Abstract
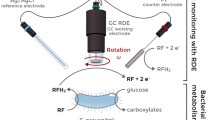
Impedance measurements using graphite electrodes were used to detect the increase in culture medium conductivity due to bacteria growth in real time along with simultaneous voltammetric monitoring of pyocyanin concentration. Electrochemical methods were compared to conventional continuous monitoring of bacterial cultures using turbidity measurement at an optical density of 600 nm (OD600). A practical osmotic system was further designed for concentrating bacterial cultures during growth to enable earlier detection using the electrochemical methods. Bacterial cultures, starting from an initial culture density of 1 × 108 cells/mL, were grown inside a sealed cellulose ester dialysis membrane, while polyethylene glycol in LB medium was used as the draw solution outside the membrane to gradually concentrate the growing cultures. 0.7-mm-diameter graphite for mechanical pencils was utilized as working and counter electrodes with a platinum wire reference electrode for electrochemical measurements with and without the osmotic system. In the absence of forward osmosis, impedance measurements detected culture growth ~ 1 h faster than conventional OD600. Integrating osmosis showed a twofold decrease in the time to detect pyocyanin production as an indicator for bacterial growth. For impedance monitoring, removal of water by osmosis was conflated with the impedance decrease due to cell growth; however, the results show a promising ability to detect bacteria growth via an observed shift in osmotic impedance profile when bacteria are present in a sample. By monitoring the deviation in the impedance profile, a threefold improvement in detection time was achieved when compared to conventional OD600 measurements.







Similar content being viewed by others
References
Heo J, Hua SZ. An overview of recent strategies in pathogen sensing. Sensors (Switzerland). 2009;9:4483–502.
Wang H, Bedard E, Prevost M, et al. Methodological approaches for monitoring opportunistic pathogens in premise plumbing: a review. Water Res. 2017;117:68–86.
Bereket W, Hemalatha K, Getenet B, et al. Update on bacterial nosocomial infections. Eur Rev Med Pharmacol Sci. 2012;16:1039–44.
Ventola CL. The antibiotic resistance crisis: part 1: causes and threats. Pharm Ther J. 2015;40(4):277.
Caliendo AM, Gilbert DN, Ginocchio CC, et al. Better tests, better care: improved diagnostics for infectious diseases. Clin Infect Dis. 2013;57:S139–70. https://doi.org/10.1093/cid/cit578.
Frieden T. Antibiotic resistance threats in the United States, 2013. Centres for Disease Control and Prevention; 2013. https://www.cdc.gov/drugresistance/pdf/ar-threats-2013-508.pdf.
Karlowsky JA, Draghi DC, Jones ME, et al. Surveillance for antimicrobial susceptibility among clinical isolates of Pseudomonas aeruginosa and Acinetobacter baumannii from hospitalized patients in the United States, 1998 to 2001. Antimicrob Agents Chemother. 2003;47:1681–8. https://doi.org/10.1128/AAC.47.5.1681-1688.2003.
CDC. Gram-negative bacteria infections in healthcare settings. Centres for Disease Control and Prevention; 2011. https://www.cdc.gov/HAI/organisms/gran-negative-bacteria.html. Accessed 14 Dec 2018.
Foweraker JE, Laughton CR, Brown DFJ, Bilton D. Phenotypic variability of Pseudomonas aeruginosa in sputa from patients with acute infective exacerbation of cystic fibrosis and its impact on the validity of antimicrobial susceptibility testing. J Antimicrob Chemother. 2005;55:921–7. https://doi.org/10.1093/jac/dki146.
Burns JL, Saiman L, Whittier S, et al. Comparison of agar diffusion methodologies for antimicrobial susceptibility testing of Pseudomonas aeruginosa isolates from cystic fibrosis patients. J Clin Microbiol. 2000;38:1818–22.
Wanger, A. Antibiotic susceptibility testing. In: Practical handbook of microbiology, 3rd ed. 2015.
Kerremans JJ, Verboom P, Stijnen T, et al. Rapid identification and antimicrobial susceptibility testing reduce antibiotic use and accelerate pathogen-directed antibiotic use. J Antimicrob Chemother. 2008;61:428–35. https://doi.org/10.1093/jac/dkm497.
