Abstract
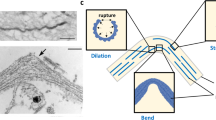
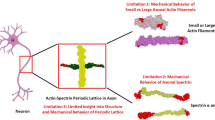
Axonal cytoskeletal components of neuron have been studied and modeled by using different approaches. The axonal cytoskeleton comprises of microtubules (MTs), which mostly control the overall property and behavior of axon. However, MTs contain crosslinks which are MT associated proteins (mainly tau proteins), intermediate filaments (mainly neurofilaments or NFs in case of brain axon), and microfilaments (MFs). Due to the micrometer level length scale of the components, experimental approaches such as microscopy are inconvenient. In this regard, several computational approaches, such as fully atomistic molecular dynamics (MD) simulation, coarse grained (CG) simulation, or finite element analysis (FEA) can be considered reasonable approaches for observing the behavior of the cytoskeletal components and determining their mechanical properties. Among the cytoskeletal components, MTs and MFs are well studied, but the behavior of taus and NFs are not studied comprehensively. Over the last two decades, the computational approaches have been improved manifold to determine and analyze numerous aspects of cytoskeletal components. Due to structurally disordered nature of tau and NF we lack sufficient literature on these components, almost all the structural and behavioral aspects have been analyzed in depth for MT and MF – for which computational studies have played vital role. This study attempts to discuss the current scenario of these computational approaches performed on cytoskeletal components, as well as recent advancements. It attempts to benchmark the current capability of computational approaches to find out properties, behavior, and dynamics of cytoskeletal components, which is required to further advance using similar maneuvers. Importantly, we also pinpoint the possibilities of future studies through which the current limitations in the tau and NF territories can be effectively addressed.





Similar content being viewed by others
Availability of data and material
The data and material are available upon reasonable request from the corresponding author.
Code availability
No software application or custom code was used to create the data and material of this manuscript.
References
R. de Rooij, E. Kuhl, Microtubule Polymerization and Cross-Link Dynamics Explain Axonal Stiffness and Damage. Biophys. J. 114(1), 201–212 (2018). https://doi.org/10.1016/j.bpj.2017.11.010
B.R. Brooks et al., CHARMM: The biomolecular simulation program. J. Comput. Chem. 30(10), 1545–1614 (Jul. 2009). https://doi.org/10.1002/jcc.21287
S.L. Mayo, B.D. Olafson, W.A. Goddard, DREIDING: a generic force field for molecular simulations. J. Phys. Chem. 94(26), 8897–8909 (1990)
J. Wang, R.M. Wolf, J.W. Caldwell, P.A. Kollman, D.A. Case, Development and testing of a general amber force field. J. Comput. Chem. 25(9), 1157–1174 (2004)
W. Damm, A. Frontera, J. Tirado-Rives, W.L. Jorgensen, OPLS all-atom force field for carbohydrates. J. Comput. Chem. 18(16), 1955–1970 (1997)
A.D. Mackerell Jr., M. Feig, C.L. Brooks III, Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J. Comput. Chem. 25(11), 1400–1415 (2004)
T. Stolarski, Y. Nakasone, and S. Yoshimoto, Engineering analysis with ANSYS software. Butterworth-Heinemann, 2018.
J.Z. Wu, W. Herzog, M. Epstein, Evaluation of the finite element software ABAQUS for biomechanical modelling of biphasic tissues. J. Biomech. 31(2), 165–169 (1997)
A. Adnan, S. Qidwai, A. Bagchi, On the atomistic-based continuum viscoelastic constitutive relations for axonal microtubules. J. Mech. Behav. Biomed. Mater. 86, 375–389 (2018)
J. Zhang, C. Wang, Molecular structural mechanics model for the mechanical properties of microtubules. Biomech. Model. Mechanobiol. 13(6), 1175–1184 (2014)
D.B. Wells, A. Aksimentiev, Mechanical properties of a complete microtubule revealed through molecular dynamics simulation. Biophys. J. 99(2), 629–637 (2010)
S. Feng, H. Liang, A coarse grain model of microtubules. Theor. Appl. Mech. Lett. 2(1), 14006 (2012)
S. Barreto, C.H. Clausen, C.M. Perrault, D.A. Fletcher, D. Lacroix, A multi-structural single cell model of force-induced interactions of cytoskeletal components. Biomaterials 34(26), 6119–6126 (2013)
A. Mitra, D. Sept, Taxol Allosterically Alters the Dynamics of the Tubulin Dimer and Increases the Flexibility of Microtubules. Biophys. J. 95(7), 3252–3258 (2008). https://doi.org/10.1529/biophysj.108.133884
Y. Gebremichael, J.-W. Chu, G.A. Voth, Intrinsic Bending and Structural Rearrangement of Tubulin Dimer: Molecular Dynamics Simulations and Coarse-Grained Analysis. Biophys. J. 95(5), 2487–2499 (2008). https://doi.org/10.1529/biophysj.108.129072
S. Enemark, M.A. Deriu, M. Soncini, A. Redaelli, Mechanical model of the tubulin dimer based on molecular dynamics simulations. J. Biomech. Eng. 130(4), 41008 (2008)
M. SONCINI et al., “MICROTUBULE-KINESIN MECHANICS BY MOLECULAR MODELING,” Biophys. Rev. Lett., vol. 04, no. 01n02, pp. 45–61, Apr. 2009, doi: https://doi.org/10.1142/S1793048009000922.
