Abstract
Stripe rust, caused by Puccinia striiformis f. sp. tritici (Pst) is one of the most destructive diseases of wheat (Triticum aestivum L.) worldwide causing huge yield losses every year. Development and deployment of resistant varieties is the most economical and environment friendly approach for controlling this disease. However, because of the continuous evolution of the pathogen, resistant genes are easily overcome by new virulent Pst races, which necessitates a continuous identification and introgression of resistance genes to develop resistant wheat varieties. To identify effective source of resistance, a genome-wide association study was performed using 426 elite bread wheat genotypes based on 5176 polymorphic Diversity Arrays Technology (DArT) markers. Adult-plant-resistance was evaluated under field conditions for yellow rust resistance for two consecutive years (2014 and 2015) at ICARDA Merchouch station, Morocco. Out of the 426 genotypes, 51.17% were highly resistant with 5–10% level of severity to yellow rust. Genome wide association studies (GWAS) using a mixed linear model (MLM) identified three DArT markers on chromosomes 1B, 2B and 7B which are significantly associated with stripe rust resistance at false discovery rate p ≤ 0.05. BLAST analysis confirmed that the marker 412,394 in chromosome 2B overlapped with two previously reported QTLs (QYrlu.cau-2BS1 Luke and QYrid.ui-2B.1_IDO444). However, the two other markers 542,318 (1B) and 583,038 (5B) were not mapped within any of the previously reported gene/QTL regions; therefore, these markers may represent novel resistance loci for yellow rust. The highly resistant elite genotypes and linked molecular markers are recommended for further gene introgression and pyramiding purposes in the wheat breeding programs after validation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bread wheat (Triticum aestivum L.) is one of the most important crops for global food security with an annual production of 760 M tons in 219 M hectares (FAO 2018). Various biotic and abiotic stresses cause significant grain yield losses every year. However, stripe rust caused by Puccinia striiformis f. sp. tritici (Pst), is considered as one of the disastrous fungal diseases of wheat worldwide, causing 5.47 million tons loss yearly (Beddow et al. 2015). During the last decade, several stripe rust epidemics have been causing significant yield reduction with an average loss ranging from 10 to 100% on highly susceptible wheat cultivars (Wang et al. 2016). Since the year 2000, new aggressive virulent races of the pathogen have been detected across the world (Chen et al. 2014; Milus et al. 2009). Epidemics have been reported in East and North Africa, with high disease pressure causing national losses estimated at 20% during 2009 (https://rusttracker.cimmyt.org/).
Resistance genes against rust infections in common wheat can be divided into two main categories: major and minor resistance genes. The major genes were used by Flor (1956) to describe the harmonizing gene-for-gene interaction. These genes have a race-specificity and involves the non-durable nature of such resistant mechanisms. Generally, these genes are implied in a host response to the invading pathogen very early in the infection process and obtain hypersensitivity in which plant host cells that are in close proximity to the invading fungus, or are being attacked by the fungus, undergo programmed cell death. This stops the fungus from setting up feeding structures within the plant, which fundamentally lead to fungal death. History showed that when a single major gene protects a large area of a wheat growing region, the fungus can overcome this resistance in a relatively brief period of time. These types of genes have also been named as seedling genes as they are effective in both seedlings and adult plants. However, more accurate description would be all-stage resistance (Lin and Chen 2007). Minor resistance genes generally have a distinct mode of action and do not supply the exception or the high levelled resistance that a single major gene does. These genes have varyingly been called horizontal, partial, non-race specific, slow-rusting, durable or adult plant resistances (Caldwell 1968: Johnson 1988; Parlevliet 1979). The mechanisms by which fungal disease is inhibited by slight resistances include a rise in the latency period, reduced uredinia size, reduced infection frequency and reduced spore production (Ohm et al. 1976). Yellow rust can be controlled through the application of fungicides, however, chemicals are costly and unfriendly to the environment and social health.
