Abstract
The net form net blotch (NFNB) is a significant disease of barley. Its causal agent, Pyrenophora teres f. teres (PTT), has an important economic impact on yield and grain quality globally. However, the molecular interaction between PTT and barley is not fully understood. The plant–pathogen encounter comprises the secretion of diverse molecules involved in plant defence, including pathogenicity-related proteins, and fungal attack, such as proteinaceous toxins called effectors. The forefront of the molecular crosstalk between plant and fungus is the space between plant cells or apoplast. To explore the suitability of studying apoplastic proteins to assist understanding the host–pathogen interaction, a mass spectrometry-based proteomics technique was used to profile apoplastic protein differences in control and NFNB-infected leaves in a susceptible cultivar. The analysis revealed 1130 barley proteins, of which 140 were found to be significantly differentially expressed. This paper presents an overview of the major protein changes induced in the barley apoplast and discusses the involvement of individual proteins in defence and disease development. Our results suggest that the fungus may be hijacking defence signalling pathways. This investigation provides the first in vivo proteomics data for a NFNB–barley interaction, setting a background for further studies.
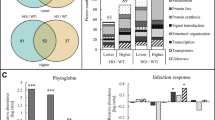
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Net blotch of barley is a significant disease caused by two closely related fungal pathogens: Pyrenophora teres f. teres (PTT) and Pyrenophora teres f. maculata (PTM). The causal agent of the net form net blotch (NFNB) is PTT, while PTM causes spot form net blotch (Smedegård-Petersen 1971). NFNB has a longer host association with barley and is genetically diverse, with numerous pathotypes and correspondingly complex genetic interactions with barley (Liu et al. 2010; Ellwood et al. 2012, 2019). PTT is a hemibiotrophic fungus, meaning it switches from biotrophy to necrotrophy, polar opposites of pathogenic lifestyles. Biotrophic fungi typically colonise healthy plant tissue by evading or suppressing the host defence responses and acquiring nutrients from living host cells, while necrotrophic fungi gain nutrition by killing the host tissue (Vleeshouwers and Oliver 2015). Biotrophic and necrotrophic fungi elicit different responses in the host plant. In a susceptible interaction, PTT induces pathogenesis-related (PR) proteins associated with the activation of the plant salicylic acid (SA) pathway (Al-Daoude et al. 2018) which is a defence pathway adapted against biotrophic fungi (Bhadauria et al. 2010). This pathway can induce a strong defence response leading to host cell death known as hypersensitive response (HR). However, PTT’s early biotrophic stage is relatively short, spending time as a necrotroph the majority of its infectious cycle (Lightfoot and Able 2010). As necrotrophic fungi rely on host cell death, they may hijack the HR pathway inducing cell ‘suicide’ mechanisms, allowing for a successful susceptible interaction (Hammond-Kosack and Rudd 2008). Both ethylene (ET) and jasmonic acid (JA) pathways are typically induced in response to necrotrophic fungi and do not result in HR, but are involved in the upregulation of alternative defence-related genes (Glazebrook 2005). These interactions take place via a complex interchange of signals, pathogenicity factors and defence response proteins primarily in the apoplast, the space between the host cells where the fungal infection develops (Mott et al. 2014). The discovery and classification of the key protein players in a pathosystem are of high importance, not only for the understanding of the plant–pathogen interactions at a molecular level, but also to the plant breeding industry as it could help to develop more resistant crops, via such techniques as effector-assisted breeding (Vleeshouwers and Oliver 2015).
Among inducible plant responses in response to fungal disease are pathogenesis-related (PR) proteins (Jin Kim and Kook Hwang 2000), covering a diverse range of structurally and functionally different proteins (Zhang et al. 2018). PR proteins are typically localised extracellularly in the apoplastic space or within the vacuole, the former being the main site of accumulation (Gorjanović 2009). Since their first discovery in 1970, through investigations into hypersensitive response reactions against tobacco mosaic virus, many PR proteins have been identified which are now categorised into 17 families (Van Loon and Van Kammen 1970). Only 13 of these families have been reported as being expressed in barley (Table 1). PR proteins are typically expressed during either compatible or incompatible interactions with an invading pathogen; however, some are also constitutively expressed during plant development and senescence (Hanfrey et al. 1996; Liljeroth et al. 2005). Currently, there is limited knowledge of the molecular mechanisms of many PR proteins; however, they are starting to play a part in the future of plant breeding efforts as targets for genetically modified crop development due to their central roles in defence against agricultural fungal pathogens (Mills and Tatham 2003). Furthermore, PR proteins are of interest as food and seed preservatives due to their antifungal properties (Dempsey et al. 1998).