D. Sept, N.A. Baker, J.A. McCammon, The physical basis of microtubule structure and stability. Protein Sci. 12(10), 2257–2261 (Oct. 2003). https://doi.org/10.1110/ps.03187503
A.S. Zeiger, B.E. Layton, Molecular modeling of the axial and circumferential elastic moduli of tubulin. Biophys. J. 95(8), 3606–3618 (2008)
Y.-T. Wu, A. Adnan, Damage and failure of axonal microtubule under extreme high strain rate: an in-silico molecular dynamics study. Sci. Rep. 8(1), 12260 (2018)
R. I. Dima and H. Joshi, “Probing the origin of tubulin rigidity with molecular simulations,” Proc. Natl. Acad. Sci., vol. 105, no. 41, pp. 15743 LP – 15748, Oct. 2008, doi: https://doi.org/10.1073/pnas.0806113105.
A. Grafmüller, G.A. Voth, Intrinsic Bending of Microtubule Protofilaments. Structure 19(3), 409–417 (2011). https://doi.org/10.1016/j.str.2010.12.020
M.A. Deriu, S. Enemark, M. Soncini, F.M. Montevecchi, A. Redaelli, Tubulin: from atomistic structure to supramolecular mechanical properties. J. Mater. Sci. 42(21), 8864–8872 (2007). https://doi.org/10.1007/s10853-007-1784-6
E.J. Carpenter, J.T. Huzil, R.F. Ludueña, J.A. Tuszynski, Homology modeling of tubulin: influence predictions for microtubule’s biophysical properties. Eur. Biophys. J. 36(1), 35–43 (2006). https://doi.org/10.1007/s00249-006-0088-0
J. W. J. Kerssemakers, E. Laura Munteanu, L. Laan, T. L. Noetzel, M. E. Janson, and M. Dogterom, “Assembly dynamics of microtubules at molecular resolution,” Nature, vol. 442, no. 7103, pp. 709–712, 2006, doi: https://doi.org/10.1038/nature04928.
J.A. Tuszyński et al., Molecular dynamics simulations of tubulin structure and calculations of electrostatic properties of microtubules. Math. Comput. Model. 41(10), 1055–1070 (2005). https://doi.org/10.1016/j.mcm.2005.05.002
J.A. Tuszyński, T. Luchko, S. Portet, J.M. Dixon, Anisotropic elastic properties of microtubules. Eur. Phys. J. E 17(1), 29–35 (2005). https://doi.org/10.1140/epje/i2004-10102-5
G. Yoon, J. Kwak, J.I. Kim, S. Na, K. Eom, Mechanical Characterization of Amyloid Fibrils Using Coarse-Grained Normal Mode Analysis. Adv. Funct. Mater. 21(18), 3454–3463 (2011)
P. Zakharov, N. Gudimchuk, V. Voevodin, A. Tikhonravov, F.I. Ataullakhanov, E.L. Grishchuk, Molecular and Mechanical Causes of Microtubule Catastrophe and Aging. Biophys. J. 109(12), 2574–2591 (2015). https://doi.org/10.1016/j.bpj.2015.10.048
P. Xiang, K.M. Liew, Dynamic behaviors of long and curved microtubules based on an atomistic-continuum model. Comput. Methods Appl. Mech. Eng. 223, 123–132 (2012)
A. ADNAN, S. QIDWAI, and A. BAGCHI, “Viscoelastic Response of Microtubule—Tau Proteins Assembly During Axonal Stretch: Combined Atomistic and Continuum Predictions,” in American Society of Composites-30th Technical Conference, 2015.