Thus, managing this disease through continuous development and deployment of resistant varieties is highly important for enhancing world wheat production in a sustained manner while protecting the natural resource. Conventional breeding methods offer an efficient and environment friendly way to control yellow rust, whereas it takes more than ten years to develop and release a wheat variety (Beddow et al. 2015; Dong et al. 2017). Therefore, there is an urgent need to breed for stable, durable and broad-spectrum resistance to avoid the quick breakdown of resistant genes due to rapid evolution of new virulent races of the pathogen. Pyramiding diverse resistance genes in new elite genotypes is a successful strategy to achieve this durability using qualitative (major genes) and/or quantitative (minor genes) combinations. Quantitative resistance, often governed by several genes, provides durable resistance (Kushalappa et al. 2016; Niks et al. 2015).
To date, 83 yellow rust resistant genes (Yr) have been permanently characterized; of which 67 have been temporarily designated. More than 300 quantitative trait loci (QTLs) have been reported and mapped to the wheat genome (Chen 2020). Among the formally designated stripe rust resistant genes, 55 genes confer seedling resistance and 24 provide adult plant resistance (APR). Some of these genes have been widely used in wheat breeding programs using functional markers in gene pyramiding programs (Yang et al. 2019).
Genome wide association studies (GWAS) offer an exciting approach to identify marker–trait associations (MTAs) and localize the genomic regions that underly the gene of interest. Association mapping has been successfully used in mapping complex agronomic traits in bread wheat, such as heat and drought tolerance traits (Ballesta et al. 2020; Tadesse et al. 2019) grain quality traits (Tadesse et al. 2015), disease resistance (Puntel et al. 2016; Kankwatsa et al. 2017), and many more.
The recent explosion of stripe rust epidemic in diverse wheat-producing countries in East and North Africa, Middle East and central west Asian countries pretends a serious warning not only to wheat production and economic resources but also to the global food security. The most viable alternative to manage the permanent risk of yellow rust disease is the development of resistant wheat cultivars. The application of modern tools for yellow rust resistance breeding and their advantages over conventional breeding methods helps in developing new cultivars with long lasting resistance (Buerstmayr et al. 2014).
In this study, we investigated the association of 5176 polymorphic DArT markers with yellow rust resistance in 426 wheat genotypes. The panel was phenotyped at Merchouch station in Morocco over two consecutive years, to (i) evaluate the adult plant resistance responses to yellow rust infection under field conditions, (ii) identify elite resistant bread wheat genotypes, (iii) determine closely associated DArT markers to YR resistance, using mixed linear model approach.
Materials and methods
Plant materials
In this study, a population of 426 spring wheat genotypes from the International Center for Agricultural Research in Dry Areas (ICARDA) advanced breeding lines was used. This population was planted using an alpha lattice design in two replications in 3 m2 plot basis at Merchouch, Morocco (33°36’N, 6°43’W, 394 masl) for two cropping seasons (2013/14 and 2014/15) under rainfed conditions. The average temperature was 17.43 °C (max: 22.86 °C; min:12 °C) with 407.48 mm of precipitation recorded for about 72 days of rain per year (https://www.historique-meteo.net). The trials management followed common agronomic practices. Details of the population are provided in Supplementary Table 1.
Disease phenotyping
The 426 wheat genotypes were screened against stripe rust under natural field infection at adult-plant stage. The response for the major infection types was recorded according to Roelfs (1992). Yellow rust severity was assessed as a percentage of leaf surface area covered following modified Cobb’s scale (Peterson et al. 1948). Field responses were assessed 2 to 3 times and the final scoring at soft-dough stage was considered for the association mapping analysis. The data recorded was combined to calculate the coefficient of infection (CI) by multiplying disease severity by a value of 0.2, 0.4, 0.6, 0.8 or 1.0 for the host response of resistance (R), moderately resistant (MR), intermediate (M-MS), moderately susceptible and susceptible (S), respectively (Pathan and Park 2006).
Genotyping
Genomic DNA was extracted as described by Ogbonnaya et al. (2001) from 2-week-old seedlings using pooled leaf samples from 5 plants per line, the plants leaves were frozen in liquid nitrogen and stored at − 80 °C before DNA extraction. Then 10 μl of a 100 ng μL-1 DNA of each sample was sent to Triticarte Pty. Ltd. Australia as a commercial service provider for whole genome scan using 13,698 Diversity Arrays Technology (DArT) markers. The significant associated markers were aligned to the International Wheat Genome Sequencing Consortium (IWGSC RefSeq v2.1) Chinese Spring to find their putative locations on the wheat genome.