To date, no proteomic study has investigated the interaction between PTT and barley in vivo. The aim of this study was to explore the proteins present in the plant apoplast, as well as the suitability of this approach in providing a better understanding of the plant responses to the pathogen. To achieve this, cellular wash fluid (CWF) from healthy and infected barley leaves was collected applying the broadly used vacuum infiltration–centrifugation method (Witzel et al. 2011). The samples were then analysed and compared by sequential window acquisition of all theoretical fragment ion spectra mass spectrometry (SWATH-MS), which is a data-independent acquisition technique allowing label-free proteomics while providing reproducible and accurate protein quantitation (Krasny et al. 2018). The study explores the strengths and limitations of this approach and discusses the significance of different PR families identified in this study during the development of NFNB disease, together with the current knowledge of the biological functions of these proteins.
Methodology
Fungal and plant material
PTT isolate NB29 (collected from Wongan Hills, WA, in 1985) was used in this study as it exhibits good sporulation under laboratory conditions and is virulent against a range of barley cultivars. The isolate was grown and maintained on V8 juice potato dextrose agar (V8-PDA; 150 ml l−1 V8 (Campbell’s, Chicago, USA), 10 g l−1 potato dextrose agar (Thermo Fisher Scientific, Massachusetts, USA), 3 g l−1 CaCO3 and 10 g l−1 agar).
The barley cultivar Baudin was used as a representative of a modern commercial line which is susceptible to NB29. Plants were grown in 36-cell propagation trays containing four seeds per cell. Trays were filled with vermiculite and fertilised with one teaspoon of Thrive all-purpose fertiliser (Yates, Auckland, New Zealand) before being placed under florescent lighting at 18 °C with a 12-h photoperiod.
Inoculum preparation
In order to induce sporulation of PTT, V8-PDA plates were inoculated with an agar plug of NB29 and incubated for 7 days at room temperature after which the plates were exposed to black light for 18 h at room temperature, followed by 24 h in the dark at 15 °C (Speakman and Pommer 1986). Spore collection was performed by adding 5 ml of water with 0.02% Tween-20 to the plates and applying a paintbrush to the whole plate to dislodge the spores. The liquid containing the spores was collected and the concentration adjusted to 1000 spores ml−1 after being measured with a haemocytometer. A control solution was prepared similarly using non-inoculated V8-PDA plates.
Plant infection
Baudin plants were grown to the third leaf stage (Zadok’s growth stage Z13.3/20) (Lancashire et al. 1991). Inoculation was performed using a method similar to that described by Kosiada (2008). Briefly, seedling trays were sprayed with 30 ml of either spore suspension or control solution using a spray bottle and allowed to dry for 30 min. Inoculated plants were then placed in a growth chamber under a 48-h period of darkness at 25 °C and > 90% humidity.
CWF collection
CWF containing the apoplastic fluid was obtained using the vacuum infiltration–centrifugation method described by Witzel et al. (2011). The second and third leaves were collected five days after infection, cut into 5–7 cm sections and the cut ends rinsed in 0.1 M sodium phosphate buffer (pH 6.5). In order to infiltrate the collected leaves, batches of around 25 leaves were loaded vertically into 50 ml centrifuge tubes and submerged in 0.1 M sodium phosphate buffer (pH 6.5). Next, the tubes were placed in a vacuum chamber for 10 min, followed by three mins at atmospheric pressure, and then another cycle of 10 min under vacuum (300 mBar) with three mins of incubation at atmospheric pressure.
To collect the CWF, infiltrated leaves were then gently dried with a paper towel to remove any liquid on the surface before placing them in 20 ml syringe barrels, which were then inserted in 50 ml centrifuge tubes similar to the method described by Agrawal et al. (2010). Tubes were centrifuged for 10 min at 1000g, and the CWF collected as recommended by Nouchi et al. (2012). A protease inhibitor (Pefabloc SC, Sigma-Aldrich) was then added to each sample to a final concentration of 0.25 mM. The protein concentration of the CWF was measured using an Invitrogen 2.0 Qubit Fluorometer and adjusted to 300 µg per sample to be submitted for SWATH-MS proteomic analysis. Each of the two variables (infected and control CWF samples) tested included four biological replicates.
Protein verification
Prior to sending the samples for SWATH-MS analysis, a tricine-based SDS-PAGE protein gel was prepared to test each sample for protein degradation and protein band differences. To do this, 1.5 ng of each protein sample was combined with 4 µl of Coomassie Blue loading dye (4 ml 4 × TrisCl/SDS pH 6.8, 4.8 ml glycerol, 1.6 g SDS, 0.6 g dithiothreitol and 4 mg Coomassie Blue G250 per 10 ml) and made up to 20 µl with MQ water. Each sample was then heated to 99 °C for three mins to allow the proteins to denature. 20 µl of each sample was loaded as well as 5 µl of Precision Plus, Protein Dual Xtra Standard ladder (Bio-Rad, California, USA) with a range of 250–2 kDa. The gel was then run for 1 h at 20 V, followed by 2 h at 120 V. The gel was then silver-stained using Invitrogen’s SilverQuest kit (Thermo Fisher Scientific, Massachusetts, USA) following the manufacturer’s protocol.