H. Jiang, L. Jiang, J.D. Posner, B.D. Vogt, Atomistic-based continuum constitutive relation for microtubules: elastic modulus prediction. Comput. Mech. 42(4), 607–618 (2008)
B. Vogt, “Atomistic-based continuum constitutive relation for microtubules: elastic modulus prediction,” 2008.
D. Sept, F.C. MacKintosh, Microtubule elasticity: connecting all-atom simulations with continuum mechanics. Phys. Rev. Lett. 104(1), 18101 (2010)
P. Xiang, K.M. Liew, A computational framework for transverse compression of microtubules based on a higher-order Cauchy-Born rule. Comput. Methods Appl. Mech. Eng. 254, 14–30 (2013). https://doi.org/10.1016/j.cma.2012.10.013
K.M. Liew, P. Xiang, Y. Sun, A continuum mechanics framework and a constitutive model for predicting the orthotropic elastic properties of microtubules. Compos. Struct. 93(7), 1809–1818 (2011). https://doi.org/10.1016/j.compstruct.2011.01.017
H. Ahmadzadeh, D.H. Smith, V.B. Shenoy, Viscoelasticity of Tau Proteins Leads to Strain Rate-Dependent Breaking of Microtubules during Axonal Stretch Injury: Predictions from a Mathematical Model. Biophys. J. 106(5), 1123–1133 (2014). https://doi.org/10.1016/j.bpj.2014.01.024
H. Ahmadzadeh, D.H. Smith, V.B. Shenoy, Mechanical effects of dynamic binding between tau proteins on microtubules during axonal injury. Biophys. J. 109(11), 2328–2337 (2015)
L. An, Y. Gao, Mechanics behavior of microtubules based on nonlocal anisotropic shell theory. IOP Conf. Ser. Mater. Sci. Eng. 10, 12181 (2010). https://doi.org/10.1088/1757-899x/10/1/012181
H. Sim, D. Sept, Properties of Microtubules with Isotropic and Anisotropic Mechanics. Cell. Mol. Bioeng. 6(4), 361–368 (2013). https://doi.org/10.1007/s12195-013-0302-y
M. Hemmat, B.T. Castle, D.J. Odde, Microtubule dynamics: moving toward a multi-scale approach. Curr. Opin. Cell Biol. 50, 8–13 (2018). https://doi.org/10.1016/j.ceb.2017.12.013
M.A. Deriu et al., Anisotropic elastic network modeling of entire microtubules. Biophys. J. 99(7), 2190–2199 (2010)
C. Lazarus, M. Soheilypour, M.R.K. Mofrad, Torsional behavior of axonal microtubule bundles. Biophys. J. 109(2), 231–239 (2015)
X.-Y. Ji, X.-Q. Feng, Coarse-grained mechanochemical model for simulating the dynamic behavior of microtubules. Phys. Rev. E 84(3), 31933 (2011)
S.J. Peter, M.R.K. Mofrad, Computational modeling of axonal microtubule bundles under tension. Biophys. J. 102(4), 749–757 (2012)
S.S. Setayandeh, A. Lohrasebi, Multi scale modeling of 2450 MHz electric field effects on microtubule mechanical properties. J. Mol. Graph. Model. 70, 122–128 (2016)
K. E. Theisen, N. J. Desai, A. M. Volski, and R. I. Dima, “Mechanics of severing for large microtubule complexes revealed by coarse-grained simulations,” J. Chem. Phys., vol. 139, no. 12, p. 09B629_1, 2013.
K.E. Theisen, A. Zhmurov, M.E. Newberry, V. Barsegov, R.I. Dima, Multiscale modeling of the nanomechanics of microtubule protofilaments. J. Phys. Chem. B 116(29), 8545–8555 (2012)
M. I. Molodtsov, E. L. Grishchuk, A. K. Efremov, J. R. McIntosh, and F. I. Ataullakhanov, “Force production by depolymerizing microtubules: A theoretical study,” Proc. Natl. Acad. Sci. U. S. A., vol. 102, no. 12, pp. 4353 LP – 4358, Mar. 2005, doi: https://doi.org/10.1073/pnas.0501142102.
