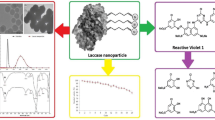
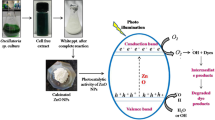
Abstract
Azo dyes released by the textile industries cause severe damage to the environment and living organisms. The degradation of azo dyes is widely studied using enzymatic methods. Laccase is a copper-containing enzyme that degrades the azo dyes into less toxic compounds. In this work, the crude laccase enzyme produced by the alkaliphile Pseudomonas mendocina in the degradation of mixed azo dye showed 0.386 U/mL activity at pH 8.5. A combination of enzymatic and green synthesized nanoparticles was used in the degradation of mixed azo dye. Laccase used in the degradation of mixed azo dyes showed 58.4% in 72 h, while the photocatalytic degradation of mixed azo dyes showed 15.9%. The degradation of azo dyes using copper iodide nanoparticles resulted in 15.8% degradation. However, it was noticed that the combined method of degradation of azo dyes involving both crude laccase and CuI nanoparticles gave a degradation of 62.3% in 60 min. Interaction of laccase enzyme with azo dyes using in silico analysis predicted the binding energy with reactive red (−7.19 kcal/mol), reactive brown (−8.57 kcal/mol), and reactive black dyes (−9.17 kcal/mol) respectively.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The increase in urbanization and population leads to an increase in dyes, plastics, and chemicals that add significant pollutants to the environment. Dyes are complex structural compounds used in the textile, food, and pharmaceutical industries [1]. Azo dyes are synthetic dyes used to give permanent color to fabrics as they are highly resistant to external factors. More than 70% of textile industries use azo dyes for dying fabrics. Its effluent released into the environment possess a significant contribution to environmental pollution [2]. The azo dye contains a chromophore azo group (N = N), which gives color to the fabrics. The azo dyes may be classified as mono azo, diazo, or poly azo dye based on several azo groups present in them [3]. The azo dyes released into the environment, either untreated or partially treated, lead to soil pollution, water, and other ecosystems as azo dyes are carcinogenic and mutagenic in nature [4]. The phytoplankton, zooplankton, fishes, aquatic plants are affected by decreased oxygen level and decreased photosynthetic activity, resulting in aquatic habitats' death. It also affects human beings by causing jaundice, tumor, skin irritation, allergies, and heart defects [5,6,7]. These effects have drawn interest in treating the azo dye contaminated sites by various physical, chemical, and biological methods. Physical methods such as adsorption and chemical methods such as Fenton's reagent and other techniques such as Advanced Oxidation Process (AOP), cavitation, ultrasound technique were studied exclusively [3]. The physical and chemical methods are highly toxic, and hence, biological methods are an alternative eco-friendly technique for treating azo dyes [2, 8]. The biological process involves aerobic and anaerobic degradation of azo dyes by microorganisms. The extremophiles, mainly alkaliphiles, have drawn interest to study the degradation of azo dyes. As the nature of azo dye contaminated sites is alkaline, it makes the survival of mesophiles difficult. Alkaliphiles are microorganisms whose cell wall is composed of acid polymers, which maintain the microorganisms internal pH and make them survive in extreme alkaline environments [9, 10]. The degradation of azo dyes by microbes occurs through enzymatic cleavage. The enzymatic degradation of azo dyes by the bacteria occurs either by extracellular or intracellular enzymes. The enzymes that are commonly involved in azo dye degradation are azoreductase, laccase, peroxidases, etc. These enzymes require appropriate environmental conditions to maintain their stability and activity [11]. Laccase is copper-containing oxidases that do not require redox mediators for degrading azo dyes [12, 13]. The substrate specificity of laccase can be widened by adding redox mediators to the reaction mixture [14]. Laccases cleave azo dyes and form nitrogen molecules that are non-toxic [15].
Photocatalytic degradation is one of the AOP used in the bulk treatment of pollutants. This process results in the mineralization of the recalcitrant compounds [16]. Recently, nanosize materials have been widely studied because of their better properties than bulk size particle. They exhibit a large specific surface area and thus have distinct catalytic and thermal properties. Copper iodide is a highly versatile compound having many applications in adsorption studies, catalysis, solar cells, etc. [17]. The green synthesis of copper iodide nanoparticles gives a highly pure product involving the use of minimum chemicals. Photo-assisted catalysis using a semiconductor has been recognized as a promising approach for eliminating many organic pollutants, such as azo dyes [18].