Proteomic analysis
Sample preparation
Proteomic analysis was carried out by the Australian Proteome Analysis Facility (APAF), Macquarie University of NSW, Australia. Samples were prepared as follows: 100 µg of each sample was diluted to 450 µl with 1% sodium deoxycholate. The samples then were reduced with 10 mM DTT for 30 min at 56 °C. This was followed by alkylation using 25 mM IAA at room temperature, for 30 min in the dark. Alkylation was quenched with 10 mM DTT for 15 min at room temperature. Trypsin (5 µg) was then used to digest each protein sample for 16 h, at 37 °C. TFA acid was then added (final concentration of 1%) to precipitate out sodium deoxycholate. The resulting peptides were recovered from the supernatant after centrifuging at 14,000g for 10 min. Peptide samples were concentrated and desalted using a solid phase extraction disk and washing with 0.2% TFA and then eluted with acetonitrile with 5% ammonium hydroxide. Peptides were then dried under vacuum.
Samples were reconstituted in loading buffer (25 µl of 2% acetonitrile, 97.9% water and 0.1% formic acid), vortexed briefly and sonicated for one min before centrifuging at 10,000g for 10 min.
Ion library generation through high pH (HpH) fractionation was done by pooling 15 µg of each respective biological replicate. The peptide mixture was fractionated by HpH reverse phase HPLC into 96 fractions, which were then consolidated into 17 fractions prior to LC-MS/MS analysis. Consolidated fractions were then vacuum dried.
Data acquisition
In order to perform information-dependent data acquisition (IDA), each HpH fraction sample was subjected to 1D-IDA nano-LC MS/MS analysis. During this, a time-of-flight mass spectrometry (TOF-MS) survey is acquired with the 20 most intense multiply charged ions being subjected to MS/MS analysis. MS/MS spectra were then accumulated for 100 ms using the rolling collision energy (mass range m/z 100–1800). In order to perform the data-independent analysis (SWATH), a TOF-MS survey scan was conducted (m/z 350–1500, 0.05 s) followed by 100 predefined m/z ranges, which were sequentially subjected to MS/MS analysis. Thirty milliseconds of MS/MS spectra data were accumulated in the mass range m/z 350–1500, with rolling collision energy optimised for low m/z in m/z window + 10%.
Data processing
The IDA data files were collectively searched with ProteinPilot (v5.0) (Sciex, Massachusetts, USA) using the ParagonTM algorithm. The UniProt database for barley plant proteins (Hordeum vulgare subsp. vulgare strain: cv. Morex, proteome ID: UP000011116) was used for searching the data. Due to the lack of any protein hits related to PTT in preliminary searches, only the barley proteome was used. A minimum of a 95% confidence for identification was used as well as unused score cut-off of 1.5.
Analysis of the ion library and SWATH data files was done in PeakView (v2.1). Retention time differences between SWATH and IDA analysis were aligned with the aid of six endogenous peptides. The top six most intense fragments of each peptide were extracted from the SWATH data sets, with any modified or shared peptides being excluded. A maximum of 100 peptides/protein with confidence greater than 99% and false discovery rate less than 1% were then used for quantification.
Identification of highly differentially expressed proteins
Additional filtering was used to identify the most differentially expressed proteins and improve peptide matching accuracy. Proteins were filtered to contain a minimum of two peptide hits, a P value of less than 0.05 and minimum fold change of 2.5. These proteins were then searched using the UniProt BLAST function with an unused cut-off score of 1e−05 for the most similar homology in order to match the identified uncharacterised proteins to known proteins. Proteins with alignments of less than 80% were assessed with caution and discussed accordingly.
Results
Plant infections and extraction of CWF
Following inoculation, PTT disease lesions showed progressive development in size indicating successful infection. Infected leaves were similar to non-inoculated control leaves until three days post-inoculation (dpi), when signs of necrosis started to appear. Necrotic lesions continued to develop, and chlorosis was visible after 4 dpi. Leaves for the vacuum infiltration were collected at five dpi to achieve a balance of adequate disease levels without excessive tissue damage. After extraction of the CWF, the protein concentration of each sample was measured. On average, the infected samples had a higher concentration compared to the uninfected (averaging 464 µg ml−1 and 612 µg ml−1, respectively, P = 0.02). The quality of the samples was then evaluated by SDS-PAGE gel electrophoresis which showed discrete bands present in each well, suggesting that the proteins in each sample were not degraded during the extraction process. Band intensity differences were visible between the CWF from infected plants and the uninfected controls (image not shown).