Y. Ding, Z. Xu, Mechanics of Microtubules from a Coarse-Grained Model. Bionanoscience 1(4), 173–182 (2011). https://doi.org/10.1007/s12668-011-0027-0
V. VanBuren, L. Cassimeris, D.J. Odde, Mechanochemical Model of Microtubule Structure and Self-Assembly Kinetics. Biophys. J. 89(5), 2911–2926 (2005). https://doi.org/10.1529/biophysj.105.060913
M. L. Gardel, J. H. Shin, F. C. MacKintosh, L. Mahadevan, P. Matsudaira, and D. A. Weitz, “Elastic Behavior of Cross-Linked and Bundled Actin Networks,” Science (80-. )., vol. 304, no. 5675, pp. 1301 LP – 1305, May 2004, doi: https://doi.org/10.1126/science.1095087.
L. Mahadevan, T.J. Mitchison, Powerful curves. Nature 435(7044), 895–897 (2005). https://doi.org/10.1038/435895a
Z. Wu, H.-W. Wang, W. Mu, Z. Ouyang, E. Nogales, J. Xing, Simulations of Tubulin Sheet Polymers as Possible Structural Intermediates in Microtubule Assembly. PLoS ONE 4(10), e7291 (Oct. 2009)
S. Kasas, A. Kis, B.M. Riederer, L. Forró, G. Dietler, S. Catsicas, Mechanical Properties of Microtubules Explored Using the Finite Elements Method. ChemPhysChem 5(2), 252–257 (Feb. 2004). https://doi.org/10.1002/cphc.200300799
A. Kis et al., Nanomechanics of Microtubules. Phys. Rev. Lett. 89(24), 248101 (Nov. 2002). https://doi.org/10.1103/PhysRevLett.89.248101
P.J. de Pablo, I.A.T. Schaap, F.C. MacKintosh, C.F. Schmidt, Deformation and Collapse of Microtubules on the Nanometer Scale. Phys. Rev. Lett. 91(9), 98101 (Aug. 2003). https://doi.org/10.1103/PhysRevLett.91.098101
I.A.T. Schaap, C. Carrasco, P.J. de Pablo, F.C. MacKintosh, C.F. Schmidt, Elastic Response, Buckling, and Instability of Microtubules under Radial Indentation. Biophys. J. 91(4), 1521–1531 (2006). https://doi.org/10.1529/biophysj.105.077826
S. Kasas et al., Oscillation modes of microtubules. Biol. Cell 96(9), 697–700 (2004). https://doi.org/10.1016/j.biolcel.2004.09.002
K.M. Liew, P. Xiang, L.W. Zhang, Mechanical properties and characteristics of microtubules: a review. Compos. Struct. 123, 98–108 (2015)
B. Gu, Y.-W. Mai, C.Q. Ru, Mechanics of microtubules modeled as orthotropic elastic shells with transverse shearing. Acta Mech. 207(3–4), 195–209 (2009)
C.Y. Wang, L.C. Zhang, Circumferential vibration of microtubules with long axial wavelength. J. Biomech. 41(9), 1892–1896 (2008). https://doi.org/10.1016/j.jbiomech.2008.03.029
X.S. Qian, J.Q. Zhang, C.Q. Ru, Wave propagation in orthotropic microtubules. J. Appl. Phys. 101(8), 84702 (Apr. 2007). https://doi.org/10.1063/1.2717573
C. Li, C.Q. Ru, A. Mioduchowski, Torsion of the central pair microtubules in eukaryotic flagella due to bending-driven lateral buckling. Biochem. Biophys. Res. Commun. 351(1), 159–164 (2006). https://doi.org/10.1016/j.bbrc.2006.10.019
Ö. Civalek, Ç. Demir, Bending analysis of microtubules using nonlocal Euler-Bernoulli beam theory. Appl. Math. Model. 35(5), 2053–2067 (2011). https://doi.org/10.1016/j.apm.2010.11.004
Ç. Demir, Ö. Civalek, Torsional and longitudinal frequency and wave response of microtubules based on the nonlocal continuum and nonlocal discrete models. Appl. Math. Model. 37(22), 9355–9367 (2013). https://doi.org/10.1016/j.apm.2013.04.050
H.-S. Shen, Buckling and postbuckling of radially loaded microtubules by nonlocal shear deformable shell model. J. Theor. Biol. 264(2), 386–394 (2010). https://doi.org/10.1016/j.jtbi.2010.02.014
H.-S. Shen, Nonlocal shear deformable shell model for postbuckling of axially compressed microtubules embedded in an elastic medium. Biomech. Model. Mechanobiol. 9(3), 345–357 (2010). https://doi.org/10.1007/s10237-009-0180-3
H.-S. Shen, Nonlocal shear deformable shell model for bending buckling of microtubules embedded in an elastic medium. Phys. Lett. A 374(39), 4030–4039 (2010). https://doi.org/10.1016/j.physleta.2010.08.006
Y.J. Shi, W.L. Guo, C.Q. Ru, Relevance of Timoshenko-beam model to microtubules of low shear modulus. Phys. E Low-dimensional Syst. Nanostructures 41(2), 213–219 (2008)
T. Li, A mechanics model of microtubule buckling in living cells. J. Biomech. 41(8), 1722–1729 (2008)
Z. Wu, E. Nogales, J. Xing, Comparative studies of microtubule mechanics with two competing models suggest functional roles of alternative tubulin lateral interactions. Biophys. J. 102(12), 2687–2696 (2012)
T. Hawkins, M. Mirigian, M.S. Yasar, J.L. Ross, Mechanics of microtubules. J. Biomech. 43(1), 23–30 (2010)
S. Nishimura et al., Microtubules modulate the stiffness of cardiomyocytes against shear stress. Circ. Res. 98(1), 81–87 (2006)
H. Wada and R. R. Netz, “Non-equilibrium hydrodynamics of a rotating filament,” EPL (Europhysics Lett., vol. 75, no. 4, p. 645, 2006.
