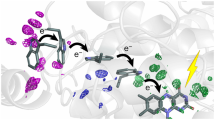
Abstract
Quenching of flavin fluorescence by electron transfer from neighboring aromatic residues is ubiquitous in flavoproteins. Apart from constituting a functional process in specific light-active systems, time-resolved spectral characterization of the process can more generally be employed as a probe for the active site configuration and dynamics. In the C51A variant of the bacterial RNA-transforming flavoenzyme TrmFO from the bacterium Thermus thermophilus, fluorescence is very short-lived (~ 1 ps), and close-by Tyr343 is known to act as the main quencher, as confirmed here by the very similar dynamics observed in protein variants with modified other potential quenchers, Trp283 and Trp214. When Tyr343 is modified to redox-inactive phenylalanine, slower and highly multiphasic kinetics are observed on the picosecond-nanosecond timescale, reflecting heterogeneous electron donor–acceptor configurations. We demonstrate that Trp214, which is located on a potentially functional flexible loop, contributes to electron donor quenching in this variant. Contrasting with observations in other nucleic acid-transforming enzymes, these kinetics are strikingly temperature-independent. This indicates (a) near-barrierless electron transfer reactions and (b) no exchange between different configurations on the timescale up to at least 2 ns, despite the presumed flexibility of Trp214. Results of extensive molecular dynamics simulations are presented to explain this unexpected finding in terms of slowly exchanging protein configurations.





Similar content being viewed by others
Data availability
Not applicable.
Code availability
Not applicable.
References
Miura, R. (2001). Versatility and specificity in flavoenzymes: control mechanisms of flavin reactivity. Chemical Record, 1(3), 183–194. https://doi.org/10.1002/tcr.1007
van den Berg, P. A. W., Feenstra, K. A., Mark, A. E., Berendsen, H. J. C., & Visser, A. J. W. G. (2002). Dynamic conformations of flavin adenine dinucleotide: simulated molecular dynamics of the flavin cofactor related to the time-resolved fluorescence characteristics. The Journal of Physical Chemistry B, 106(34), 8858–8869. https://doi.org/10.1021/jp020356s
Brazard, J., Usman, A., Lacombat, F., Ley, C., Martin, M. M., & Plaza, P. (2011). New insights into the ultrafast photophysics of oxidized and reduced FAD in solution. Journal of Physical Chemistry A, 115(15), 3251–3262. https://doi.org/10.1021/jp110741y
Aubert, C., Mathis, P., Eker, A. P. M., & Brettel, K. (1999). Intraprotein electron transfer between tyrosine and tryptophan in DNA photolyase from Anacystis nidulans. Proceedings of the National academy of Sciences of the United States of America, 96(10), 5423–5427
Giovani, B., Byrdin, M., Ahmad, M., & Brettel, K. (2003). Light-induced electron transfer in a cryptochrome blue-light photoreceptor. Natural Structural Biology, 10(6), 489–490
Lacombat, F., Espagne, A., Dozova, N., Plaza, P., Muller, P., Brettel, K., et al. (2019). Ultrafast oxidation of a tyrosine by proton-coupled electron transfer promotes light activation of an animal-like cryptochrome. Journal of the American Chemical Society, 141(34), 13394–13409. https://doi.org/10.1021/jacs.9b03680
Lukacs, A., Brust, R., Haigney, A., Laptenok, S. P., Addison, K., Gil, A., et al. (2014). BLUF domain function does not require a metastable radical intermediate state. Journal of the American Chemical Society, 136(12), 4605–4615. https://doi.org/10.1021/ja4121082
Mathes, T., van Stokkum, I. H., Kennis, J. T. (2014). Photoactivation mechanisms of flavin-binding photoreceptors revealed through ultrafast spectroscopy and global analysis methods. In Weber, S., Schleicher, E. (Eds.), Flavins and Flavoproteins: Methods and Protocols. Springer
Sorigué, D., Légeret, B., Cuiné, S., Blangy, S., Moulin, S., Billon, E., et al. (2017). An algal photoenzyme converts fatty acids to hydrocarbons. Science, 357(6354), 903–907. https://doi.org/10.1126/science.aan6349
Sorigué, D., Hadjidemetriou, K., Blangy, S., Gotthard, G., Bonvalet, A., Coquelle, N., et al. (2021). Mechanism and dynamics of fatty acid photodecarboxylase. Science, 372, eabd5687
