Abstract
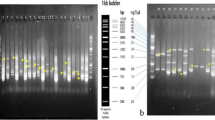
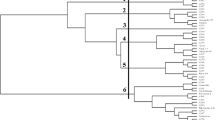
Frequent changes in ecosystems and environments through the change of climate have modified the rainfall patterns and seasons. This unpredictability has placed new emphasis on breeding resilient wheat varieties alongside higher yield and better nutritional quality. Use of few successful varieties as parents for breeding new varieties led to the loss of locally adaptive genetic diversity. Therefore, selection and use of diverse genotypes in a breeding program is required to create genetic variability. This study was initiated to study genomic diversity of 50 accessions of wheat at NIAB with an aim to breed new and resilient cultivars of wheat using sequence-related amplified polymorphism (SRAP). SRAP markers divided the germplasm into three main clusters comprising of 21 (Cluster 1), 19 (Cluster 2) and 10 genotypes (Cluster 3) with futher sub-gorups within them. More importantly, out of 10 cultivars developed in Pakistan, 6 fall in same cluster 1 (e.g., TAKBEER, LALMA, KT-335, INQALAB-91, NIA-SUNHARI and GALAXY-13), 1 line (NIA-AMBER in sub-group 2d) falls in cluster 2 and the rest of three lines (FAISALABAD-08 (sub-group 3a), PUNJAB-11 and BENAZIR-12 (both in sub-group 3b) fall in cluster 3. This study also identified a NIAB advanced line NW-1-20 distinctive from the other advanced lines indicating diverse parents in its pedigree. Thus, we found that SRAP markers were helpful in revealing the hidden population structure in a germplasm collection. To add to it, this study will help in selecting diverse parents for future wheat breeding at NIAB.


Similar content being viewed by others
Notes
Further details about the Electronic Supplementary Material (ESM) can be found at the end of the article.
References
Abdelkhaliq SM, Salem AKM, Abdelaziz AR, Ammar MH (2016) Morphological and sequence-related amplified polymorphism-based molecular diversity of local and exotic wheat genotypes. Genet Mol Res 15:1–9
Alghamdi SS, Al-Faifi SA, Migdadi HM, Khan MA, El-Harty EH, Ammar MH (2012) Molecular diversity assessment using sequence related amplified polymorphism (SRAP) markers in Vicia faba L. Int J Mol Sci 13:16457–16471
Ayala FJ, Kiger J (1984) Modern Genetics. Benjamin, Cummings, Menlo Park, pp 123–134
Baranwal D, Mishra V, Singh T (2013) Genetic diversity based on cluster and principal component analyses for yield and its contributing characters in wheat (Triticum aestivum L.). Madras Agric J 100:320–323
Bar-Hen A, Charcosset A, Bourgoin M, Guiard J (1995) Relationship between genetic markers and morphological traits in a maize inbred lines collection. Euphytica 84:145–154
Bernardo R (2013) Estimation of coefficient of coancestry using molecular markers in maize. Theor Appl Genet 85:1055–1062
Borrill P, Fahy B, Smith AM, Uauy C (2015) Wheat grain filling is limited by grain filling capacity rather than the duration of flag leaf photosynthesis: a case study using NAM RNAi plants. PLoS ONE 10:e0134947
Bux H, Ashraf M, Chen XM, Mumtaz AS (2011) Effective genes for resistance to stripe rust and virulence of Puccinia striiformis f. sp. tritici in Pakistan. Afr J Biotechnol 10:5489–5495
da Silva EF, de Sousa SB, da Silva GF, Sousa NR, do Nascimento Filho FJ, Hanada RE (2016) TRAP and SRAP markers to find genetic variability in complex polyploid Paullinia cupana var. sorbilis. Plant Gene 6:43–47
Doyle JJ (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Enghiad A, Danielle U, Countryman AM, Thilmany DD (2017) An overview of global wheat market fundamentals in an era of climate concerns. Int J Agron 2017:1–15
FAO (2017) The State of Food and Agriculture 2017. ISSN 0081-4539
Fouilloux G, Bannerot H (1988) Selection methods in the common bean (Phaseolous vulgaris). In: Gepts P (ed) Genetic resources of Phaseolous beans. Kluwer, Dordrecht, pp 503–542
Gulnaz S, Zulkiffal M, Sajjad M, Ahmed J, Musa M, Abdullah M, Ahsan A, Rehman A (2019) Identifying Pakistani wheat landraces as genetic resources for yield potential, heat tolerance and rust resistance. Int J Agric Biol 21:520–526