C.Y. Wang, C.Q. Ru, A. Mioduchowski, Vibration of microtubules as orthotropic elastic shells. Phys. E Low-dimensional Syst. Nanostructures 35(1), 48–56 (2006)
C.Y. Wang, C.Q. Ru, A. Mioduchowski, Orthotropic elastic shell model for buckling of microtubules. Phys. Rev. E 74(5), 52901 (2006)
Ö. Civalek, C. Demir, A simple mathematical model of microtubules surrounded by an elastic matrix by nonlocal finite element method. Appl. Math. Comput. 289, 335–352 (2016)
M.A. Deriu, T.C. Bidone, G. Grasso, A. Acquaviva, U. Morbiducci, Multiscale modeling of microtubules and actin filaments. IFAC Proc. 45(2), 1023–1028 (2012)
M. Soheilypour, M. Peyro, S.J. Peter, M.R.K. Mofrad, Buckling behavior of individual and bundled microtubules. Biophys. J. 108(7), 1718–1726 (2015)
A. Shahinnejad, M. Haghpanahi, F. Farmanzad, Finite Element Analysis of Axonal Microtubule Bundle under Tension and Torsion. Procedia Eng. 59, 16–24 (2013). https://doi.org/10.1016/j.proeng.2013.05.088
Y. Gao, J. Wang, H. Gao, Persistence length of microtubules based on a continuum anisotropic shell model. J. Comput. Theor. Nanosci. 7(7), 1227–1237 (2010)
A. Shamloo, F. Manuchehrfar, H. Rafii-Tabar, A viscoelastic model for axonal microtubule rupture. J. Biomech. 48(7), 1241–1247 (2015)
M. Kolarova, F. García-Sierra, A. Bartos, J. Ricny, and D. Ripova, “Structure and pathology of tau protein in Alzheimer disease,” Int. J. Alzheimer’s Dis., vol. 2012, 2012.