In this present study, the degradation of individual and mixed textile azo dyes was studied using a crude laccase enzyme-producing alkaliphilic bacterium isolated from an azo dye contaminated site. Although there are few reports on photodegradation using green synthesized metal oxides nanoparticles, there is hardly any data on dye degradation involving a combination of green synthesized nanoparticles and bacterial laccase. So, it was thought to enhance mixed azo dye degradation by immobilizing bacterial laccase on CuI nanoparticles. The analysis of the degradation of azo dyes was done using UV–visible Spectrophotometer and HPLC. The dyes were also analyzed using the in silico analysis, which showed the binding of laccase enzyme with reactive red (RR), reactive brown (RB), and reactive black (RBL). The binding also correlates with the degradation of the azo dyes.
2 Materials and methods
2.1 Sample collection
The textile dye contaminated soil samples were collected from Erode (textile industry). Three different soil (different colors) were collected and used to isolate azo dye degrading bacterial strains. The textile dyes RR, RB, and RBL were purchased from Bengal ChemiColor and Co, Parrys, Chennai, India.
2.2 Isolation of alkaliphilic bacterial strain
The bacterial strains were isolated from the textile effluent contaminated soil samples enriched in the Mineral Salts Medium (MSM) with 100 mg/L of mixed azo dye. The MSM was prepared using disodium hydrogen phosphate (Na2HPO4—12.8 g/L), potassium dihydrogen phosphate (KH2PO4—3 g/L), ammonium chloride (NH4Cl – 1 g/L), sodium chloride (NaCl – 0.5 g/L), magnesium sulphate (MgSO4—10 mg/L), 0.01 M calcium chloride (CaCl2) and 20% glucose (pH 8.5). The medium was sterilized, cooled with a mixed azo dye-containing RR, RB, and RBL; each of 100 mg/L was added in 250 mL Erlenmeyer flask. The bacterial strains were isolated on MSM Agar, and each strain's pure culture was stored in 15% glycerol at −20 °C for further studies [19].
2.3 Degradation assay
The dye's maximum absorbance (RR, RB, RBL, and Mixed) was determined using a UV Vis spectrophotometer. The culture was provided with 100 mg/L of mixed azo dye in MSM, and the decolorization was monitored every 24 h for 4 days. The degradation percentage was calculated using the formula [15].
where D–degradation, Ai–initial absorbance, At–Absorbance after time t.
2.4 Biodegradation of azo dyes
The biological method for mixed and individual azo dye degradation was carried out using a laccase enzyme produced by isolated bacterial strain DY2. The enzymatic degradation was studied further by the extraction of extracellular and intracellular proteins. The efficiency of the degradation of mixed and individual azo dyes was studied.
2.5 Extraction of laccase enzyme
The presence of laccase enzyme was screened in extracellular and intracellular extracts of the isolated bacterial strain. The laccase production by the bacterial strain DY2KVG was determined by inoculating the bacterium in 100 mL MSM broth was amended with 100 mg/L of mixed azo dye. The culture was incubated at 37 °C and centrifuged at 10,000 g for 10 min; then, the supernatant was filtered and used for further studies [20]. The intracellular laccase enzyme was extracted by suspending the pellet in 50 mmol−1 of potassium phosphate buffer of pH 7.4, then sonicated using ultra probe sonicator at 60 amp for 4 min with 1 min interval at 4 °C. The supernatant was filtered using a 0.44 µ millipore filter. The filtrate was then assayed for laccase enzyme activity [21].
Crude laccase activity was determined by 2,2ˈ-azino-bis (3-ethylbenzothiazoline-6-sulphonic acid) (ABTS) substrate oxidation. The assay was carried out using 0.5 mL of ABTS dissolved in 0.1 M citrate buffer (pH—4.5) and 0.5 mL of crude enzyme. The reaction mixture was then incubated at 37 °C, 120 rpm, for 10 min [12]. Absorbance was observed at 379 nm by a UV/Vis spectrophotometer. One enzyme activity unit was defined as the amount of enzyme required to reduce the absorbance of 0.01/sec [22]. The crude laccase enzyme activity in U/mL was calculated using the formula given below
where, E.A–enzyme activity, A–absorbance at 379 nm, V–the total volume of a mixture (mL), t–incubation time, e–extinction coefficient, v–the volume of the enzyme (mL). The molar extinction coefficient was calculated and found to be 1.398 M−1 cm−1 [23].
2.6 Effect of pH and temperature on laccase
The pH and temperature were optimized to determine the maximum enzyme activity of crude laccase produced by DY2KVG bacterial strain by varying the buffer's pH and the incubation temperature used in the assay. The crude laccase activity for pH of various ranges (5.8–8.0) using 50 mM potassium phosphate buffer and pH of 9.0 using 50 mM glycine –NaOH buffer was used for optimization [24]. The temperature ranging from 20 to 50 °C was used to study the effect of temperature on the laccase activity. The enzyme activity of crude laccase was monitored using a standard laccase assay procedure after 10 min of incubation [23].