Proteomic data analysis
CWF samples were analysed by SWATH-MS to determine protein level differences between the infected and non-infected barley plants to identify induced host response proteins and proteins secreted by PTT into the barley apoplastic space. The analysis revealed 1130 proteins which were differentially expressed and linked to barley UniProt identifiers across all eight samples (supplementary data 1). No fungal proteins were identified, the reasons for which are suggested in the discussion. The results were then restricted to P values < 0.05 and a minimum fold change of ± 1.5. This filtering left 140 proteins, of which 52 were significantly upregulated and 88 significantly downregulated (Fig. 1a, supplementary data 2). However, the average fold increase in the upregulated proteins was approximately 11, while the change of those that were downregulated was approximately three (Fig. 1b).
Extracellular defence-related proteins are highly upregulated in CWF
The biological roles and cellular localisation of 20 most upregulated and the 20 most downregulated proteins were determined using UniProt BLASTP, their gene ontology (GO) annotations and their homology to known proteins, and/or found in previous reports. The majority of the 20 most upregulated proteins had extracellular localisation signals (Fig. 2a) and consisted of PR proteins and a single non-PR ripening protein (Fig. 3). The highest difference in expression levels was related to three PR-5 proteins, homologues of thaumatin-like proteins (TLP). These proteins showed strong homologies to TLP 5, 7 and 8 and were previously reported to be secreted into the apoplast (Wang 2018). The next most upregulated proteins were two PR-1 proteins, known to be secreted into the apoplast. One of these shared strong homology with a generic unknown PR-1 protein, whereas the other shared strong homology with the known PR-1a protein. PR-6 proteins consisted of two protease inhibitors, one of which showed strong homology with a form of trypsin inhibitor and the other to a xylanase inhibitor. The trypsin inhibitor is known to be secreted into the apoplast; however, the localisation of the xylanase inhibitor is currently unreported. PR-17 proteins were homologues to PR-17c and PR-17d. However, the PR-17d homologues, despite having a strong e-value (1.7e−101), shared only 67% identity. Both PR-17d and PR-17c are also known to be secreted into the apoplast (Zhang et al. 2012). The PR categories 3, 4 and 11, all consisting of chitinases, contributed to 7% of the overall fold change. One of these is a known PR-4 protein, but there was difficulty in assigning a single PR category to the remaining chitinases due to insufficient annotation of the homologues. Two glucan endo-1,3-beta-glucosidase homologues contributed to the PR-2 upregulation of 7% and are commonly found within the vacuole or as part of the cellular membrane (The UniProt Consortium 2017). Two PR-14 proteins contributed to 5% of the total fold change, both of which were classed as non-specific lipid-transfer proteins found in the membrane or extracellular regions of the plant (Kader 1997). Two PR-9 proteins contributed to another 7% of the total upregulation, both of which were peroxidases secreted into the apoplast. A final non-PR protein was also found within the top 20 most upregulated proteins and shared homology with a plant ripening-related protein.
On the other hand, the 20 most downregulated proteins ranged from a 3.4- to a 5.1-fold decrease and cellular functions could not be assigned to half of them. Of those annotated, four were chloroplast proteins, two were cytoplasmic proteins involved in the Krebs cycle, a puttive superoxide dismutase localised extracellularly and in the chloroplast, a single extracellular protein, a homologue of glucan endo-1,3-beta-glucosidase 8 (PR-2) related to plant secondary metabolism, and a single cell membrane protein related to cell wall synthesis (Fig. 2b).
Discussion
Protein composition of barley CWF
The methodology used in this study allowed the detection of 1130 plant proteins in the CWF, and the identification of differentially expressed proteins within our samples helps to profile the changes that occur in the apoplast after a fungal infection. Nonetheless, since the apoplast is the forefront of the plant–fungal interaction, it was initially expected that both plant and fungal proteins would be identified in the CWF, but none of the latter could be detected. Contributing reasons for the apparent absence of fungal proteins may be biological or technical. Some secreted fungal proteins are known to be readily taken up by host cells or specifically localised and therefore absent or at very low abundance in the apoplast. Examples of this are Pyrenophora tritici-repentis ToxA, which is internalised within mesophyll cells (Manning and Ciuffetti 2005), and Parastagonospora nodorum SnTox1, which binds to fungal cell wall chitin to protect against host chitinases (Liu et al. 2016). Given that no fungal proteins were detected at all, a more likely explanation is that the plant proteins contained in the CWF, given their comparatively much higher concentration, mask those secreted by the fungus. This is a common limitation of proteomic approaches as less abundant protein signals are not detected due to the dynamic range inherent to the mass spectrometer (Zubarev 2013), blending them with the basal noise. However, as plant defence-related proteins were well represented in the apoplastic fluid, the technique used in this study denotes an effective means to investigate extracellular proteomic responses to fungal infection in barley since it enriched for apoplastic proteins. This technique complements alternate methods which rely on the detection of secretion signals as required by whole proteome analysis or RNASeq, which ignore secreted proteins without a signal (Regente et al. 2012).