K.J. Rosenberg, J.L. Ross, H.E. Feinstein, S.C. Feinstein, J. Israelachvili, Complementary dimerization of microtubule-associated tau protein: Implications for microtubule bundling and tau-mediated pathogenesis. Proc. Natl. Acad. Sci. 105(21), 7445–7450 (2008)
B. Isralewitz, M. Gao, K. Schulten, Steered molecular dynamics and mechanical functions of proteins. Curr. Opin. Struct. Biol. 11(2), 224–230 (2001)
Y. Fichou, M. Heyden, G. Zaccai, M. Weik, D.J. Tobias, Molecular dynamics simulations of a powder model of the intrinsically disordered protein Tau. J. Phys. Chem. B 119(39), 12580–12589 (2015)
J. Li et al., An overview of predictors for intrinsically disordered proteins over 2010–2014. Int. J. Mol. Sci. 16(10), 23446–23462 (2015)
L.A. Kelley, S. Mezulis, C.M. Yates, M.N. Wass, M.J.E. Sternberg, The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10(6), 845 (2015)
Y. Zhang, I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9(1), 40 (2008)
Y. Luo et al., Molecular insights into the reversible formation of tau protein fibrils. Chem. Commun. 49(34), 3582–3584 (2013)
S. Wegmann, J. Schöler, C.A. Bippes, E. Mandelkow, D.J. Muller, Competing interactions stabilize pro-and anti-aggregant conformations of human Tau. J. Biol. Chem. 286(23), 20512–20524 (2011)
Y.S. Jho, E.B. Zhulina, M.-W. Kim, P.A. Pincus, Monte carlo simulations of tau proteins: effect of phosphorylation. Biophys. J. 99(8), 2387–2397 (2010)
A.J. Lyons, N.S. Gandhi, R.L. Mancera, Molecular dynamics simulation of the phosphorylation-induced conformational changes of a tau peptide fragment. Proteins Struct. Funct. Bioinforma. 82(9), 1907–1923 (2014)
T.G. Castro, F.-D. Munteanu, A. Cavaco-Paulo, Electrostatics of tau protein by molecular dynamics. Biomolecules 9(3), 116 (2019)
A. Battisti, A. Tenenbaum, Molecular dynamics simulation of intrinsically disordered proteins. Mol. Simul. 38(2), 139–143 (2012)
D. Dułak et al., Filamentous aggregates of tau proteins fulfil standard amyloid criteria provided by the fuzzy oil drop (FOD) model. Int. J. Mol. Sci. 19(10), 2910 (2018)
L. Jayanthi, “Computational Investigation On The Structural Properties Of Neurofilaments And Their Sidearms,” 2014.
R. Beck, J. Deek, and C. R. Safinya, “Structures and interactions in ‘bottlebrush’neurofilaments: the role of charged disordered proteins in forming hydrogel networks.” Portland Press Limited, 2012.
S.P. Adiga, D.W. Brenner, Molecular Basis for Neurofilament Heavy Chain Side Arm Structure Modulation by Phosphorylation. J. Phys. Chem. C 114(12), 5410–5416 (Apr. 2010). https://doi.org/10.1021/jp905671u
L. Jayanthi, W. Stevenson, Y. Kwak, R. Chang, Y. Gebremichael, Conformational properties of interacting neurofilaments: Monte Carlo simulations of cylindrically grafted apposing neurofilament brushes. J. Biol. Phys. 39(3), 343–362 (2013)
W. Stevenson, R. Chang, Y. Gebremichael, Phosphorylation-mediated conformational changes in the mouse neurofilament architecture: insight from a neurofilament brush model. J. Mol. Biol. 405(4), 1101–1118 (2011)
J. Lee, S. Kim, R. Chang, L. Jayanthi, Y. Gebremichael, Effects of molecular model, ionic strength, divalent ions, and hydrophobic interaction on human neurofilament conformation. J. Chem. Phys. 138(1), 01B604 (2013)
R. Chang, Y. Kwak, Y. Gebremichael, Structural properties of neurofilament sidearms: sequence-based modeling of neurofilament architecture. J. Mol. Biol. 391(3), 648–660 (2009)
S. Kumar, X. Yin, B.D. Trapp, J.H. Hoh, M.E. Paulaitis, Relating interactions between neurofilaments to the structure of axonal neurofilament distributions through polymer brush models. Biophys. J. 82(5), 2360–2372 (2002)
M.J. Stevens, J.H. Hoh, Conformational dynamics of neurofilament side-arms. J. Phys. Chem. B 114(27), 8879–8886 (2010)
M.J. Stevens, J.H. Hoh, Interactions between planar grafted neurofilament side-arms. J. Phys. Chem. B 115(23), 7541–7549 (2011)
S. Kim, R. Chang, C. Teunissen, Y. Gebremichael, A. Petzold, Neurofilament stoichiometry simulations during neurodegeneration suggest a remarkable self-sufficient and stable in vivo protein structure. J. Neurol. Sci. 307(1–2), 132–138 (2011)
E.B. Zhulina, F.A.M. Leermakers, Effect of the ionic strength and pH on the equilibrium structure of a neurofilament brush. Biophys. J. 93(5), 1452–1463 (2007)
E.B. Zhulina, F.A.M. Leermakers, A self-consistent field analysis of the neurofilament brush with amino-acid resolution. Biophys. J. 93(5), 1421–1430 (2007)
E.B. Zhulina, F.A.M. Leermakers, The polymer brush model of neurofilament projections: effect of protein composition. Biophys. J. 98(3), 462–469 (2010)
S. Kumar, A. Mansson, Covalent and non-covalent chemical engineering of actin for biotechnological applications. Biotechnol. Adv. 35(7), 867–888 (2017)
T. Splettstoesser, K.C. Holmes, F. Noé, J.C. Smith, Structural modeling and molecular dynamics simulation of the actin filament. Proteins Struct. Funct. Bioinforma. 79(7), 2033–2043 (2011)
J.-W. Chu, G.A. Voth, Coarse-grained modeling of the actin filament derived from atomistic-scale simulations. Biophys. J. 90(5), 1572–1582 (2006)
T. Oda, M. Iwasa, T. Aihara, Y. Maéda, A. Narita, The nature of the globular-to fibrous-actin transition. Nature 457(7228), 441 (2009)
R. Dominguez, K.C. Holmes, Actin structure and function. Annu. Rev. Biophys. 40, 169–186 (2011)
J. Pfaendtner, E. Lyman, T.D. Pollard, G.A. Voth, Structure and dynamics of the actin filament. J. Mol. Biol. 396(2), 252–263 (2010)
J. Pfaendtner, E. M. De La Cruz, and G. A. Voth, “Actin filament remodeling by actin depolymerization factor/cofilin,” Proc. Natl. Acad. Sci., vol. 107, no. 16, pp. 7299 LP – 7304, Apr. 2010, doi: https://doi.org/10.1073/pnas.0911675107.