2.7 Azo dye degradation by photocatalysis
The photocatalytic degradation of azo dyes (RR, RB, RBL, and mixed dyes) was carried out in a 200 mL capacity batch reactor. The reactor was fed with 50 mL of MSM at neutral pH and 50 mL of 100 mg/L azo dye (RR, RB, RBL, and mixed dye). The degradation was carried out with UV irradiation placed parallel to the quartz tube with continuous stirring. The temperature was maintained at 21 °C ± 0.2 °C. The samples were withdrawn every 10 min up to 1 h, and the samples were scanned from 200–800 nm [25].
2.8 Synthesis of CuI nanoparticles using Hibiscus rosa—Sinensis L. flower extract
AnalR-Grade CuSO4.5H2O and KI were used as precursors without further purification. CuI nanoparticles were prepared using Hibiscus rosa—Sinensis L. flower extract by the method reported in the literature [26, 27]. The flower extract was added dropwise into CuSO4.5H2O solution under magnetic stirring followed by dropwise addition of KI solution. The obtained mixture was stirred for a further 30 min at room temperature. The resultant off-white precipitate was filtered, washed thoroughly with 50% ethanol, and dried at 60 °C. The synthesized CuI nanoparticles were used in the degradation of azo dyes.
2.9 Degradation of azo dyes by CuI nanoparticles
The photocatalytic degradation of azo dyes (RR, RB, RBL, and mixed dyes) was carried out in the same reactor used for photocatalysis. In this process of degradation, 25 mg of CuI nanoparticles were added in addition to MSM (50 mL) and 100 mg/L of azo dyes (50 mL) as in photocatalytic degradation using UV light [25].
2.10 Combined method for degradation of azo dyes
The efficiency of the crude laccase enzyme to degrade azo dyes was increased by immobilizing the laccase enzyme onto CuI nanoparticle. CuI NP—10 mg, 1 mL of crude laccase enzyme with 0.341 U/mL of enzyme activity, and 1 mL of potassium phosphate buffer (pH—8) were added and incubated in a shaker for 24 h. The mixture was centrifuged at 10,000 g for 10 min and washed twice with a phosphate buffer of pH 8 [28, 29]. The immobilized crude laccase enzyme was introduced into the reactor containing an equal volume of azo dyes, MSM, and the degradation percentage was calculated [25].
2.11 Characterization of CuI nanoparticle and metabolites of degradation
The synthesized and enzyme immobilized CuI nanoparticles were characterized morphologically using HR-SEM analysis (FEI-Quanta FEG 200F). The degradation of RR, RB, RBL, and mixed azo dye were analyzed by high-performance liquid chromatography (Shimadzu Prominence binary gradient HPLC system). Structural analysis of XRD was performed using X-ray powder diffraction (Bruker D8 advance P–XRD), growth, enzyme assay, and preliminary degradation were carried out using UV spectrophotometer (Hitachi, double beam).
2.12 Analysis of azo dye degradation
The high-performance liquid chromatography (HPLC) was carried out in the C18 column by isocratic elution using methanol. The samples were prepared using ethyl acetate (equal volume of sample and solvent). The solvent phase was extracted and filtered using a membrane filter (0.44µ) and then condensed in a rotary vacuum evaporator. Then 20 µl of the sample was injected manually into the column with methanol as the mobile phase [3].
2.13 Molecular identification of isolated bacterium
The genomic DNA was isolated from the bacterial strain, and PCR amplification was performed. The unknown bacterial strain was given for 16S rRNA sequencing in Amnion Biosequences, Bangalore. The evolutionary relationship between the bacterial strains was determined by constructing a phylogenetic tree using the Maximum Likelihood Method in MEGA7.
2.14 In Silico analysis
The laccase protein sequence of Pseudomonas mendocina was retrieved from the NCBI database with GenBank ID KER98563.1. This sequence was used to model the 3D structure using a Swiss-Model online server with PDB ID 1RW0 as a template [29]. The model was evaluated using the RAMPAGE server [30]. The active site of the protein was predicted using the CastP server [31]. The 3D structures of RR, RB and RBL were retrieved from the PubChem database with CIDs 87291, 137233370, and 135442967 respectively. Molecular dockings were performed twice using AutoDock standalone package [32]. The hydrogen and necessary charges were introduced to the laccase protein, and torsions were set to the reactive dyes. The grid of 60 × 60 × 60 cubic along the x, y, and z-axis were fixed around the laccase active site. AutoGrid and AutoDock were performed consecutively using a Lamarckian Genetic Algorithm. The average binding energy and laccase dyes complexes were obtained as the output. These complexes were visualized using Schrodinger Maestro.