Statistical analysis
The scoring of stripe rust was collected in two consecutive years (2014/2015). The collected data was analyzed using R environment software (Allaire 2012). Analysis of variance (ANOVA) were performed across years using a mixed linear model. Genotype and genotype × year were considered as fixed to get the means and their standard errors and were taken as random to estimate the genetic variance and heritability. The heritability (H2) was calculated using the formula as per Knapp et al. (1985) (1):
where \({{\varvec{\sigma}}{\varvec{g}}}^{2}\): genetic variance, \({{\varvec{\sigma}}{\varvec{e}}}^{2}\): error variance, \({\varvec{l}}\): number of environments (years), \({\varvec{l}}{\varvec{r}}\): number of replicates.
Linkage disequilibrium (LD) and population structure analysis
Population structure was assessed by “LEA” R core package (Frichot and Francois 2015) with 272 unlinked markers distributed across the wheat genome with at least two loci on each chromosome. The number of ancestral populations (K) was chosen based on the evaluation of a cross entropy criterion for each K.
Linkage disequilibrium was estimated on filtered genotypic data using Plink1.9 software (Frichot and Francois 2015). We used 5176 polymorphic markers out of 13,698 (MAF 5% and missing data 10%). Allele frequency correlations (R2) were calculated using the LD function according to Hill and Weir (1988). The significance of pair-wise LDs (p values) was computed using 1000 permutations on inter-chromosomal pairs. The LD decay analysis was done using the procedure from Hill and Weir (1988). The genomic relationship matrix (G-matrix) was obtained according to VanRaden (2007) and implemented in the R package GAPIT (Wang and Zhang 2021).
Genome-wide association study (GWAS)
To identify markers associated with responses to yellow rust, GWAS was carried out using 5176 polymorphic markers and the two years average phenotypic stripe rust coefficient of infection were used. The association of the markers retained after quality filtering was carried out using GAPIT v3 (Wang and Zhang 2021). Significant MTAs, with minor allele frequency MAF ≥ 5%, were identified using the mixed linear models (MLM) taking into consideration both kinship (K) and population structure (Q) matrices as covariates. The K matrix was generated within TASSEL using 5176 DArT markers. To eliminate linear dependence between columns of the Q matrix, the last column was removed prior before running the GWAS. Manhattan plot was produced using ‘CMplot’ package; Marker-trait associations were considered significant at the threshold of a positive false discovery rate (FDR) less than 5%.
Putative candidate genes
Linking the significant markers with putative candidate genes was processed using a position-dependent strategy. The genomic region upstream and downstream (± 500 Kb) of the identified polymorphic sites was scanned to identify genes that are in the physical proximity to the reference genome IWGSC v1.0.
Only genes in the same genetic position were considered. The candidate genes were obtained using variant effect predictor at the EnsemblPlants website (https://plants.ensembl.org/Triticum_aestivum/Tools/VEP). For each wheat gene retrieved, the coding gene ID was subjected to blasting against the Panther (http://www.pantherdb.org/) and the Uniprot databases (www.uniprot.org). For further functional annotation on a genome-wide scale, we used the functions GO (gene ontology) and pathway. The list of genomic sequences harboring the polymorphic sites to allow follow-up marker development studies was generated.
Results
Phenotypic, genotypic, and environment variances and heritability
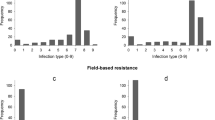
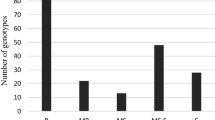
Phenotypic variation was observed for the infection types and level of YR severity for the 426 genotypes scored at Merchouch, during the two cropping seasons (Fig. 1).
Figure 1 shows the different types of genotype reactions to yellow rust on the x-axis, and the number of genotypes that share the same reaction on the y-axis. The final score ranged from 5 R (resistant) to 70 S (highly susceptible). Based on the coefficient of infection, the different rusts severity classes were classified. Out of 426 genotypes, 51.17% genotypes were resistant (exhibited resistance reaction response CI = 0 to 20); 25.82% genotypes were moderately resistant (CI = 20 to 30), 9.62% were moderately susceptible (CI = 30 to 40); 10.09% were moderately susceptible to susceptible (CI = 40 to 60) and 3.29% genotypes were susceptible to Puccinia striiformis f. sp. tritici (CI = 60 to 100) (Fig. 1).