T. Li, Y. Gu, X.-Q. Feng, P.K.D.V. Yarlagadda, A. Oloyede, Hierarchical multiscale model for biomechanics analysis of microfilament networks. J. Appl. Phys. 113(19), 194701 (2013)
T. Kim, W. Hwang, R.D. Kamm, Computational Analysis of a Cross-linked Actin-like Network. Exp. Mech. 49(1), 91–104 (2009). https://doi.org/10.1007/s11340-007-9091-3
M.M.A.E. Claessens, M. Bathe, E. Frey, A.R. Bausch, Actin-binding proteins sensitively mediate F-actin bundle stiffness. Nat. Mater. 5(9), 748–753 (2006). https://doi.org/10.1038/nmat1718
X. Zheng, K. Diraviyam, D. Sept, Nucleotide Effects on the Structure and Dynamics of Actin. Biophys. J. 93(4), 1277–1283 (2007). https://doi.org/10.1529/biophysj.107.109215
J. Pfaendtner, D. Branduardi, M. Parrinello, T. D. Pollard, and G. A. Voth, “Nucleotide-dependent conformational states of actin,” Proc. Natl. Acad. Sci., vol. 106, no. 31, pp. 12723 LP – 12728, Aug. 2009, doi: https://doi.org/10.1073/pnas.0902092106.
P. Dalhaimer, T.D. Pollard, B.J. Nolen, Nucleotide-Mediated Conformational Changes of Monomeric Actin and Arp3 Studied by Molecular Dynamics Simulations. J. Mol. Biol. 376(1), 166–183 (2008). https://doi.org/10.1016/j.jmb.2007.11.068
J.Y. Lee, T.M. Iverson, R.I. Dima, Molecular Investigations into the Mechanics of Actin in Different Nucleotide States. J. Phys. Chem. B 115(1), 186–195 (Jan. 2011). https://doi.org/10.1021/jp108249g
S. Matsushita, Y. Inoue, T. Adachi, Quantitative analysis of extension–torsion coupling of actin filaments. Biochem. Biophys. Res. Commun. 420(4), 710–713 (2012)
S. Matsushita, T. Adachi, Y. Inoue, M. Hojo, M. Sokabe, Evaluation of extensional and torsional stiffness of single actin filaments by molecular dynamics analysis. J. Biomech. 43(16), 3162–3167 (2010)
J.I. Kim, J. Kwon, I. Baek, S. Na, Steered molecular dynamics analysis of the role of cofilin in increasing the flexibility of actin filaments. Biophys. Chem. 218, 27–35 (2016). https://doi.org/10.1016/j.bpc.2016.08.002
J. Jeon, N.R. Alexander, A.M. Weaver, P.T. Cummings, Protrusion of a Virtual Model Lamellipodium by Actin Polymerization: A Coarse-Grained Langevin Dynamics Model. J. Stat. Phys. 133(1), 79 (2008). https://doi.org/10.1007/s10955-008-9600-5
D. Ming, Y. Kong, Y. Wu, J. Ma, Simulation of F-Actin Filaments of Several Microns. Biophys. J. 85(1), 27–35 (2003). https://doi.org/10.1016/S0006-3495(03)74451-8
M.G. Saunders, G.A. Voth, Comparison between Actin Filament Models: Coarse-Graining Reveals Essential Differences. Structure 20(4), 641–653 (2012). https://doi.org/10.1016/j.str.2012.02.008
M.A. Deriu et al., Multiscale modeling of cellular actin filaments: From atomistic molecular to coarse-grained dynamics. Proteins Struct. Funct. Bioinforma. 80(6), 1598–1609 (2012)
J. Fan, M.G. Saunders, G.A. Voth, Coarse-graining provides insights on the essential nature of heterogeneity in actin filaments. Biophys. J. 103(6), 1334–1342 (2012)
O.N. Yogurtcu, J.S. Kim, S.X. Sun, A mechanochemical model of actin filaments. Biophys. J. 103(4), 719–727 (2012)