Kassim A, Omuse G, Premji Z, Revathi G. Comparison of Clinical Laboratory Standards Institute and European Committee on Antimicrobial Susceptibility Testing guidelines for the interpretation of antibiotic susceptibility at a University Teaching Hospital in Nairobi, Kenya: a cross-sectional stud. Ann Clin Microbiol Antimicrob. 2016;11:15–21. https://doi.org/10.1186/s12941-016-0135-3.
Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing, 27th ed. CLSI supplement M100. Wayne: Clinical and Laboratory Standards Institute; 2017.
European Committee on Antimicrobial. The European Committee on Antimicrobial Susceptibility Testing. Breakpoint tables for interpretation of MICs and zone diameters/EUCAST; 2017.
Lambert RJW, Pearson J. Susceptibility testing: accurate and reproducible minimum inhibitory concentration (MIC) and non-inhibitory concentration (NIC) values. J Appl Microbiol. 2000;88:784–90. https://doi.org/10.1046/j.1365-2672.2000.01017.x.
Jorgensen JH, Ferraro MJ. Antimicrobial susceptibility testing: a review of general principles and contemporary practices. Clin Infect Dis. 2009;49:1749–55. https://doi.org/10.1086/647952.
Yang L, Bashir R. Electrical/electrochemical impedance for rapid detection of foodborne pathogenic bacteria. Biotechnol Adv. 2008;26:135–50. https://doi.org/10.1016/j.biotechadv.2007.10.003.
Oziat J, Elsen S, Owens RM, et al. Electrochemistry provides a simple way to monitor Pseudomonas aeruginosa metabolites. In: Proceedings of the annual international conference of the IEEE engineering in medicine and biology society, EMBS; 2015.
Santiveri CR, Sismaet HJ, Kimani M, Goluch ED. Electrochemical detection of Pseudomonas aeruginosa in polymicrobial environments. ChemistrySelect. 2018;3:2926–30. https://doi.org/10.1002/slct.201800569.
Buzid A, Shang F, Reen FJ, et al. Molecular signature of Pseudomonas aeruginosa with simultaneous nanomolar detection of quorum sensing signaling molecules at a boron-doped diamond electrode. Sci Rep. 2016;6:30001. https://doi.org/10.1038/srep30001.
Wawerla M, Stolle A, Schalch B, Eisgruber H. Impedance microbiology: applications in food hygiene. J Food Prot. 1999;62:1488–96. https://doi.org/10.4315/0362-028X-62.12.1488.
Cheng X, Liu YS, Irimia D, et al. Cell detection and counting through cell lysate impedance spectroscopy in microfluidic devices. Lab Chip. 2007;7:746–55. https://doi.org/10.1039/b705082h.
Silley P, Forsythie S. Impedance microbiology-a rapid change for microbiologists. J Appl Bacteriol. 1996;80:233–43. https://doi.org/10.1111/j.1365-2672.1996.tb03215.x.
Yang L, Li Y. Detection of viable Salmonella using microelectrode-based capacitance measurement coupled with immunomagnetic separation. J Microbiol Methods. 2006;64:9–16. https://doi.org/10.1016/j.mimet.2005.04.022.
Seviour T, Doyle LE, Lauw SJL, et al. Voltammetric profiling of redox-active metabolites expressed by Pseudomonas aeruginosa for diagnostic purposes. Chem Commun. 2015;51:3789–92. https://doi.org/10.1039/c4cc08590f.
Webster TA, Sismaet HJ, Conte JL, et al. Electrochemical detection of Pseudomonas aeruginosa in human fluid samples via pyocyanin. Biosens Bioelectron. 2014;60:265–70. https://doi.org/10.1016/j.bios.2014.04.028.
Sismaet HJ, Goluch ED. Electrochemical probes of microbial community behavior. Annu Rev Anal Chem. 2018;11:441–61. https://doi.org/10.1146/annurev-anchem-061417-125627.
Beyenal H, Chen SN, Lewandowski Z. The double substrate growth kinetics of Pseudomonas aeruginosa. Enzyme Microb Technol. 2003;32:92–8. https://doi.org/10.1016/S0141-0229(02)00246-6.
Acknowledgements
This work was supported in part by award #1740961 from the National Science Foundation.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Kimani, M.K., Mwangi, J. & Goluch, E.D. Bacterial Sample Concentration and Culture Monitoring Using a PEG-Based Osmotic System with Inline Impedance and Voltammetry Measurements. J. Anal. Test. 3, 166–174 (2019). https://doi.org/10.1007/s41664-019-00096-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41664-019-00096-x