3 Results and discussion
The azo dyes are degraded using conventional methods of which biodegradation is considered eco-friendly. The enzymes produced by the bacterial strains degrade the azo dyes based on the enzyme’s nature and the bacterial strain producing it. The present study focuses on the degradation of azo dyes (RR, RB, RBL, and mixed azo dye) by the laccase enzyme produced by an alkaliphilic bacterial strain. The degradation of azo dyes by photocatalysis and by nanoparticles was also studied. This gave rise to the idea of immobilizing the nanoparticle with the laccase enzyme for azo dye degradation.
3.1 Isolation of azo dye degrading alkaliphile
The azo degrading bacterial strain was enriched using 100 mg/L of a mixed azo dye, a sole carbon source at alkaline pH (8.5) in MSM agar. The isolated bacterium was designated as DY2KVG as given in Figure S1. The growth of the isolated bacterial strain DY2KVG was monitored for four days at 540 nm. The bacterium enters the log phase at the 48th hour, and the decrease in the growth was observed at the 96th hour. The absorbance of the mixed azo dye decreased at the 48th hour of degradation is shown in Figure S2 (a & b). The extracellular crude laccase enzyme activity was found to be 0.1 U/mL at 72 h, which was higher than the enzyme activity of the intracellular enzyme (0.072 U/mL) produced by the bacterium DY2KVG as in Figure S3. Thus, further studies were carried out using the extracellular crude laccase enzyme.
3.2 Optimization of pH and temperature
The pH and temperature of the crude laccase enzyme produced by the bacterial strain isolated from the textile effluent contaminated soil were optimized. The results obtained show that the laccase enzyme activity increased at pH 5.8 (0.386 U/mL) and decreased with an increase in pH. The laccase enzyme activity was found to increase at the pH of 8.0, whose enzyme activity was (0.341 U/mL). This might be due to the bacterium's alkaliphilic nature and the pH of the medium in which the bacterium was isolated. The graph of pH optimization was plotted as in Figure S4. The enzyme activity of the laccase enzyme was also monitored at various temperatures. The enzyme activity (0.769 U/mL) was maximum at 20 °C and decreased with increased temperature and given in Figure S5.
It was reported that laccase enzyme-producing Bacillus sp. with 0. 981 U/mL of enzyme activity at pH 9 and peptone as substrate [23]. The extraction of intracellular laccase from P.desmolyticum but the optimum pH and temperature for the degradation of reactive red and reactive green was 4.5 and 60 °C respectively [20].The laccase enzyme produced by the Podoscypha elegans, which resulted in maximum activity in the pH ranging between 5.5 and 7.0 [33], correlates with the present study results.
3.3 Analysis of azo dyes degradation
In this study, crude laccase enzyme produced by an alkaliphilic bacterial strain degraded 2.8% for RR, 16.4% for RB, 6.2% for RBL, and 58.4% for mixed dyes at the end of 72 h in a single cycle. Laccase enzyme on the degradation of aromatic compounds produces cations that form stable metabolites in the enzyme's presence. The catalytic ability of laccase is due to the presence of four copper atoms as a prosthetic group. Thus the degradation of aromatic compounds by laccase is reported to follow the redox reaction [34]. There was no degradation reported in direct blue 15 by the laccase enzyme produced by the fungus Podoscypha elegans. The maximum degradation of 70% of Congo red and rose Bengal was observed in 72 days, whereas direct blue 15 resulted only in 24.8% degradation [33]. This contradicts the present study, as the crude laccase enzyme produced by an alkaliphilic bacterial strain degraded the reactive azo dyes in 72 h, while the laccase produced by the fungi Podoscypha elegans required 21 days, which resulted in minimum degradation of dyes. The bacterial consortium of P. rettgeri strain HSL1 and Pseudomonas sp. SUK1 showed only up to 22% of 100 mg/L of azo dye degradation in 48 h [3]. In this study, the metabolites produced by the degradation of azo dyes by laccase enzyme was analyzed. The HPLC chromatogram of the RR (Fig. 1), RB (Fig. 2), RBL (Fig. 3), and mixed dye (Fig. 4) showed adecrease in the peak intensity and shift in the retention time of the peaks.
