB. Mutations of SGO2 and CLDN14 collectively cause coincidental Perrault syndrome. Clin Genet. 2017;91(2):328–32.
Hammami A, Elgazzeh M, Chalbi N, Mansour BA. Endogamy and consanguinity in Mauritania. Tunis Med. 2005;83(1):38–42.
Hussain R, Bittles AH. Sociodemographic correlates of consanguineous marriage in the Muslim population of India. J Biosoc Sci. 2000;32(4):433–42.
Jouali F, Laarabi FZ, Marchoudi N, Ratbi I, Elalaoui SC, Rhaissi H, Fekkak J, Sefiani A. First application of next-generation sequencing in Moroccan breast/ovarian cancer families and report of a novel frameshift mutation of the BRCA1 gene. Oncol Lett. 2016;12(2):1192–6.
Lal D, Neubauer BA, Toliat MR, Altmüller J, Thiele H, Nürnberg P, Kamrath C, Schänzer A, Sander T, Hahn A, Nothnagel M. Increased probability of co-occurrence of two rare diseases in consanguineous families and resolution of a complex phenotype by next generation sequencing. PLoS One. 2016;11(1):e0146040.
Liascovich R, Rittler M, Castilla EE. Consanguinity in South America: demographic aspects. Hum Hered. 2001;51(1-2):27–34.
Masri AT, Oweis L, Qudah AA, El-Shanti H. Congenital muscle dystrophies: role of singleton whole exome sequencing in countries with limited resources. Clin Neurol Neurosurg. 2022;217:107271.
McCullough JM, O’Rourke DH. Geographic distribution of consanguinity in Europe. Ann Hum Biol. 1986;13(4):359–67.
Medimegh I, Troudi W, Omrane I, Ayari H, Uhrhummer N, Majoul H, Benayed F, Mezlini A, Bignon YJ, Sibille C, Elgaaied AB. Consanguinity protecting effect against breast cancer among Tunisian women: analysis of BRCA1 haplotypes. Asian Pac J Cancer Prev. 2015;16(9):4051–5.
Meyre D, Delplanque J, Chèvre JC, Lecoeur C, Lobbens S, Gallina S, Durand E, Vatin V, Degraeve F, Proença C, Gaget S, Körner A, Kovacs P, Kiess W, Tichet J, Marre M, Hartikainen AL, Horber F, Potoczna N, Hercberg S, Levy-Marchal C, Pattou F, Heude B, Tauber M, McCarthy MI, Blakemore AI, Montpetit A, Polychronakos C, Weill J, Coin LJ, Asher J, Elliott P, Järvelin MR, Visvikis-Siest S, Balkau B, Sladek R, Balding D, Walley A, Dina C, Froguel P. Genome-wide association study for early-onset and morbid adult obesity identifies three new risk loci in European populations. Nat Genet. 2009;41(2):157–9.
Mohamed M, Tellez J, Bergmann C, Gale DP, Sayer JA, Olinger E. Pseudodominant Alport syndrome caused by pathogenic homozygous and compound heterozygous COL4A3 splicing variants. Ann Hum Genet. 2022;86(3):145–52.
Nead KT, Li A, Wehner MR, Neupane B, Gustafsson S, Butterworth A, Engert JC, Davis AD, Hegele RA, Miller R, den Hoed M, Khaw KT, Kilpeläinen TO, Wareham N, Edwards TL, Hallmans G, Varga TV, Kardia SL, Smith JA, Zhao W, Faul JD, Weir D, Mi J, Xi B, Quinteros SC, Cooper C, Sayer AA, Jameson K, Grøntved A, Fornage M, Sidney S, Hanis CL, Highland HM, Häring HU, Heni M, Lasky-Su J, Weiss ST, Gerhard GS, Still C, Melka MM, Pausova Z, Paus T, Grant SF, Hakonarson H, Price RA, Wang K, Scherag A, Hebebrand J, Hinney A, Franks PW, Frayling TM, McCarthy MI, Hirschhorn JN, Loos RJ, Ingelsson E, Gerstein HC, Yusuf S, Beyene J, Anand SS, Meyre D. Contribution of common non-synonymous variants in PCSK1 to body mass index variation and risk of obesity: a systematic review and meta-analysis with evidence from up to 331 175 individuals. Hum Mol Genet. 2015;24(12):3582–94.