J.-W. Chu, G.A. Voth, Allostery of actin filaments: molecular dynamics simulations and coarse-grained analysis. Proc. Natl. Acad. Sci. 102(37), 13111–13116 (2005)
G.A. Holzapfel, M.J. Unterberger, R.W. Ogden, An affine continuum mechanical model for cross-linked F-actin networks with compliant linker proteins. J. Mech. Behav. Biomed. Mater. 38, 78–90 (2014). https://doi.org/10.1016/j.jmbbm.2014.05.014
M.J. Unterberger, K.M. Schmoller, A.R. Bausch, G.A. Holzapfel, A new approach to model cross-linked actin networks: multi-scale continuum formulation and computational analysis. J. Mech. Behav. Biomed. Mater. 22, 95–114 (2013)
T. Li, “Cross-scale biophysics modelling of F-actin cytoskeleton in cell.” Queensland University of Technology, 2015.
M.J. Unterberger, K.M. Schmoller, C. Wurm, A.R. Bausch, G.A. Holzapfel, Viscoelasticity of cross-linked actin networks: Experimental tests, mechanical modeling and finite-element analysis. Acta Biomater. 9(7), 7343–7353 (2013). https://doi.org/10.1016/j.actbio.2013.03.008
S. Suresh, Biomechanics and biophysics of cancer cells. Acta Mater. 55(12), 3989–4014 (2007)
F. Gittes, B. Mickey, J. Nettleton, J. Howard, Flexural rigidity of microtubules and actin filaments measured from thermal fluctuations in shape. J. Cell Biol. 120(4), 923–934 (1993)
N.Y. Yao, C.P. Broedersz, Y.-C. Lin, K.E. Kasza, F.C. MacKintosh, D.A. Weitz, Elasticity in ionically cross-linked neurofilament networks. Biophys. J. 98(10), 2147–2153 (2010)
J.F. Leterrier, J. Käs, J. Hartwig, R. Vegners, P.A. Janmey, Mechanical effects of neurofilament cross-bridges modulation by phosphorylation, lipids, and interactions with f-actin. J. Biol. Chem. 271(26), 15687–15694 (1996)
O.I. Wagner, S. Rammensee, N. Korde, Q. Wen, J.-F. Leterrier, P.A. Janmey, Softness, strength and self-repair in intermediate filament networks. Exp. Cell Res. 313(10), 2228–2235 (2007)
J.I. Kim, J. Kwon, I. Baek, H.S. Park, S. Na, Cofilin reduces the mechanical properties of actin filaments: approach with coarse-grained methods. Phys. Chem. Chem. Phys. 17(12), 8148–8158 (2015). https://doi.org/10.1039/C4CP06100D
A. Battisti, G. Ciasca, A. Grottesi, A. Bianconi, A. Tenenbaum, Temporary secondary structures in tau, an intrinsically disordered protein. Mol. Simul. 38(7), 525–533 (2012)
Acknowledgements and Funding
This work has been funded by the Computational Cellular Biology of Blast (C2B2) program through the Office of Naval Research (ONR) (Award # N00014-18–1-2082- Dr. Timothy Bentley, Program Manager).
Author information
Authors and Affiliations
Contributions
M.I.K. collected and interpreted the literature reviews. A.A. contributed in revising the manuscript. F.H. and K.A.M. contributed in reviewing the manuscript.
Corresponding author
Ethics declarations
Conflict of Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Khan, M.I., Hasan, F., Mahmud, K.A.H.A. et al. Recent Computational Approaches on Mechanical Behavior of Axonal Cytoskeletal Components of Neuron: A Brief Review. Multiscale Sci. Eng. 2, 199–213 (2020). https://doi.org/10.1007/s42493-020-00043-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42493-020-00043-4