The degradation of azo dyes by the photocatalytic breakdown using UV light was also observed. The percentage of azo dyes' degradation by the UV light in a photoreactor was observed to be 2.6% for RR dye, 23.7% for RB, 3.3% for RBL, and 15.9% for mixed dye at the end of 1 h. Thus, the degradation of reactive azo dyes using the photocatalytic process was not effective. The photodegradation process was reported as an effective method for the degradation of various organic compounds [35]. The complete degradation of 50 mg/L of orange II by photocatalysis was reported in 90 min, while the increase in dye concentration increased the time of degradation of orange II [36].
3.4 Azo dyes degradation using cui nanoparticle
CuI nanoparticles were prepared using Hibiscus rosa-Sinensis L. flower extract as a reducing and capping agent. The structural characterization of CuI was performed using powder X-ray diffractometry analysis. Figure 5a depicts the XRD pattern of synthesized CuI. The XRD pattern shows peaks corresponding to (111), (200), (220), (311), (222), (400), (331), (420), and (422) planes and was found to fit well in the face-centered cubic crystal structure. Using Scherrer's formula, the average crystallite size was calculated to be 89.01 nm. The SEM image (Fig. 5b) clearly shows the flaky triangular morphology of CuI nanoparticles. EDAX in Fig. 5 c shows the presence of only elemental Cu and I indicating the high purity of CuI nanoparticles [26]. The surface morphologies significantly influence the adsorption of molecules. The synthesized CuI nanoparticles triangular in shape can act as a good adsorbent due to their geometrically advantageous properties [37, 38].
3.5 Application of CuI in azo dye degradation
The literature review showed the use of CuI in 95% removal of Cd[33]. The present investigation is the first to employ and report CuI for mixed azo dye degradation. The degradation of azo dyes using CuI nanoparticles resulted in 3.2% for RR, 16.7% for RB, 16.9% for RBL, and 15.8% for mixed dye at the end of 1 h, followed by an increase in absorbance. As degradation increases linearly with the amount of CuI, the increase in absorbance after 1 h could be reduced by increasing the concentration of CuI nanoparticles. It has been reported that the use of strontium titanate in direct and reactive dyes resulted in increased degradation with an increase in the strontium titanate concentration [39].
3.6 Combined method for azo dyes degradation
The degradation of azo dyes by the laccase enzyme, photocatalysis, and using CuI nanoparticles are not sufficient for degrading the high concentration of azo dyes. An effort to enhance the process of azo dye degradation was attempted by immobilizing CuI in laccase enzyme, as shown in Fig. 5d. The EDAX shows the presence of Cu, I, C, O. The presence of K is due to the usage of potassium phosphate buffer during the immobilization process. The degradation of RR, RB, RBL, and mixed azo dyes was increased by immobilizing the laccase enzyme in CuI nanoparticle.
The degradation of RR, RB, RBL, and mixed azo dyes by different degradation methods are tabulated in Table 1.
This method enhanced the percentage of degradation to 29.1%, 75.2%, 75%, and 62.3% for RR, RB, RBL, and Mixed azo dye of 100 mg/L at 60th minute. The degradation of mixed dye was less than that of individual dyes, which may be due to the high concentration (three times the optimum concentration) of mixed dye.
The HPLC analysis of the RR shows the presence of a single peak at RT 2.4. The supplementary Figure S6(a) and Figure S6(b) shows the degraded RR of the less intense peak at 3.4 RT. The RB dye showed an intense peak at RT 2.2, a small peak at RT 3 for non-degraded RB, which is shown in Figure S7 (a) supplementary data shifted to RT 4.2, and the intense peak was significantly reduced at RT 2.5 after degradation by the combined method as shown in Figure S7 (b). The RBL dye (control) showed peaks at 2.5 and 6.1 RT (Figure S8 a), and the degraded RBL dye formed a peak at RT 3.4 in Figure S8 (b). The mixed dye showed an intense peak, as shown in Figure S9 (a & b) at RT 2.3 and 3.0. It was reported by that the reactive black dye (100 mg/L) degradation by the biological-photocatalytic method resulted in 74.9% in 4 h without changes in the UV region [25]. The literature stated that the combination of biological and photocatalytic methods was more efficient than a separate treatment process. This supports the present study in which the RBL and RB showed 75% degradation in 1 h by the combined method.
Biological degradation showed 85% of degradation of RB, while in the combined process, the degradation percentage was found to be 74% without aromatic amine formation. The combined process enhanced the degradation of individual azo dyes compared to mixed azo dyes. The percentage of degradation is also higher compared to biological and physical methods. Thus, the metabolites produced by biological and combined methods of degradation of reactive azo dyes are different, therefore presumed to have different pathways.