Hamrick J, Godt M (1997) Allozyme diversity in cultivated crops. Crop Sci 37:26–30
Karp A, Edwards KJ, Bruford M, Funk S, Vosman B, Morgante M, Seberg O, Kremer A, Boursot P, Arctander P (1997) Molecular technologies for biodiversity evaluation: opportunities and challenges. Nat Biotechnol 15:625–628
Li G, Quiros CF (2001) Sequence-related amplified polymorphism (SRAP), a new marker system based on a simple PCR reaction: its application to mapping and gene tagging in Brassica. Theor Appl Genet 103:455–461
Li QY, Dong SJ, Zhang WY, Lin RQC, Wang R, Qian DX, Lun ZR, Song HQ, Zhu XQ (2009) Sequence-related amplified polymorphism, an effective molecular approach for studying genetic variation in Fasciola spp. of human and animal health significance. Electrophoresis 30:403–409
Lopes SM, El-Basyoni I, Baenziger PS, Singh S, Royo C, Ozbek K (2015) Exploiting genetic diversity from landraces in wheat breeding for adaptation to climate change. J Exp Bot 66:3625–3638
Messmer MM, Melchinger AE, Herrmann RG, Boppenmaier J (1993) Relationships among early European maize inbreds: II. Comparison of pedigree and RFLP data. Crop Sci 33:944–950
Mukhtar S, Rahman M, Zafar Y (2002) Assessment of genetic diversity among wheat cultivars using random amplified polymorphic DNA (RAPD) analysis. Euphytica 128:417–425
Rao VR, Hodgkin T (2002) Genetic diversity and conservation and utilization of plant genetic resources. Plant Cell Tiss Org 68:1–19
Rehman Arif MA, Bux H, Kazi AG, Rasheed A, Napar AA, Riaz A, Mujeeb- Kazi A (2012) Stripe rust analysis of D-genome synthetic wheats (2n = 6x = 42) and their molecular diversity. Arch Phytopathol Pfl 45:1479–1487
Robarts DW, Wolfe AD (2014) Sequence-related amplified polymorphism (SRAP) markers: a potential resource for studies in plant molecular biology. Appl Plant Sci 2(7):1400017
Sajjad M, Xiaoling M, Khan SH, Shoaib M, Song Y, Yang W, Zhang A, Liu D (2017) TaFlo2-A1, an ortholog of rice Flo2, is associated with thousand grain weight in bread wheat (Triticum aestivum L.). BMC Plant Biol 17:164
Sajjad M, Khan SH, Shahzad M (2018) Patterns of allelic diversity in spring wheat populations by SSR markers. Cytol Genet 52:155–160
Smale M, Hartell J, Heisey PW, Senauer B (1998) The contribution of genetic resources and diversity to wheat production in the Punjab of Pakistan. Am J Agr Econ 80:482–493
Smith HW, Huggins MB, Shaw KM (1987) Factors influencing the survival and multiplication of bacteriophages in calves and in their environment. Microbiology 133:1127–1135
Smith O, Smith J (1992) Measurrement of genetic diversity among maize hybrids-a comparison of isozymic, RFLP, pedigree, and heterosis data. Maydica 37:53–60
Van-Hintum TJ, Haalman D (1994) Pedigree analysis for composing a core collection of modern cultivars, with examples from barley (Hordeum vulgare s. lat.). Theor Appl Genet 88:70–74
Yeboah MA, Xuehao C, Feng CR, Liang G, Gu M (2007) A genetic linkage map of cucumber (Cucumis sativus L) combining SRAP and ISSR markers. Afr J Biotechnol 6:2784–2791
Zeb B, Khan I, Ali S, Bacha S, Mumtaz S, Swati Z (2009) Study on genetic diversity in Pakistani wheat varieties using simple sequence repeat (SSR) markers. Afr J Biotechnol 8:4016–4019
Zeshan A, Afzal M, Alghamdi SS, Kettener K, Ali M, Mubushar M, Ahmad S (2016) Evaluation of genetic diversity among the Pakistani Wheat (Triticum aestivum L.) lines through random molecular markers. Braz Arch Biol Technol 59:1
Zhang W, He H, Guan Y, Du H, Yuan L, Li Z, Yao D, Pan J, Cai R (2010) Identification and mapping of molecular markers linked to the tuberculate fruit gene in the cucumber (Cucumis sativus L.). Theor Appl Genet 120:645–654
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Börner.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hassan, R., Waheed, M.Q., Shokat, S. et al. Estimation of genomic diversity using sequence related amplified polymorphism (SRAP) markers in a mini core collection of wheat germplasm from Pakistan. CEREAL RESEARCH COMMUNICATIONS 48, 33–40 (2020). https://doi.org/10.1007/s42976-019-00006-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42976-019-00006-y