Induced barley PR protein homologues
Among the 20 most highly upregulated proteins in the susceptible barley cultivar Baudin infected with PTT, 19 belonged to PR protein families. These are part of a complex response and signalling network that intertwines with the SA, JA and ET phytohormone defence pathways which involve a significant amount of crosstalk (Kunkel and Brooks 2002). The interactions between the pathways are either antagonistic or synergistic, resulting in a cascade of reactions with the endogenous accumulation of SA-dependent defences often leading to the downregulation of JA-dependent ones (Leon-Reyes et al. 2010). Compatible interactions between plants and necrotrophic fungi seem to encompass the SA pathway, leading to HR (Al-Daoude et al. 2018). For example, necrotrophic effectors secreted into the wheat apoplast by P. nodorum have been linked to the induction of PR-1 proteins (Breen et al. 2016). PR-1 proteins are linked to the SA signalling cascade, which allows the necrotrophic fungi to succeed in its infection, and have been shown to successfully inhibit biotrophic fungi in barley (Mouradov et al. 1994).
The most highly expressed family of PR proteins in this study was PR-5, all of which belonged to the thaumatin-like protein (TLP) class. Three homologues of TLP5, TLP7 and TLP8 had a minimum alignment homology of 99.1%, and were represented in the six most upregulated proteins. TLPs are expressed in response to either infection or abiotic stress such as drought or frost. These proteins are known to rapidly accumulate in high levels and demonstrate strong antifungal activity in several plant species (Petre et al. 2011). Despite this, specific mechanisms regarding how they affect the invading fungal pathogen remain unknown (Zhang et al. 2018). Previous studies have demonstrated the lack of accumulation of wheat PR-5 in the apoplast leads to compromised resistance against the biotrophic pathogen Puccinia triticina (Zhang et al. 2018).
Furthermore, PR-5 proteins are induced as a part of the SA defence signalling pathway (Lemarié et al. 2015) which is typically activated in response to biotrophic fungi and leads to localised cell death as a part of the host HR (Bhadauria et al. 2010). This is an effective defence response against biotrophic fungi owing to their requirement of living host cells as part of their life cycle. However, the SA signalling pathway is not an efficient method of plant defence against necrotrophic fungi, considering that HR results in host cell death, supplying a necrotrophic pathogen with the nutrients required to progress infection (Glazebrook 2005). Given the hemibiotrophic nature of PTT and its main necrotrophic stage, this mechanism is probably an important feature of the compatible interaction shown by the particular isolate and host cultivar used in this study and an indicator of a weak point in a susceptible host’s defence response mechanisms.
The second most upregulated PR protein family was PR-1, containing a generic PR-1 homologue and a PR-1a homologue, the latter being the most upregulated with a 40-fold increase in expression. PR-1a is a 17 kDa protein originally described in tobacco, but also found in the compatible interaction between barley and the obligate biotroph Blumeria graminis, resulting in HR (Bryngelsson et al. 1994). In other plant species, PR-1 proteins have been identified as playing key roles in systemic acquired resistance (SAR) (Dietrich et al. 1994). SAR is a form of host resistance to an invading pathogen which acts by warning neighbouring host tissues of an impending attack through the use of long distance signalling (Kiefer and Slusarenko 2003). Moreover, SAR is a response to an accumulation of endogenous SA and, like PR-5 proteins, leads to HR. Perhaps a similar mechanism to the aforementioned interaction between P. nodorum and wheat is observed within the PTT–barley pathosystem, in which the activation of a defence pathway against biotrophs results in the benefit of the necrotrophic fungi. Both PR-1 and PR-5 proteins account for almost half of the upregulation observed in the late-stage interaction with PTT and may contribute to a successful infection.
The function of PR-1 proteins in plant defence is still poorly understood (Breen et al. 2017). However, recent reports suggest PR-1 proteins possess sterol-binding activity and could bind to environmental sterols inhibiting an invading pathogen’s growth (Gamir et al. 2017). Interestingly, most fungi are sterol prototrophs, meaning that they can produce their own sterols without relying on those from the plant. Gamir et al. (2017) suggested that vacuole-targeted specific PR-1 proteins may yield an answer: when undergoing HR, a large payload of PR-1 proteins contained within the vacuole is released into the extracellular environment, sequestering more of the fungi’s sterols than can be replaced by its own sterol biosynthesis machinery, therefore explaining the antifungal properties of these proteins. However, more studies are needed to confirm this hypothesis (Gamir et al. 2017).