3.7 Phylogenetic analysis of Pseudomonas mendocina
The 16S rRNA sequence of DY2KVG was blasted, and it was found to be Pseudomonas mendocina (Genbank Accession number MK346304). The evolutionary history of DY2KVG is illustrated in Figure S10, which was inferred using the UPGMA method [40]. The optimal tree with the sum of branch length = 1759.30814508 is shown. (next to the branches). The evolutionary distances were computed using the Maximum Composite Likelihood method [41] and are in the units of the number of base substitutions per site. The analysis involved 11 nucleotide sequences. Codon positions included were 1st + 2nd + 3rd + Noncoding. All positions containing gaps and missing data were eliminated. There were a total of 944 positions in the final dataset. Evolutionary analyses were conducted in MEGA7 [42].
3.8 In silico analysis
From the Ramachandran plot evaluation using RAMPAGE server, the protein model was found to have 92.7% of amino acids in the favored region. Following amino acids were predicted to be the active site amino acids using CastP server: GLN63, PRO64, ALA65, TRP66, LEU67, VAL74, VAL75, ALA76, GLU77, ALA78, ASP79, PRO80, ALA81, THR82, VAL83, ILE84, ASN89, SER95, ILE96, and ALA97. From the molecular docking analysis, the average binding energies between the laccase and RR, RB, and RBL, were found to be −7.19 kcal/mol, −8.57 kcal/mol, and −9.17 kcal/mol, respectively. The amino acids involved in the binding and degradation of dyes are shown in Fig. 6 a, b, and c. The binding between the protein and the compound is stronger with lesser AutoDock binding energy [32].
From the docking results, we observed that the strongest and weakest binding of laccase was with RBL and RR, respectively. The binding between laccase and RB was found to be in between the RR and RBL. The strongest binding observed between the laccase and RBL could be the reason for the higher degradation of RBL that was seen in our experimental findings.
4 Conclusion
Azo dyes are aromatic compounds used in various industries, and they contribute as primary synthetic dyes in most of the textile industry. These dyes have severe effects on living organisms and pose a significant hazard to the environment. Hence there is a need to remove these anthropogenic compounds from the contaminated environment. There are several methods used for the removal, but still, an eco-friendly and cost-effective way for the removal remains in demand. In the current study, Azo dye degradation was studied by integrating a combination of green synthesized nanoparticles and laccase enzyme produced by a bacterial strain Pseudomonas mendocina. Bacterial laccase produced by Pseudomonas mendocina was immobilized with the green synthesized nanoparticle to improve the mixed azo dye degradation. The degradation of mixed azo dye at the end of 60th minute using laccase enzyme, copper iodide nanoparticles, UV irradiation, and a combined method showed varied responses. By using the crude laccase enzyme produced by Pseudomonas mendocina, an alkaliphilic bacterial strain, the degradation was up to be 58.5%. The photocatalytic method (UV light irradiation) showed only 15.9% of degradation of mixed azo dye. The mixed azo dye degradation using only the CuI nanoparticle synthesized from Hibiscus rosa Sinensis flowers also resulted in only 15.8%. When the laccase enzyme was immobilized with copper iodide nanoparticle, the degradation efficiency of mixed azo dye was 62.3%. The pathway for degradation of azo dyes using laccase enzyme, CuI nanoparticle, and the combined method proved to be best among the individual techniques. Thus, it is concluded that integrating the green synthesized nanoparticle immobilized with the bacterial laccase proves to be more efficient, and the work also indicates a new novel approach for treating the recalcitrant compounds. Pilot-scale studies on the treatment of industrial effluents with the mixed dyes using this integrated approach will provide a better understanding of the mechanisms involved in the removal process. The free laccase enzyme's reusability and the combined CuI and laccase enzyme are yet to be studied. The kinetics of the degradation of individual dyes are in study to determine the order of reaction involved in the azo dye degradation by the enzymatic and combined method.