Important difference in grain yield performance among the tested genotypes was observed, the average yield ranged from 9.51 and 3.50 t/ha. Among the genotypes in the resistant group, 20 elite lines have combined resistance to yellow rust, as well as a good yield potential (Table 1). These lines have been distributed to the national partners as part of the international nurseries for release or further use as parents.
The summary statics for yellow rust response is shown in Table 2. The analysis of variance for CI scores showed significance of the two years, and between the genotypes, but no genotypes × year interaction was observed. Hight heritability (H2) (0.887) was observed for the combined CI scores.
Population structure and linkage disequilibrium
To construct the population structure two hundred and seventy-two non-redundant markers across the entire genome were used. We estimated ancestry coefficients for each genotype and we computed the cross-entropy criterion. The minimum value was obtained when k = 10 (Fig. 2). The population structure of the 426 bread wheat genotypes showed stratification represented by 10 different colors. Genotypes with similar pedigree were mostly grouped in the same sub-group, as indicated in the heatmap of the relationship and ancestry matrices in Fig. 3.
For LD, most approaches are based on the coefficient of correlation (r2) calculated between marker pairs after giving numerical values to allele states. The scatter plots of LD (r2) as a function of the inter-marker distance for every genome indicated a clear LD decay for each genome (Fig. 4). LDs with mean r2 > 0.3 extended to distances up to 2.6 Mbp, 2.9 Mbp and 8 Mbp for A, B and D genomes, respectively.
Plot of pair-wise single-nucleotide polymorphism LD R2 values as a function of inter-marker map distance (Mbp). The blue curve represents the model fit to LD decay. The red line represents the extended to distances up to 2.6 Mbp, 2.9 Mbp and 8 Mbp for A, B and D genomes, respectively. for the quantitative trait loci regions in which LD R2 = 0.3 beyond which LD is likely due to linkage
GWAS for stripe rust resistance using DArT markers
Out of the 13,698 DArT markers obtained from genotyping, 5176 polymorphic DArT markers were used to run the GWAS using Q + K: MLM method. Genome B recorded the highest number of polymorphic markers with 1914 markers followed by D and A genomes with 1745 and 1325 markers, respectively. While, 164 markers had unknown positions. Chromosome 7D contained the largest number of markers (349), whereas chromosome 4D showed the fewest number (145).
A total of 19 significant marker trait associations (MTAs), representing 10 chromosomes regions were identified to be significantly associated (p < 0.001) with stripe rust resistance at adult plant stage (Table 3); These significant MTAs were located on chromosomes 2A, 7A, 1B, 2B, 3B, 5B, 1D, 2D, 3D, and 6D (Fig. 5).
The percentage of phenotypic variation explained by the significant markers ranged from 5.1% to 9.3%. The marker 583,038 in 5B exhibited the highest level of contribution to YR resistance (9.3%) with LOD of 5.326. Out of the significantly associated markers, only 3 passed a threshold of FDR ≤ 0.05. The marker 412,394 in chromosome 2B was mapped within the genomic regions of previously identified QTL (QYrlu.cau-2BS1 Luke and QYrid.ui-2B.1_IDO444) while the two other markers 542,318 (1B) and 583,038 (5B) were not overlapping with any of the previously reported genes/QTLs. Hence, these two markers most likely tag new Pst resistance locus.
Candidate genes and function annotations
The genetic frame of the significant markers was constructed using BLAST analysis of the International Wheat Genome Sequencing Consortium (IWGSC) RefSeq v1.0 genome. Only seven of the nineteen discovered markers contain related proteins, and only three of them passed the criteria (FDR p ≤ 0.05), as shown in Table 3.