Bialas, C., Barnard, D. T., Auman, D. B., Mcbride, R. A., Jarocha, L. E., Hore, P. J., et al. (2019). Ultrafast flavin/tryptophan radical pair kinetics in a magnetically sensitive artificial protein. Physical Chemistry, 21(25), 13453–13461
Nag, L., Lukacs, A., & Vos, M. H. (2019). Short-lived radical intermediates in the photochemistry of glucose oxidase. ChemPhysChem, 20(14), 1793–1798. https://doi.org/10.1002/cphc.201900329
Ernst, S., Rovida, S., Mattevi, A., Fetzner, S., & Drees, S. L. (2020). Photoinduced monooxygenation involving NAD(P)H-FAD sequential single-electron transfer. Nature Comm, 11(1), 2600. https://doi.org/10.1038/s41467-020-16450-y
Dozova, N., Lacombat, F., Bou-Nader, C., Hamdane, D., & Plaza, P. (2019). Ultrafast photoinduced flavin dynamics in the unusual active site of the tRNA methyltransferase TrmFO. Physical Chemistry Chemical Physics: PCCP, 21(17), 8743–8756. https://doi.org/10.1039/c8cp06072j
Nag, L., Sournia, P., Myllykallio, H., Liebl, U., & Vos, M. H. (2017). Identification of the TyrOH●+ radical cation in the flavoenzyme TrmFO. Journal of the American Chemical Society, 139(33), 11500–11505. https://doi.org/10.1021/jacs.7b04586
Mataga, N., Chosrowjan, H., Shibata, Y., Tanaka, F., Nishina, Y., & Shiga, K. (2000). Dynamics and mechanisms of ultrafast fluorescence quenching reactions of flavin chromophores in protein nanospace. The Journal of Physical Chemistry B, 104(45), 10667–10677
Zhong, D., & Zewail, A. H. (2001). Femtosecond dynamics of flavoproteins: charge separation and recombination in riboflavin (vitamin B2)-binding protein and in glucose oxidase enzyme. Proceedings of the National academy of Sciences of the United States of America, 98, 11867–11872
Laptenok, S. P., Bouzhir-Sima, L., Lambry, J.-C., Myllykallio, H., Liebl, U., & Vos, M. H. (2013). Ultrafast real time visualization of the active site flexibility of the flavoenzyme thymidylate synthase ThyX. Proceedings of the National academy of Sciences of the United States of America, 110, 8924–8929
Yang, H., Luo, G., Karnchanaphanurach, P., Louie, T.-M., Rech, I., Cova, S., et al. (2003). Protein conformational dynamics probed by single-molecule electron transfer. Science, 302(5643), 262–266. https://doi.org/10.1126/science.1086911
Moser, C. C., Anderson, J. L. R., & Dutton, P. L. (2010). Guidelines for tunneling in enzymes. Biochimica et Biophysica Acta, 1797(9), 1573–1586
Zhuang, B., Seo, D., Aleksandrov, A., & Vos, M. H. (2021). Characterization of light-induced short-lived interacting radicals in the active site of flavoprotein ferredoxin-NADP+ oxidoreductase. Journal of the American Chemical Society, 143, 2457–2768
Pirisi, K., Nag, L., Fekete, Z., Iuliano, J. N., Tollentino Collado, J., Clark, I. P., et al. (2021). Identification of the vibrational marker of tyrosine cation radical using ultrafast transient infrared spectroscopy of flavoprotein systems. Photochemical and Photobiological Sciences, 20, 369–378
Hamdane, D., Argentini, M., Cornu, D., Myllykallio, H., Skouloubris, S., Hui-Bon-Hoa, G., et al. (2011). Insights into Folate/FAD-dependent tRNA methyltransferase mechanism: role of two highly conserved cysteines in catalysis. Journal of Biological Chemistry, 286(42), 36268–36280. https://doi.org/10.1074/jbc.M111.256966
Urbonavicius, J., Skouloubris, S., Myllykallio, H., & Grosjean, H. (2005). Identification of a novel gene encoding a flavin-dependent tRNA:m5U methyltransferase in bacteria -evolutionary implications. Nucleic Acids Research, 33(13), 3955–3964
Nishimasu, H., Ishitani, R., Yamashita, K., Iwashita, C., Hirata, A., Hori, H., et al. (2009). Atomic structure of a folate/FAD-dependent tRNA T54 methyltransferase. Proceedings of the National academy of Sciences of the United States of America, 106(20), 8180–8185. https://doi.org/10.1073/pnas.0901330106
Sournia, P. (2016). La méthylation flavine-dépendante d’acides nucléiques : aspects évolutifs, métaboliques, biochimiques et spectroscopiques. Ecole Polytechnique.
Hamdane, D., Bruch, E., Un, S., Field, M., & Fontecave, M. (2013). Activation of a unique flavin-dependent tRNA-methylating agent. Biochemistry, 52(49), 8949–8956. https://doi.org/10.1021/bi4013879
Renger, G. (2013). Apparatus and mechanism of photosynthetic water splitting as nature’s blueprint for efficient solar energy exploitation. In R. Razeghifard (Ed.), Natural and artificial photosynthesis.John Wiley and Son.