Three PR-6 protein homologues were also identified among the 20 most upregulated proteins in this study and are characterised by exhibiting protease inhibitor-like activity (Carr 2010). These included a Bowman–Birk-type trypsin inhibitor (BBTI), a putative protease inhibitor similar to Bsi1 predicted to be a BBTI-like protein too, and a xylanase inhibitor which were upregulated 19-fold, 11-fold and 10-fold, respectively. Trypsin inhibitors have been shown to inhibit insect-gut digestion enzymes as well as providing some degree of defence against Fusarium head blight (FHB, Fusarium graminearum), a devastating barley disease which attacks host storage proteins through the use of subtilisin and other trypsin-like proteases (Pekkarinen et al. 2003). Possibly, barley trypsin-like proteinases have a similar role against PTT.
The third most upregulated PR-6 protein was a xylanase inhibitor homologue. An in vitro study on NFNB identified a fungal xylanase to be constitutively secreted and was proposed to play a role in symptom development (Ismail et al. 2014). Xylanase proteins have been identified as playing key roles in many other fungi in their ability to break down the host cell wall (Belien et al. 2006). This action is especially helpful for necrotrophic fungi which rely on host cell contents in order to proliferate. Therefore, the host xylanase could be suppressing PTT activity.
PR-17c and PR-17d were among the 20 most upregulated proteins, with a 22-fold and 15-fold increase, respectively. The PR-17 family of proteins is the most recently described one and contains oxalate oxidase-like and germin-like proteins (Gorjanović 2009). Barley PR-17 proteins are secreted into the apoplastic environment where they congregate and then accumulate in the mesophyll and epidermis of barley leaves (Christensen et al. 2002). Little research has been done on PR-17 proteins, although they have been associated with penetration resistance to wheat and barley powdery mildew (B. graminis) (Schweizer et al. 1999; Zhang et al. 2012). However, in these instances, upregulation of PR-17 was not found until 24 h post-infection, suggesting that the genes are expressed too late to suppress the primary penetration, but may help preventing secondary penetration events (Zhang et al. 2012).
A generic chitinase and an endochitinase 1 were detected in the upregulated group of proteins, with 16-fold and 12-fold increases, respectively. Chitinases represent a plant defence tactic which cover four main PR families: 3, 4, 8 and 11. Assigning a particular PR class to the first two most upregulated proteins is difficult due to the lack of information regarding the presence or absence of C-terminal modifications. Chitin is a vital constituent of fungal cell walls, and PR chitinases are able to hydrolyse the 1,4-linkages between N-acetylglucosamines of chitin. A range of stress factors can trigger the upregulation of chitinases such as microbial attack and plant wounding (Gorjanović 2009). There are two proposed roles of chitinases in suppressing fungal growth: the generation of elicitor molecules and the degradation of newly formed chitin chains produced by an invading fungus (de Gerhardt et al. 1997; Collinge et al. 1993).
A barwin protein showed eightfold increase in expression after PTT infection. Barwin is a class II PR-4 protein which possesses RNase activity (Caporale et al. 2004), similar to wheatwin1 and wheatwin2 RNases, two PR-4 class II proteins which have shown antifungal activity toward Fusarium culmorum (Caruso et al. 1996). In a recent in vitro study, a link has been formed between the upregulation of class II PR-4s and the upregulation of both SA and JA signalling pathways. The study also found that the class II PR-4s targeted single-stranded RNA directly linked to hyphal growth, leading to significant growth inhibition of Glomerella cingulata (Bai et al. 2013). This could suggest that the barwin protein may be involved in interfering with fungal growth by damaging the pathogen’s RNA related to its development.
Upregulated PR-2 proteins in this study included two forms of glucan endo-1,3-beta-glucosidases. Endo-glucanases hydrolyse the linkages of (1,3)-β-glucans found in fungal cell walls, resulting in cell lysis. Endo-glucanases and chitinases are thought to work synergistically amplifying their effectiveness regarding cell wall degradation (Gorjanović 2009). Products of fungal cell lysis can then be identified by the host plant as elicitors of plant defence responses. Endo-glucanases are classified into three structurally different classes differing in their enzymatic and antifungal activity.
Two PR-14 non-specific lipid-transfer proteins (ns-LTP) were upregulated 10- and 8.5-fold. Ns-LTPs aid in the transfer of different lipid classes between membranes and have broad target specificity. Upregulation of ns-LTPs can occur as a result of abiotic and environmental stresses including pathogen attack (Jung et al. 2003). It has been proposed that ns-LTP1 proteins in areal plant tissues work as transporters of cutin proteins to form a protective cuticle layer (Sterk et al. 1991). Another suggested role for ns-LTPs is to alter the membrane permeability of the fungus, as in the case of Fusarium solani (Gorjanovic et al. 2005). Both proteins identified in the present study have molecular masses above 9 kDa, suggesting that they are of the ns-LTP1 family, but their exact biological role remains to be determined.