5 Biorixv preprint
Sridharan, R., Krishnaswamy, V., Murali, A., & Rajagopal, R., Thirumal Kumar, D., George Priya Doss, C. (2019). Integrated Approach On Degaradtion Of Azo Dyes Using Laccase Enzyme And Nanoparticle With Its Interaction By In Silco Analysis. bioRxiv, 677690. https://doi.org/10.1101/677690
References
Shyamala Gowri R, Vijayaraghavan R, Meenambigai P (2014) Microbial degradation of azo dyes-review. Int J Curr Microbiol App Sci 3(3):421–436
Saratale RG, Saratale GD, Kalyani DC, Chang IS, Govindwar SP (2011) Bacterial decolourisation and degradation of azo dyes—a review. J Taiwan Inst Chem Eng 42:138–157
Lade Harshad, Govinwar Sanjay, Paul Diby (2015) Mineralization and detoxification of the carcinogenic azo dye congo red and real textile effluent by a polyurethane foam immobilized microbial consortium in an upflow column bioreactor. Int J Environ Res Public Health 12(6):6894–6918
Kumar Raven GN, Bhat Sumangala K (2012) Fungal degradation of azo dye- Red 3BN and optimization of physic-chemical parameters. Int J Environ Sci 1:17–24
Ramírez-Montoya Luis A, Hernández-Montoya Virginia, Montes-Morán Miguel A, Jáuregui-Rincón Juan, Cervantes Francisco J (2015) Decolourisation of dyes with different molecular properties using free and immobilized laccases from Trametes versicolo. J Mol Liq 212:30–37
Kumar PG, Bhat SK (2012) Fungal degradation of azo dye-Red 3BN and optimization of physico-chemical parameters. Int Res J Biol Sci 2:17–24
Lavanya C, Rajesh D, Sunil C, Sarita S (2014) Degradation of toxic dyes: a review. Int J Curr Microbiol Appl Sci 3(6):189–99
Kobayashi H, Rittmann BE (1982) Microbial removal of hazardous organic compounds. Environ Sci Technol 16:170–183
Boopalan Sujatha, Gayathri Veena, Vasudhevan Vidhya (2017) Alkaliphilic degradation of mixed reactive dyes (C.I. RY 125 and C.I. RB 52) by a moderately alkaliphilic bacterial consortium—optimization and bench scale studies. Basic Res J Microbiol 4:2354–4082
Alain K, Joël Q, Françoise L, Patricia P, Philippe C, Gérard R, Valérie C, Marie-Anne CB (2002) Caminibacter hydrogeniphilus gen. nov., sp. nov., a novel thermophilic, hydrogen-oxidizing bacterium isolated from an East Pacific Rise hydrothermal vent. Intern J Systemat Evolut Microbiol 52(4):1317–1323
Joshni TC, Subramaniam K (2011) Enzymatic degradation of azo dyes- a-review. Int Jounal Envo Sci 6:0976–4402
Asadgol Z, Forootanfar H, Rezaei S, Mahvi AH, Faramarzi MA (2014) Removal of phenol and bisphenol-A catalysed by laccase in aqueous solution. J Environ Health Sci 12(1):93
Pereira L, Coelho AV, Viegas CA, dos Santos MM, Robalo MP, Martins LO (2009) Enzymatic biotransformation od azo dye suan orange G with bacterial cotA—laccase. J Biotechnol 139(1):68–77
Fabbrini M, Glli C, Gentili P (2002) Comparing the catalytic efficiency of some mediators of laccase. J Mol Catal B Enzyme 16:213–240
Zille A, Gornacka B, Rehorek A, Cavaco Paulo A (2005) Degradation of azo dyes by tremates villosa laccase over long periods of oxidative conditions. Appl Environ Microbiol 71(11):6711–6718
Comparelli R, Fanizza E, Curri ML, Cozzoli PD, Mascolo G, Agostiano A (2005) UV induced photocatalytic degradation of azo dyes by organic capped ZnO nanocrystals immobilized onto substrates. Appl Catal B 60(1–2):1–11
Tavakoli F, Salavati-Niasari M, Mohandes F (2013) Green synthesis of flower-like CuI microstructures composed of trigonal nanostructures using pomegranate juice. Mater Lett 100:133–136
Vijayakumar A, Rajagopal R (2016) Green synthesis and characterisation of Copper (I) Iodide nanoparticles using kidney bean seed extract and its anti-bacterial activity. J Sci Eng Res 7:602–609
Agarwal T, Singh R (2012) Bioremedial potentials of a moderately halophilic soil bacterium. JPBMS 19:1–6
Kalme SD, Parshetti GK, Jadhav SU, Govindwar SP (2007) Biodegradation of benzidine based dye direct blue-6 by pseudomonas desmolyticum NCIM 2112. Bioresour Technol 98(7):1405–1410
Dwakar VV, Jadhav UU, Jadhav SU, Govindwar SP (2008) Biodegradation of disperse textile dye brown 3REL by newly isolated Bacillus Sps VUS. J Appl Microbiol 105(1):14–24
Mukhopadhyay A, Dasgupta AK, Chakrabarti K (2013) Thermostability, pH stability and dye degrading activity of a bacterial laccase are enhanced in the presence of Cu2O nanoparticles. Bioresour Technol 127:25–36
