Abstract
Background
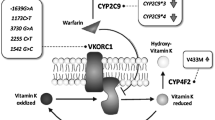
Several studies optimized the warfarin dose based on CYP2C9*2, CYP2C9*3, VKORC1 -1639 G > A, CYP4F2 V433M. But, the information on the rare variants is lacking. In this study, we have explored the prevalence of common and rare pharmacogenetic determinants of warfarin and determined their damaging nature.
Methods
We have analyzed 2000 healthy adults using the Infinium global screening array (GSA) for 15 pharmacogenetic determinants of warfarin. In addition, we have elucidated the impact of these variants on protein function, stability, dynamics, evolutionary preservation, and ligand binding propensity.
Results
The GSA Analysis has revealed that CYP4F2 V433M (MAF: 39.425%), VKORC1 -1639 G > A (MAF: 20.5%), CYP2C9*3 (MAF:9.925%), and CYP2C9*2 (MAF:4.575%) are common, while CYP2C9*14 (MAF: 1.475%), CYP2C9*4 (0.175%), CYP2C9*5 (0.125%), and CYP2C9*11 (0.125%) are rare. Position-specific evolutionary preservation (PSEP) analysis has revealed that CYP2C9*4 is possibly damaging, while CYP2C9*5, CYP2C9*11, and CYP2C9*14 are probably damaging. CYP2C9*4 has high thermolability (−10.14 kcal/mol). Among the rare CYP2C9 variants, CYP2C9*4 and CYP2C9*11 exert destabilizing effects and may have increased molecular flexibility, while CYP2C9*5 and CYP2C9*14 exert stabilizing effects and may have decreased molecular flexibility. DNase I footprint analysis has revealed the loss of the E-box consensus sequence due to VKORC1 -1639 G > A polymorphism.
Conclusion
CYP2C9*2, CYP2C9*3, VKORC1 -1639 G > A and CYP4F2 V433M are common; CYP2C9*4, CYP2C9*5, CYP2C9*11, and CYP2C9*14 variants are rare in Indian subjects. All the CYP2C9 variants are found to be damaging. DNase I footprint analysis provided the mechanistic rationale for the association of VKORC1 -1639 G > A with warfarin sensitivity.



Similar content being viewed by others
Abbreviations
- AFR:
-
Africans
- AMR:
-
Americans
- CUPSAT:
-
Cologne University Protein Stability Analysis Tool
- CYP2C9:
-
Cytochrome P450 2 C9
- CYP4F2:
-
Cytochrome P450 4 F2
- DDIG-in:
-
Detecting disease-causing genetic variants due to indels
- EAS:
-
East Asians
- EUR:
-
Europeans
- GGCX:
-
Gamma-glutamyl carboxylase
- PSEP:
-
Position-specific evolutionary preservation
- MAF:
-
Minor allele frequency
- SAS:
-
South Asians
- SFR:
-
Shoulder-to-foot ratio
- VKORC1:
-
Vitamin K epoxide reductase complex 1
References
Saksena D, Muralidharan S, Mishra YK, Kanhere V, Mohanty BB, Srivastava CP, et al. Anticoagulation management in patients with valve replacement. J Assoc Physicians Ind. 2018;66(1):59–74.
Sawhney JP, Kothiwale VA, Bisne V, Durgaprasad R, Jadhav P, Chopda M, et al. Risk profiles and 1-year outcomes of patients with newly diagnosed atrial fibrillation in India: insights from the GARFIELD-AF registry. Ind Heart J. 2018;70(6):828–35.
Krishna Kumar D, Shewade DG, Loriot MA, Beaune P, Balachander J, Sai Chandran BV, et al. Effect of CYP2C9, VKORC1, CYP4F2 and GGCX genetic variants on warfarin maintenance dose and explicating a new pharmacogenetic algorithm in South Indian population. Eur J Clin Pharmacol. 2014;70(1):47–56.