Finally, among the PR proteins within the 20 most upregulated proteins, two peroxidases classified as PR-9 were identified. These proteins were induced 15- and 11-fold, and both proteins shared 99.4% and 83.7% homology, respectively, with the same UniProt identifier (P27337), barley peroxidase 1 (BP1) (Rasmussen et al. 1997). Peroxidases are widely distributed in the plant kingdom and have been reported to be involved in the reinforcement of the cell wall in response to pathogen invasion stimuli, cross-linking phenolic compounds to the plant cell (Almagro et al. 2009). This process has been related to disease resistance in wheat against FHB and may play a similar role in barley (Mohammadi and Kazemi 2002).
PR proteins were also overrepresented beyond the top 20 most upregulated, and the majority of the remaining upregulated thirty-two highly induced proteins were PR-related proteins. The most common of these were homologues of PR-9 (seven representatives), PR-6 (five representatives) and PR-3 (four representatives).
A highly upregulated uncharacterised protein
Within the 20 most upregulated proteins, only one non-PR protein was identified, an uncharacterised protein induced approximately 15-fold with 78.6% (e-value: 8e−92) homology to a putative rice ripening protein 2. Ripening protein gene expression has been previously described in P. teres-infected barley (Krupinska et al. 2002). It is thought that the degradation of host chloroplasts during infection leads to the phenomenon. There are no known defence functions related to barley ripening proteins; rather, the protein’s increased concentration in the CWF may be due to host cellular death, but further research is required.
Downregulated proteins
On the other side of the spectrum, homologues for the majority of the downregulated proteins (> 2.5-fold) in this study were annotated as intracellular. Although the methodology for obtaining the protein samples was designed to minimise the presence of non-secreted proteins, cytoplasmic contamination was evidenced by the detection of ribulose-1,5-bisphosphate carboxylase/oxygenase (RuBisCO) at similar levels in each sample. Therefore, the localisation of differentially expressed intracellular proteins would require further validation using different leaf tissue types and fluorescence antibody binding methods.
Among the 20 most downregulated proteins, six have roles in the photosynthetic process or are localised in the chloroplast or involved in carbon metabolism; these are the 30S ribosomal protein S10, a molecular chaperone trigger factor, an enoyl reductase 2, an ACT domain-containing protein DS12, a superoxide dismutase and a fructose-bisphosphate aldolase (Uematsu et al. 2012; The UniProt Consortium 2017). In barley infected with Puccinia (rust), and B. graminis (powdery mildew), downregulation of photosynthesis and carbohydrate metabolism-related proteins has been described (Zellerhoff et al. 2010; Molitor et al. 2011). It is thought that this photosynthesis-related downregulation is part of a shift from photosynthetic carbon assimilation to defence (Zellerhoff et al. 2010). Several major carbohydrate metabolism-related proteins were found to be downregulated, such as a ‘not fully characterised protein’, glucan endo-1,3-beta-glucosidase 8, alpha-glucosidase-like protein and a putative beta-D-xylosidase. Minor carbohydrate metabolism is also overrepresented in the downregulated proteins and includes proteins involved in the biosynthesis of the cell wall such as a xyloglucan endotransglucosylase/hydrolase.
Another highly downregulated protein was uroporphyrinogen decarboxylase. In tobacco plants, the indirect action of downregulating uroporphyrinogen decarboxylase using antisense RNA leads to the upregulation of other enzymes involved tetrapyrrole biosynthesis, resulting in cell necrosis (Mock and Grimm 1997). This is consistent with the symptoms observed in a compatible PTT interaction with barley and suggests that the necrosis may be aided in this case due to uroporphyrinogen decarboxylase downregulation.
A glucan endo-1,3-beta-glucosidase 8 protein was also found highly downregulated in the 20 most downregulated proteins. Despite glucan endo-1,3-beta-glucosidase being considered a PR-2 protein, some forms of glucan endo-1,3-beta-glucosidases are constitutively expressed (Dietz et al. 2000) possibly contributing to the basal resistance of healthy plants and providing a broad spectrum resistance against an array of pathogens or pests (Gorjanović 2009). An alignment of the two upregulated endo-glucosidases with the downregulated glucosidase shows the upregulated proteins shared approximately 53% homology with one another, whereas the downregulated endo-glucosidase only shared up to 23% with either upregulated protein. This may suggest that upon infection with a pathogen, different forms of glucosidases are preferentially upregulated to combat the invading organism.