Olinger E, Alawi IA, Al Riyami MS, Salmi IA, Molinari E, Faqeih EA, Al-Hamed MH, Barroso-Gil M, Powell L, Al-Hussaini AA, Rahim KA, Almontashiri NAM, Miles C, Shril S, Hildebrandt F, G. E. R. Consortium, Wilson IJ, Sayer JA. A discarded synonymous variant in NPHP3 explains nephronophthisis and congenital hepatic fibrosis in several families. Hum Mutat. 2021;42(10):1221–8.
Oniya O, Neves K, Ahmed B, Konje JC. A review of the reproductive consequences of consanguinity. Eur J Obstet Gynecol Reprod Biol. 2019;232:87–96.
Otto EA, Tory K, Attanasio M, Zhou W, Chaki M, Paruchuri Y, Wise EL, Wolf MT, Utsch B, Becker C, Nürnberg G, Nürnberg P, Nayir A, Saunier S, Antignac C, Hildebrandt F. Hypomorphic mutations in meckelin (MKS3/TMEM67) cause nephronophthisis with liver fibrosis (NPHP11). J Med Genet. 2009;46(10):663–70.
Pauli S, Altmüller J, Schröder S, Ohlenbusch A, Dreha-Kulaczewski S, Bergmann C, Nürnberg P, Thiele H, Li Y, Wollnik B, Brockmann K. Homozygosity for the c.428delG variant in KIAA0586 in a healthy individual: implications for molecular testing in patients with Joubert syndrome. J Med Genet. 2019;56(4):261–4.
Radha Rama Devi A, Naushad SM, Lingappa L. Clinical and molecular diagnosis of Joubert syndrome and related disorders. Pediatr Neurol. 2020;106:43–9.
Ripperger T, Beger C, Rahner N, Sykora KW, Bockmeyer CL, Lehmann U, Kreipe HH, Schlegelberger B. Constitutional mismatch repair deficiency and childhood leukemia/lymphoma--report on a novel biallelic MSH6 mutation. Haematologica. 2010;95(5):841–4.
Romdhane L, Mezzi N, Hamdi Y, El-Kamah G, Barakat A, Abdelhak S. Consanguinity and inbreeding in health and disease in North African populations. Annu Rev Genomics Hum Genet. 2019;20:155–79.
Rousseau-Nepton I, Okubo M, Grabs R, Mitchell J, Polychronakos C, Rodd C. A founder AGL mutation causing glycogen storage disease type IIIa in Inuit identified through whole-exome sequencing: a case series. CMAJ. 2015;187(2):E68–e73.
Saeed S, Arslan M, Froguel P. Genetics of obesity in consanguineous populations: toward precision medicine and the discovery of novel obesity genes. Obesity (Silver Spring). 2018;26(3):474–84.
Scherag A, Dina C, Hinney A, Vatin V, Scherag S, Vogel CI, Müller TD, Grallert H, Wichmann HE, Balkau B, Heude B, Jarvelin MR, Hartikainen AL, Levy-Marchal C, Weill J, Delplanque J, Körner A, Kiess W, Kovacs P, Rayner NW, Prokopenko I, McCarthy MI, Schäfer H, Jarick I, Boeing H, Fisher E, Reinehr T, Heinrich J, Rzehak P, Berdel D, Borte M, Biebermann H, Krude H, Rosskopf D, Rimmbach C, Rief W, Fromme T, Klingenspor M, Schürmann A, Schulz N, Nöthen MM, Mühleisen TW, Erbel R, Jöckel KH, Moebus S, Boes T, Illig T, Froguel P, Hebebrand J, Meyre D. Two new Loci for body-weight regulation identified in a joint analysis of genome-wide association studies for early-onset extreme obesity in French and german study groups. PLoS Genet. 2010;6(4):e1000916.