Takeuchi F, McGinnis R, Bourgeois S, Barnes C, Eriksson N, Soranzo N, et al. A genome-wide association study confirms VKORC1, CYP2C9, and CYP4F2 as principal genetic determinants of warfarin dose. PLoS Genet. 2009;5(3):1000433.
International Warfarin Pharmacogenetics Consortium, Klein TE, Altman RB, Eriksson N, Gage BF, Kimmel SE, et al. Estimation of the warfarin dose with clinical and pharmacogenetic data. N Engl J Med. 2009;360(8):753–64.
Gage BF, Eby C, Johnson JA, Deych E, Rieder MJ, Ridker PM, et al. Use of pharmacogenetic and clinical factors to predict the therapeutic dose of warfarin. Clin Pharmacol Ther. 2008;84(3):326–31.
Wadelius M, Chen LY, Eriksson N, Bumpstead S, Ghori J, Wadelius C, et al. Association of warfarin dose with genes involved in its action and metabolism. Hum Genet. 2007;121(1):23–34. https://doi.org/10.1007/s00439-006-0260-8 (Epub 2006 Oct 18).
Pavani A, Naushad SM, Rupasree Y, Kumar TR, Malempati AR, Pinjala RK, et al. Optimization of warfarin dose by population-specific pharmacogenomic algorithm. Pharmacogenomics J. 2012;12(4):306–11.
Pavani A, Naushad SM, Mishra RC, Malempati AR, Pinjala R, Kumar TR, Kutala VK. Retrospective evidence for clinical validity of expanded genetic model in warfarin dose optimization in a South Indian population. Pharmacogenomics. 2012;13(8):869–78.
Nizamuddin S, Dubey S, Singh S, Sharma S, Machha P, Thangaraj K. CYP2C9 variations and their pharmacogenetic implications among diverse South Asian populations. Pharmgen Pers Med. 2021;27(14):135–47.
Chaudhary N, Kabra M, Gulati S, Gupta YK, Pandey RM, Bhatia BD. Frequencies of CYP2C9 polymorphisms in North Indian population and their association with drug levels in children on phenytoin monotherapy. BMC Pediatr. 2016;14(16):66.
Zhao F, Loke C, Rankin SC, Guo JY, Lee HS, Wu TS, et al. Novel CYP2C9 genetic variants in Asian subjects and their influence on maintenance warfarin dose. Clin Pharmacol Ther. 2004;76(3):210–9.
Zhang L, Sarangi V, Moon I, Yu J, Liu D, Devarajan S, Reid JM, Kalari KR, Wang L, Weinshilboum R. CYP2C9 and CYP2C19: deep mutational scanning and functional characterization of genomic missense variants. Clin Transl Sci. 2020;13(4):727–42.
Naushad SM, Janaki Ramaiah M, Kutala VK, Hussain T, Alrokayan SA. Pharmacogenetic determinants of thiopurines in an Indian cohort. Pharmacol Rep. 2021;73(1):278–87. https://doi.org/10.1007/s43440-020-00158-3.
Naushad SM, Hussain T, Alrokayan SA, Kutala VK. Pharmacogenetic profiling of dihydropyrimidine dehydrogenase (DPYD) variants in the Indian population. J Gene Med. 2021;23(1):e3289. https://doi.org/10.1002/jgm.3289.
Dickmann LJ, Rettie AE, Kneller MB, Kim RB, Wood AJ, Stein CM, Wilkinson GR, Schwarz UI. Identification and functional characterization of a new CYP2C9 variant (CYP2C9*5) expressed among African Americans. Mol Pharmacol. 2001;60(2):382–7.
Niinuma Y, Saito T, Takahashi M, Tsukada C, Ito M, Hirasawa N, Hiratsuka M. Functional characterization of 32 CYP2C9 allelic variants. Pharmacogenomics J. 2014;14(2):107–14.