Gauden, M., van Stokkum, I. H. M., Key, J. M., Lührs, D. C., van Grondelle, R., Hegemann, P., et al. (2006). Hydrogen-bond switching through a radical pair mechanism in a flavin-binding photoreceptor. Proceedings of the National academy of Sciences of the United States of America, 103(29), 10895–10900. https://doi.org/10.1073/pnas.0600720103
Laptenok, S. P., Nuernberger, P., Lukacs, A., & Vos, M. H. (2014). Subpicosecond Kerr-gate spectrofluorometry. In Y. Engelborghs & A. J. W. G. Visser (Eds.), Methods in molecular biology, fluorescence spectroscopy and microscopy: methods and protocols.Humana Press.
Lambry, J.-C., Stranava, M., Lobato, L., Martinkova, M., Shimizu, T., Liebl, U., et al. (2016). Ultrafast spectroscopy evidence for picosecond ligand exchange at the binding site of a heme protein: heme-based sensor YddV. Journal of Physical Chemistry Letters, 7(1), 69–74. https://doi.org/10.1021/acs.jpclett.5b02517
Snellenburg, J. J., Laptenok, S. P., Seger, R., Mullen, K. M., & van Stokkum, I. H. M. (2012). Glotaran: a java-based graphical user interface for the R package TIMP. J Stat Software, 49(3), 2
Webb, B., & Sali, A. (2016). Comparative protein structure modeling using modeller. Curr Protoc Bioinform, 54(1), 561–5637. https://doi.org/10.1002/cpbi.3
Laskowski, R. A., MacArthur, M. W., Moss, D. S., & Thornton, J. M. (1993). PROCHECK: a program to check the stereochemical quality of protein structures. Journal of Applied Crystallography, 26(2), 283–291. https://doi.org/10.1107/S0021889892009944
Brooks, B. R., Brooks, C. L., Mackerell, A. D., Nilsson, L., Petrella, R. J., Roux, B., et al. (2009). CHARMM: the biomolecular simulation program. J Comp Chem, 30(10), 1545–1614
Darden, T., York, D., & Pedersen, L. (1993). Particle mesh Ewald: An N⋅log(N) method for Ewald sums in large systems. The Journal of Chemical Physics, 98(12), 10089–10092. https://doi.org/10.1063/1.464397
Phillips, J. C., Braun, R., Wang, W., Gumbart, J., Tajkhorshid, E., Villa, E., et al. (2005). Scalable molecular dynamics with NAMD. J Comp Chem, 26(16), 1781–1802. https://doi.org/10.1002/jcc.20289
Huang, J., & MacKerell, A. D., Jr. (2013). CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data. J Comp Chem, 34(25), 2135–2145. https://doi.org/10.1002/jcc.23354
Jorgensen, W. L., Chandrasekhar, J., Madura, J. D., Impey, R. W., & Klein, M. L. (1983). Comparison of simple potential functions for simulating liquid water. The Journal of Chemical Physics, 79(2), 926–935. https://doi.org/10.1063/1.445869
Neria, E., Fischer, S., & Karplus, M. (1996). Simulation of activation free energies in molecular systems. The Journal of Chemical Physics, 105(5), 1902–1921. https://doi.org/10.1063/1.472061
Aleksandrov, A. (2019). A molecular mechanics model for Flavins. J Comp Chem, 40(32), 2834–2842. https://doi.org/10.1002/jcc.26061
Sugita, Y., & Okamoto, Y. (1999). Replica-exchange molecular dynamics method for protein folding. Chemical Physics Letters, 314(1), 141–151. https://doi.org/10.1016/S0009-2614(99)01123-9
Patriksson, A., & van Spoel, D. (2008). A temperature predictor for parallel tempering simulations. Physical Chemistry Chemical Physics: PCCP, 10(15), 2073–2077. https://doi.org/10.1039/B716554D
Acknowledgements
B. Z. is supported by a China Scholarship Council PhD scholarship. Part of this work was performed using High Performance Computing resources from Grand Equipement National pour le Calcul Intensif [Centre de Calcul Recherche et Technologie/Centre Informatique National de l’Enseignement Supérieur/Institut du Développement et des Ressources en Informatique Scientifique] (Grant 2018-2019, project number A0020706913).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Not applicable.
Additional information
Pushing the limits of flash photolysis to unravel the secrets of biological electron and proton transfer - a topical issue in honour of Klaus Brettel.
Rights and permissions
About this article
Cite this article
Zhuang, B., Nag, L., Sournia, P. et al. Photochemical processes in flavo-enzymes as a probe for active site dynamics: TrmFO of Thermus thermophilus. Photochem Photobiol Sci 20, 663–670 (2021). https://doi.org/10.1007/s43630-021-00052-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43630-021-00052-8