Sheikh AB, Nasrullah A, Haq S, Akhtar A, Ghazanfar H, Nasir A, Afzal RM, Bukhari MM, Chaudhary AY, Naqvi SW. The interplay of genetics and environmental factors in the development of obesity. Cureus. 2017;9(7):e1435.
Small N, Bittles AH, Petherick ES, Wright J. Endogamy, consanguinity and the health implications of changing marital choices in the UK Pakistani community. J Biosoc Sci. 2017;49(4):435–46.
Smedley D, Smith KR, Martin A, Thomas EA, McDonagh EM, Cipriani V, Ellingford JM, Arno G, Tucci A, Vandrovcova J, Chan G, Williams HJ, Ratnaike T, Wei W, Stirrups K, Ibanez K, Moutsianas L, Wielscher M, Need A, Barnes MR, Vestito L, Buchanan J, Wordsworth S, Ashford S, Rehmström K, Li E, Fuller G, Twiss P, Spasic-Boskovic O, Halsall S, Floto RA, Poole K, Wagner A, Mehta SG, Gurnell M, Burrows N, James R, Penkett C, Dewhurst E, Gräf S, Mapeta R, Kasanicki M, Haworth A, Savage H, Babcock M, Reese MG, Bale M, Baple E, Boustred C, Brittain H, de Burca A, Bleda M, Devereau A, Halai D, Haraldsdottir E, Hyder Z, Kasperaviciute D, Patch C, Polychronopoulos D, Matchan A, Sultana R, Ryten M, Tavares ALT, Tregidgo C, Turnbull C, Welland M, Wood S, Snow C, Williams E, Leigh S, Foulger RE, Daugherty LC, Niblock O, Leong IUS, Wright CF, Davies J, Crichton C, Welch J, Woods K, Abulhoul L, Aurora P, Bockenhauer D, Broomfield A, Cleary MA, Lam T, Dattani M, Footitt E, Ganesan V, Grunewald S, Compeyrot-Lacassagne S, Muntoni F, Pilkington C, Quinlivan R, Thapar N, Wallis C, Wedderburn LR, Worth A, Bueser T, Compton C, Deshpande C, Fassihi H, Haque E, Izatt L, Josifova D, Mohammed S, Robert L, Rose S, Ruddy D, Sarkany R, Say G, Shaw AC, Wolejko A, Habib B, Burns G, Hunter S, Grocock RJ, Humphray SJ, Robinson PN, Haendel M, Simpson MA, Banka S, Clayton-Smith J, Douzgou S, Hall G, Thomas HB, O'Keefe RT, Michaelides M, Moore AT, Malka S, Pontikos N, Browning AC, Straub V, Gorman GS, Horvath R, Quinton R, Schaefer AM, Yu-Wai-Man P, Turnbull DM, McFarland R, Taylor RW, O'Connor E, Yip J, Newland K, Morris HR, Polke J, Wood NW, Campbell C, Camps C, Gibson K, Koelling N, Lester T, Németh AH, Palles C, Patel S, Roy NBA, Sen A, Taylor J, Cacheiro P, Jacobsen JO, Seaby EG, Davison V, Chitty L, Douglas A, Naresh K, McMullan D, Ellard S, Temple IK, Mumford AD, Wilson G, Beales P, Bitner-Glindzicz M, Black G, Bradley JR, Brennan P, Burn J, Chinnery PF, Elliott P, Flinter F, Houlden H, Irving M, Newman W, Rahman S, Sayer JA, Taylor JC, Webster AR, Wilkie AOM, Ouwehand WH, Raymond FL, Chisholm J, Hill S, Bentley D, Scott RH, Fowler T, Rendon A, Caulfield M. 100,000 genomes pilot on rare-disease diagnosis in health care - preliminary report. N Engl J Med. 2021;385(20):1868–80.