Wang D, Chen H, Momary KM, Cavallari LH, Johnson JA, Sadée W. Regulatory polymorphism in vitamin K epoxide reductase complex subunit 1 (VKORC1) affects gene expression and warfarin dose requirement. Blood. 2008;112(4):1013–21.
Yuan HY, Chen JJ, Lee MT, Wung JC, Chen YF, Charng MJ, Lu MJ, Hung CR, Wei CY, Chen CH, Wu JY, Chen YT. A novel functional VKORC1 promoter polymorphism is associated with inter-individual and inter-ethnic differences in warfarin sensitivity. Hum Mol Genet. 2005;14(13):1745–51.
McDonald MG, Rieder MJ, Nakano M, Hsia CK, Rettie AE. CYP4F2 is a vitamin K1 oxidase: an explanation for altered warfarin dose in carriers of the V433M variant. Mol Pharmacol. 2009;75(6):1337–46.
Danese E, Raimondi S, Montagnana M, Tagetti A, Langaee T, Borgiani P, et al. Effect of CYP4F2, VKORC1, and CYP2C9 in influencing coumarin dose: a single-patient data meta-analysis in more than 15,000 individuals. Clin Pharmacol Ther. 2019;105(6):1477–91.
Oner Ozgon G, Langaee TY, Feng H, Buyru N, Ulutin T, Hatemi AC, Siva A, Saip S, Johnson JA. VKORC1 and CYP2C9 polymorphisms are associated with warfarin dose requirements in Turkish patients. Eur J Clin Pharmacol. 2008;64(9):889–94.
Cho HJ, Sohn KH, Park HM, Lee KH, Choi B, Kim S, et al. Factors affecting the interindividual variability of warfarin dose requirement in adult Korean patients. Pharmacogenomics. 2007;8(4):329–37.
Quinn AL, Liko I, Lee JC. Clinical effect of CYP2C9*5/*6 genotype on a patient’s warfarin dose requirement. Pharmacogenomics. 2017;18(11):1051–7.
Tai G, Farin F, Rieder MJ, Dreisbach AW, Veenstra DL, Verlinde CL, et al. In-vitro and in-vivo effects of the CYP2C9*11 polymorphism on warfarin metabolism and dose. Pharmacogenet Genomics. 2005;15(7):475–81.
Quinn AL, Bhat S, Lee JC. Effect of CYP2C9 *11/*11 genotype on initial and long-term warfarin dose requirement and therapeutic response. Pharmacogenomics. 2020;21(18):1271–7.
Lee YM, Eggen J, Soni V, Drozda K, Nutescu EA, Cavallari LH. Warfarin dose requirements in a patient with the CYP2C9*14 allele. Pharmacogenomics. 2014;15(7):909–14.
Johnson JA, Caudle KE, Gong L, Whirl-Carrillo M, Stein CM, Scott SA, Lee MT, Gage BF, Kimmel SE, Perera MA, Anderson JL, Pirmohamed M, Klein TE, Limdi NA, Cavallari LH, Wadelius M. Clinical pharmacogenetics implementation consortium (CPIC) guideline for pharmacogenetics-guided warfarin dosing: 2017 update. Clin Pharmacol Ther. 2017;102(3):397–404.
Funding
The authors are grateful to the Deanship of Scientific Research, King Saud University for funding through Vice Deanship of Scientific Research Chairs.
Author information
Authors and Affiliations
Contributions
SMN and VKK conceptualized and designed the study. SMN, VKK, TH, SAA acquired, analyzed, and interpreted the data. SMN and VKK drafted the manuscript. TH and SAA revised it critically. The final version of the manuscript was approved by all the authors. The accuracy and integrity of the work are ascertained by all the authors.
Corresponding author
Ethics declarations
Conflicts of interest
All the authors hereby declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Naushad, S.M., Kutala, V.K., Hussain, T. et al. Pharmacogenetic determinants of warfarin in the Indian population. Pharmacol. Rep 73, 1396–1404 (2021). https://doi.org/10.1007/s43440-021-00297-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43440-021-00297-1