An underlying breakthrough is to think about what would be the foremost effective approach to acknowledge stripe rust resistance within the germplasm, that would improve practical utilization in wheat breeding programs. Standard bi-parental crosses have been started using some of the potentially diverse germplasm supported diverging disease reactions. However, the development of cost-effective, high-throughput marker systems such as DArT, SNP and QTL mapping efforts in individual bi-parental populations will not expose, in the most efficient way, the diverse alleles present in large germplasm collections and their chromosomal locations (Roy et al. 2010). The GWAS relies frequently on the historical pattern of recombination that has been employed using different genetic backgrounds such as the elite germplasm used for this study (Vinod 2011). Field screening of the ICARDA elite panel confirmed that the majority of genotypes possessed adult resistance genes (APR). Within the present study, we found that from the germplasm screened at Merchouch station, 77% of the genotypes were resistant to moderately resistant. The results of GWAS might be considerably influenced by population structure (Kang et al. 2008). Therefore, in our study, most of the genotypes with similar pedigrees were clustered within the same group. Although 10 sub-populations were obtained using DArT markers.
The effectiveness of whole-genome association studies, for rust resistance and other traits, depends on the decay value. LD with mean r2 > 0.3 extended to distances up to 2.6 Mbp, 2.9 Mbp and 8 Mbp for A, B and D genomes, respectively (Fig. 4). DArT markers showed a higher percentage of significant pairs in the intrachromosomal LDs. These discrepancies may be assigned to the multi-loci nature of some DArT markers, which resulted in significant LD between markers located on different chromosomes or between distant markers on the same chromosome.
Discussion
The recent outbreak of stripe rust in many wheat-producing regions in East and North Africa, the Middle East, and Central West Asia poses a severe danger to wheat production and economic resources, as well as world food security. The most viable alternative to manage the permanent risk of yellow rust disease is within the development of resistant wheat cultivars. The application of modern tools for yellow rust resistance breeding and their advantages over conventional breeding methods helps in developing new cultivars with long lasting resistance (Buerstmayr et al. 2014).
Many territories in Central West Asia and North Africa (CWANA) and Sub-Saharan Africa (SSA) have previously reported severe yield losses ranging from 10 to 80% as a result of the breakdown of major gene resistance in some commonly planted cultivars due to the development of novel pathotypes (Solh et al. 2012; Zegeye et al. 2014). As a result, future outbreaks of yellow rust represent a persistent danger to yield stability. For example, the breakdown in resistance by Yr27 prompted Ethiopian farmers to spend over 3.2 million dollars on fungicides in 2010 to protect their production (Ali et al. 2017). In 2017, serious epidemics were observed in major wheat-growing areas in the CWANA region, including Algeria, Morocco, Syria, and Turkey (Morgounov et al. 2012).
In Morocco, during 2014/2015, two yellow rust races were registered, the Warrior (PstS7) race was confirmed in 2013 in Morocco and continue to be spread broadly in the Northern African countries, Europe, and East Asia (Milus et al. 2009; EL-Sabagh et al. 2017). The PstS10 known as Warrior (-) (virulence pattern: [1,2,3,4,-,6,7,-,9,-,-,17,-,25,-,32,Sp,AvS,-]) was also reported in Algeria and Spain during 2014 and possibly in Morocco as well, but no available data to verify (Hovmøller et al. 2018; El Hanafi et al. 2021).
ICARDA has committed 40 years working with communities and national research centers in the CWANA region to establish genotypes with novel sources of YR resistance in order to stay ahead of the pathogens evolutionary potential outbreak. Therefore, the possibility of novel pathotypes overcoming resistance will be followed and reduced by combining both minor and major resistance genes (Pedersen and Leath 1988).
An underlying breakthrough is to think about what the foremost effective approach would be to acknowledge stripe rust resistance within the germplasm, that would improve practical utilization in wheat breeding programs. The continuous field screening of the genetic material is crucial to track and identify the resources of resistance that would be deployed in modern plant breeding programs. In the present study, we could show that 77% of the 426 genotypes used were resistant to moderately resistant and most likely were distributed as international nurseries and used already in previous crossing plans. Hence, we applied GWAS using 5549 polymorphic DArT markers to detect the associated markers with the resistance to yellow rust. A total of 19 MTAs at different wheat chromosomes were detected significatively associated with stripe rust resistance at LOD ≤ 3. However, only 3 markers (542,318, 412,394 and 583,038) were significant at experiment-wise FDR ≤ 0.05 on chromosomes 1B, 2B and 5B, respectively. A comparable study conducted in the same region of Merchouch in 2014/2015 discovered 23 markers on chromosomes 2A, 2B, 2D, and 7B that were highly related with adult plant resistance at FDR ≤ 0.05 (El Hanafi et al. 2021). Unfortunately, none of those associated markers was in common or overlapped with the genomic region of the markers found in the current study.