Shah AI, Jobanputra JB (2017) Production and partial purification of laccase produced by bacillus species isolated from contaminated soil. Int J Innov Res Sci Eng Technol 6:17862–17869
Neifar M, Chouchane H, Mahjoubi M, Jaouani A, Cherif A (2016) Pseudomonasextremorientalis BU118: a new salt-tolerant laccase-secreting bacterium with biotechnological potential in textile azo dye decolourization. Biotech 6(1):107
Shah MP (2013) Combined application of biological-photocatalytic process in degradation of reactive black dye: an excellent outcome. Am J Microbiol Res 1(4):92–97
Archana KM, Yogalakshmi D, Rajagopal R (2019) Application of green synthesized nanocrystalline CuI in the removal of aqueous Mn(VII) and Cr(VI) ions. SN App Sciences 1(6):522
Indubala E, Dhanasekar M, Sudha V, Malar EP, Divya P, Sherin J, Rajagopal R, Bhat SV, Harinipriya S (2018) L-Alanine capping of ZnO nanorods: increased carrier concentration in ZnO/CuI heterojunction diode. RSC Adv 8(10):5350–5361
Patel SK, Kalia VC, Choi JH, Haw JR, Kim IW, Lee JK (2014) Immobilization of laccase on SiO2 nanocarriers improves its stability and reusability. J Microbiol Biotechnol 24(5):639–647
Biasini M, Bienert S, Waterhouse A, Arnold K, Studer G, Schmidt T, Kiefer F, Cassarino TG, Bertoni M, Bordoli L, Schwede T (2014) SWISS-MODEL: modelling protein tertiary and quaternary structure using evolutionary information. Nucleic acids Res 1(42):W252–W8
Lovell SC, Davis IW, Arendall III WB, De Bakker PI, Word JM, Prisant MG, Richardson JS, Richardson DC (2003) Structure validation by Cα geometry: ϕ, ψ and Cβ deviation. Proteins: Struct Funct Bioinf 50(3): 437–50
Binkowski TA, Naghibzadeh S, Liang J (2003) CASTp: computed atlas of surface topography of proteins. Nucleic Acids Res 31(13):3352–3355
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30(16):2785–2791
Pramanik S, Chaudhuri S (2018) Laccase activity and azo dye decolorization potential of podoscypha elegans. Mycobiology 46(1):79–83
Rubilar O, Diez MC, Gianfreda L (2008) Transformation of chlorinated phenolic compounds by white rot fungi. Crit Rev Environ Sci Technol 38(4):227–268
Daneshvar N, Salari D, Khataee AR (2004) Photocatalytic degradation of azo dye acid red 14 in water on ZnO as an alternative catalyst to TiO2. J Photochem Photobiol A 162(2–3):317–322
Divya N, Bansal A, Jana AK (2013) Photocatalytic degradation of azo dye orange II in aqueous solutions using copper-impregnated titania. Int J Environ Sci Technol 10(6):1265–1274
Guo S, Cao Y, Sun T, Zhang J, Gu L, Zhang C, Xu Z (2018) Influence of surface morphology on adsorption of potassium stearate molecules on diamond-like carbon substrate: molecular dynamics study. Appl Surf Sci 441:708–717
Jiang Y, Gao S, Li Z, Jia X, Chen Y (2011) Cauliflower-like CuI nanostructures: green synthesis and applications as catalyst and adsorbent. Mater SciEng B 176(13):1021–1027
Karimi L, Zohoori S, Yazdanshenas ME (2014) Photocatalytic degradation of azo dyes in aqueous solutions under UV irradiation using nano-strontium titanate as the nanophotocatalyst. J Saudi Chem Soc 18(5):581–588
Sneath PH, Sokal RR (1973) Numerical taxonomy. San Francisco, W.H. Freeman and Company, The principles and practice of numerical classification
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. ProNatAcad Sci 101(30):11030–11035
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets: Mol Biol Evol 33(7): 1870–1874
Acknowledgements
We would like to thank UGC-MRP for funding this project. We would like to thank DST-FIST CRIST LAB, Stella Maris College (Autonomous) for providing the instrumentation facility. We would like to extend our gratitude to SAIF IITM, Department of Chemistry IITM for SEM and XRD analysis and VIT, Vellore.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
There is no conflict of interest for the manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sridharan, R., Krishnaswamy, V., Archana, K.M. et al. Integrated approach on azo dyes degradation using laccase enzyme and Cul nanoparticle. SN Appl. Sci. 3, 370 (2021). https://doi.org/10.1007/s42452-021-04164-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42452-021-04164-9