Speakman JR, Levitsky DA, Allison DB, Bray MS, de Castro JM, Clegg DJ, Clapham JC, Dulloo AG, Gruer L, Haw S, Hebebrand J, Hetherington MM, Higgs S, Jebb SA, Loos RJ, Luckman S, Luke A, Mohammed-Ali V, O'Rahilly S, Pereira M, Perusse L, Robinson TN, Rolls B, Symonds ME, Westerterp-Plantenga MS. Set points, settling points and some alternative models: theoretical options to understand how genes and environments combine to regulate body adiposity. Dis Model Mech. 2011;4(6):733–45.
Sukenik-Halevy R, Leil-Zoabi UA, Peled-Perez L, Zlotogora J, Allon-Shalev S. Compliance for genetic screening in the Arab population in Israel. Isr Med Assoc J. 2012;14(9):538–42.
Tadmouri GO, Nair P, Obeid T, Al Ali MT, Al Khaja N, Hamamy HA. Consanguinity and reproductive health among Arabs. Reprod Health. 2009;6:17.
Warsy AS, Al-Jaser MH, Albdass A, Al-Daihan S, Alanazi M. Is consanguinity prevalence decreasing in Saudis?: a study in two generations. Afr Health Sci. 2014;14(2):314–21.
Wheeler E, Huang N, Bochukova EG, Keogh JM, Lindsay S, Garg S, Henning E, Blackburn H, Loos RJ, Wareham NJ, O'Rahilly S, Hurles ME, Barroso I, Farooqi IS. Genome-wide SNP and CNV analysis identifies common and low-frequency variants associated with severe early-onset obesity. Nat Genet. 2013;45(5):513–7.
Ye Z, Jia X, Liu X, Zhang Q, Wang K, Chen M. Case report: the first reported concurrence of wilson disease and bilateral retinitis pigmentosa. Front Med (Lausanne). 2022;9:877752.
Zayoud K, Kraoua I, Chikhaoui A, Calmels N, Bouchoucha S, Obringer C, Crochemore C, Najjar D, Zarrouk S, Miladi N, Laugel V, Ricchetti M, Turki I, Yacoub-Youssef H. Identification and characterization of a novel recurrent ERCC6 variant in patients with a severe form of Cockayne syndrome B. Genes (Basel). 2021;12(12):1922.
Zeevi DA, Chung WK, Levi C, Scher SY, Bringer R, Kahan Y, Muallem H, Benel R, Hirsch Y, Weiden T, Ekstein A, Ekstein J. Recommendation of premarital genetic screening in the Syrian Jewish community based on mutation carrier frequencies within Syrian Jewish cohorts. Mol Genet Genomic Med. 2021;9(8):e1756.
Acknowledgements
JAS is supported by Kidney Research UK and the Northern Counties Kidney Research Fund.
Funding
JAS is funded by Kidney Research UK (Paed_RP_001_20180925) and the Northern Counties Kidney Research Fund (01/19).
Author information
Authors and Affiliations
Contributions
GT conceived the idea and wrote the first draft. NN and JAS edited the manuscript. All authors approved of the final version.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Ethical approval and consent to participate was not required for this study as it did not involve patients directly.
Consent for publication
Not applicable.
Competing interests
Professor John Sayer is a co-author of this study and an Editorial Board member of the journal. He was not involved in handling this manuscript during the review process. The rest of the authors have no conflict of interest to declare.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Temaj, G., Nuhii, N. & Sayer, J.A. The impact of consanguinity on human health and disease with an emphasis on rare diseases. J Rare Dis 1, 2 (2022). https://doi.org/10.1007/s44162-022-00004-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44162-022-00004-5