In order to determine the genes/QTLs associated with stripe rust resistance previously reported, we have compared the physical location of the three markers found at FDR ≤ 0.05 by BLAST analysis of the International Wheat Genome Sequencing (IWGSC RefSeq v2.1 genome). Numerous Yr genes and QTL have been mapped on chromosome 1B, including Yr9, Yr10, Yr15, Yr24/Yr26/YrCh42, Yr64, Yr65, YrTr1, YrAlp, YrH52, YrH122, YrL693, YrC142, YrMY41, QYr.cau-1BS, QYrco.wpg-1BS.1 and QYrco.wpg-1BS.2 (Feng et al. 2018). However, none of the genes/QTLs are within the genomic region of the marker 542,318 detected in this study, thus, likely represents novel resistance loci for stripe rust. In fact, QYr.cim-1BSb reported to have a small effect QTL by wPT-8168 and wPT-6240 DArt markers in cultivar Pastor close to the genomic region of the marker 542,318 (Maccaferri et al. 2015). BLAST analysis of the marker 583,038 identified one gene TraesCS5B02G374300 that encodes for Glutaredoxin protein having role as detoxification of xenobiotics and heavy metals, signaling in plant-pathogen interactions as well as plant development processes and cell cycle control (Ziemann et al. 2008).
On chromosome 2B, 112 genes were mapped so far. Among the previously published QTL, QYrlu.cau‐2BS1_Luke and QYrid.ui‐2B.1_IDO444 conferring large effects were located within the flanking region of the marker 412,394 (64.09 Mbp) (Chen et al. 2012; Mallard et al. 2005). QYrlu.cau‐2BS1 Luke is mapped in 36.4 Mbp and 100.8 Mbp covering almost 64.4 Mbp. While, QYrid.ui‐2B.1_IDO444 appeared in the genomic region between 49.8 Mbp and 72.6 Mbp according to the annotation of Chinese Spring reference sequence. BLAST analysis and anchoring the flanking marker 412,394 on the reference genomes allowed the identification of the candidate gene TraesCS2B02G103200 reported with unknown function.
In chromosome 5B, the marker 583,038 at 551 Mbp reported to be associated to yellow rust resistance at adult plant stage having the highest LOD = 5.33. This marker was far away from many QTL (QYr.caas-5BL.2_Libellula, QYr.tem5B.2_Flinor, and QYr.inra-5BL.2_CampRemy) for about 44 Mbp, hence tagging novel genomic region. BLAST analysis of the marker 583,038 identified functional protein TraesCS5B02G374300 with unknown function.
The significantly associated markers with yellow rust resistance in this study should be validated using other elite set of wheat genotypes and could be used for marker assisted selection in wheat breeding programs to pyramid resistance genes and develop varieties with durable resistance.
References
Ali S, Rodriguez-Algaba J, Thach T, Sørensen CK, Hansen JG, Lassen P, Hovmøller MS (2017) Yellow rust epidemics worldwide were caused by pathogen races from divergent genetic lineages. Front Plant Sci 8:1057. https://doi.org/10.3389/fpls.2017.01057
Allaire J (2012) RStudio: integrated development environment for R. Boston, MA. 770(394):165–171
Ballesta S, Shi W, Conen KE, Padoa-Schioppa C (2020) Values encoded in orbitofrontal cortex are causally related to economic choices. Nature 588(7838):450–453
Beddow JM, Pardey PG, Chai Y, Hurley TM, Kriticos DJ, Braun HJ, Yonow T (2015) Research investment implications of shifts in the global geography of wheat stripe rust. Nature Plants 1(10):1-5
Buerstmayr M, Matiasch L, Mascher F et al (2014) Mapping of quantitative adult plant field resistance to leaf rust and stripe rust in two European winter wheat populations reveals co-location of three QTL conferring resistance to both rust pathogens. Theo Appl Genet 127(9):2011–2028
Caldwell RM (1968) Breeding for general and/or specific plant disease resistance. In Proceedings of the Third International Wheat Genetics Symposium, Canberra, Australia. Canberra: Australian Academy of Sciences 236–272
Chen J, Chu C, Souza EJ et al (2012) Genome-wide identification of QTL conferring high-temperature adult-plant (HTAP) resistance to stripe rust (Puccinia striiformis f. sp. tritici) in wheat. Mole Breed 29(3):791–800
Chen W, Wellings C, Chen X et al (2014) Wheat stripe (yellow) rust caused by P uccinia striiformis f. sp. tritici. Mole Plant Pathol 15(5):433–446
Chen X (2020) Pathogens which threaten food security: Puccinia striiformis, the wheat stripe rust pathogen. Food Sec 12(2):239–251
Dong Z, Hegarty JM, Zhang J et al (2017) Validation and characterization of a QTL for adult plant resistance to stripe rust on wheat chromosome arm 6BS (Yr78). Theo Appl Genet 130(10):2127–2137
El-Sabagh A, Barutcular C, Islam MS (2017) Relationships between stomatal conductance and yield under deficit irrigation in maize (Zea mays L.). J Exp Biol Agric Sci 5(1):14–21
El Hanafi S, Backhaus, A, Bendaou N et al (2021) Genome-wide association study for adult plant resistance to yellow rust in spring bread wheat (Triticum aestivum L.). Euphytica 217(5):1-14
FAO Crop Prospects and Food Situation (2018) Food and Agricultural Organization of the United Nations (FAO)
Feng J, Wang M, See DR et al (2018) Characterization of novel gene Yr79 and four additional quantitative trait loci for all-stage and high-temperature adult-plant resistance to stripe rust in spring wheat PI 182103. Phytopathol 108(6):737–747
Flor HH (1956) The complementary genic systems in flax and flax rust. In Adv Genet Academic Press 29–54
Frichot E, François O (2015) LEA: An R package for landscape and ecological association studies. Methods in Ecology and Evolution (MEE) 6(8):925–929
Hill WG, Weir BS (1988) Variances and covariances of squared linkage disequilibria in finite populations. Theor Popul Biol 33(1):54–78
Hovmøller MS, Rodriguez-Algaba J, Thach T et al (2018) Report for Puccinia striiformis race analyses and molecular genotyping 2017, Global Rust Reference Center (GRRC), Aarhus University, Flakkebjerg, DK-4200 Slagelse Denmark. https://doi.org/10.1094/phyto-85-567
https://www.historique-meteo.net. Accessed 14 Mar 2022
Johnson R (1988) Durable resistance to yellow (stripe) rust in wheat and its implications in plant breeding. In Breeding Strategies for Resistance to the Rusts of Wheat, El Batan, Mexico (Mexico), 29 Jun-1 Jul 1987. CIMMYT
Kang HM, Zaitlen NA, Wade CM (2008) Efficient control of population structure in model organism association mapping. Genetics 178(3):1709–1723
Kankwatsa P, Singh D, Thomson PC et al (2017) Characterization and genome-wide association mapping of resistance to leaf rust, stem rust and stripe rust in a geographically diverse collection of spring wheat landraces. Mole Breed 37(9):1–24
Knapp SJ, Stroup WW, Ross WM (1985) Exact confidence intervals for heritability on a progeny mean basis 1. Crop Sci 25(1):192–194
Kushalappa AC, Yogendra KN, Karre S (2016) Plant innate immune response: qualitative and quantitative resistance. Crit Rev Plant Sci 35(1):38–55
Lin F, Chen XM (2007) Genetics and molecular mapping of genes for race-specific all-stage resistance and non-race-specific high-temperature adult-plant resistance to stripe rust in spring wheat cultivar Alpowa. Theo Appl Genet 114(7):1277–1287
Maccaferri M, Zhang J, Bulli P et al (2015). A genome-wide association study of resistance to stripe rust (Puccinia striiformis f. sp. tritici) in a worldwide collection of hexaploid spring wheat (Triticum aestivum L.). G3: Genes Genom Genet 5(3):449–465
Mallard S, Gaudet D, Aldeia A, Abelard C, Besnard AL, Sourdille P, Dedryver F (2005) Genetic analysis of durable resistance to yellow rust in bread wheat. Theo Appl Genet 110(8):1401–1409
Milus EA, Kristensen K, Hovmøller MS (2009) Evidence for increased aggressiveness in a recent widespread strain of Puccinia striiformis f. sp. tritici causing stripe rust of wheat. Phytopathol 99(1):89–94
Morgounov A, Tufan HA, Sharma R et al (2012) Global incidence of wheat rusts and powdery mildew during 1969–2010 and durability of resistance of winter wheat variety Bezostaya 1. Eur J Plant Pathol 132(3):323–340. https://doi.org/10.1007/s10658-011-9879-y
Niks RE, Alemu SK, Marcel TC et al (2015) Mapping genes in barley for resistance to Puccinia coronata from couch grass and to P. striiformis from brome, wheat and barley. Euphytica 206(2):87–499
Ogbonnaya FC, Seah S, Delibes A et al (2001) Molecular-genetic characterisation of a new nematode resistance gene in wheat. Theor Appl Genet 102(4):623–629. https://doi.org/10.1007/s001220051689
Ohm HW (1976) Three components of slow leaf-rusting at different growth stages in wheat.
Parlevliet JE (1979) Components of resistance that reduce the rate of epidemic development. Ann Rev Phytopathol 17(1):203–222
Pathan AK, Park RF (2006) Evaluation of seedling and adult plant resistance to leaf rust in European wheat cultivars. Euphytica 149(3):327–342
Pedersen WL, Leath S (1988) Pyramiding major genes for resistance to maintain residual effects. Annual Rev Phytopathol 26(1):369–378
Peterson RF, Campbell AB, Hannah AE (1948) A diagrammatic scale for estimating rust intensity on leaves and stems of cereals. Canadian J Res 26(5):496–500. https://doi.org/10.1139/cjr48c-033
Puntel LA, Sawyer JE, Barker DW (2016) Modeling long-term corn yield response to nitrogen rate and crop rotation. Front Plant Sci 7:1630
Roelfs AP (1992) Rust diseases of wheat: concepts and methods of disease management. Cimmyt
Roy JK, Smith KP, Muehlbauer GJ et al (2010). Association mapping of spot blotch resistance in wild barley. Mole Breed 26(2):243–256
Solh M, Nazari K, Tadesse W, Wellings CR (2012) The growing threat of stripe rust worldwide. In Borlaug Global Rust Initiative (BGRI) conference, Beijing, China (pp. 1–4)
Tadesse W, Bishaw Z, Assefa S (2019) Wheat production and breeding in Sub-Saharan Africa: Challenges and opportunities in the face of climate change. Int J Clim Change Strateg Manag https://doi.org/10.1108/IJCCSM-02-2018-0015.
Tadesse W, Ogbonnaya FC, Jighly A et al (2015) Genome-wide association mapping of yield and grain quality traits in winter wheat genotypes. PLoS ONE 10(10):e0141339
Vanraden PM (2007) Efficient estimation of breeding values from dense genomic data. J Dairy Sci 90(1):374–375
Vinod KK (2011) Association mapping in crop plants. Bioinform Tools Genom Res
Wang J, Zhang Z (2021) GAPIT Version 3: boosting power and accuracy for genomic association and prediction. Genom Proteom Bioinform
Wang SB, Feng JY, Ren WL et al (2016) Improving power and accuracy of genome-wide association studies via a multi-locus mixed linear model methodology. Sci Rep 6(1):1–10. https://doi.org/10.1038/srep19444
Yang M, Li G, Wan H et al (2019) Identification of QTLs for stripe rust resistance in a recombinant inbred line population. Int Journal Molecular Sciences 20(14):3410. https://doi.org/10.3390/ijms20143410
Zegeye H, Rasheed A, Makdis F et al (2014) Genome-wide association mapping for seedling and adult plant resistance to stripe rust in synthetic hexaploid wheat. PloS One 9(8):e105593
Ziemann M, Zachgo S, Bhave M (2008) The glutaredoxin gene family in wheat functions beyond redox homeostasis regulation
Acknowledgements
We would like to thank the International Center for Agricultural Research in the Dry Areas (ICARDA) for its support and funding this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
El Messoadi, K., El Hanafi, S., Gataa, Z.E. et al. Genome wide association study for stripe rust resistance in spring bread wheat (Triticum aestivum L.). J Plant Pathol 104, 1049–1059 (2022). https://doi.org/10.1007/s42161-022-01132-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42161-022-01132-z