Concluding remarks
The aim of this study was to characterise barley–PTT molecular interaction in the apoplast. Despite no fungal proteins being identified due to the reasons described above, our work contributes to the research on the barley–PTT pathosystem by describing the major apoplastic host protein changes during PTT infection and suggests some of the molecular mechanisms involved in disease development.
The most significant changes in the analysed proteome induced by PTT infection were the upregulation of PR proteins. The upregulation of PR-1 and PR-5 homologues may help to explain the susceptibility to PTT in this interaction given that these are involved in inducing plant cell death or a hypersensitive response. Conversely, proteins involved in photosynthesis and carbon metabolism were downregulated, which appears to represent a shift in photosynthetic carbon assimilation and catabolism to defence. In addition, an uroporphyrinogen decarboxylase homologue was downregulated, the action of which may promote upregulation of enzymes involved tetrapyrrole biosynthesis leading to cell necrosis. These examples help form the hypothesis that the compatible interaction between barley and PTT may be due to incorrect signal cascades being activated or hijacked, by the pathogen.
Many aspects of the molecular interaction in the barley–PTT pathosystem remain elusive. The use of SWATH-MS to compare the apoplastic-enriched proteomes of PTT-infected and non-infected barley plants has provided a glimpse of the molecular responses in a susceptible response to the pathogen. By identifying the weaknesses and strengths of the techniques in this study, a platform has been provided for further apoplastic protein expression research in the barley–PTT pathosystem. Potential future directions utilising this technique would be to include resistant barley lines, track PR protein expression changes over time during infection and refine the technique to focus on the effector molecules secreted by the pathogen.
References
Agrawal GK, Jwa N-S, Lebrun M-H, Job D, Rakwal R (2010) Plant secretome: unlocking secrets of the secreted proteins. Proteomics 10:799–827. https://doi.org/10.1002/pmic.200900514
Al-Daoude A, Jawhar M, Al-Shehadah E, Shoaib A, Orfi M, Arabi MIE (2018) Changes in salicylic acid content and pathogenesis-related (PR2) gene expression during barley-Pyrenophora teres interaction. Hell Plant Prot J 11(2):71–77. https://doi.org/10.2478/hppj-2018-0010
Almagro L, Gómez Ros LV, Belchi-Navarro S, Bru R, Ros Barceló A, Pedreño MA (2009) Class III peroxidases in plant defence reactions. J Exp Bot 60(2):377–390. https://doi.org/10.1093/jxb/ern277
Bai S, Dong C, Li B, Dai H (2013) A PR-4 gene identified from Malus domestica is involved in the defense responses against Botryosphaeria dothidea. Plant Physiol Biochem 62:23–32. https://doi.org/10.1016/j.plaphy.2012.10.016
Belien T, Robben J, Volckaert G (2006) Microbial endoxylanases: effective weapons to breach the plant cell-wall barrier or, rather, triggers of plant defense systems? Mol Plant Microbe Interact 19(10):1072–1081. https://doi.org/10.1094/MPMI-19-1072
Bhadauria V, Banniza S, Wang L-X, Wei Y-D, Peng Y-L (2010) Proteomic studies of phytopathogenic fungi, oomycetes and their interactions with hosts. Eur J Plant Pathol 126(1):81–95. https://doi.org/10.1007/s10658-009-9521-4
Breen S, Williams SJ, Winterberg B, Kobe B, Solomon PS (2016) Wheat PR-1 proteins are targeted by necrotrophic pathogen effector proteins. Plant J 88(1):13–25. https://doi.org/10.1111/tpj.13228
Breen S, Williams SJ, Outram M, Kobe B, Solomon PS (2017) Emerging insights into the functions of pathogenesis-related protein 1. Trends Plant Sci 22(10):871–879. https://doi.org/10.1016/j.tplants.2017.06.013
Bryngelsson T, Sommer-Knudsen J, Gregersen PL, Collinge DB, Ek B, Thordal-Christensen H (1994) Purification, characterization, and molecular cloning of basic PR-1-type pathogenesis-related proteins from barley. Mol Plant Microbe Interact 7(2):267. https://doi.org/10.1094/MPMI-7-0267
Caporale C, Di Berardino I, Leonardi L, Bertini L, Cascone A, Buonocore V et al (2004) Wheat pathogenesis-related proteins of class 4 have ribonuclease activity. FEBS Lett 575(1–3):71–76. https://doi.org/10.1016/j.febslet.2004.07.091
Carr JP (2010) Natural and engineered resistance to plant viruses, part II. Preface. Adv Virus Res 76:vii
Caruso C, Caporale C, Chilosi G, Vacca F, Bertini L, Magro P et al (1996) Structural and antifungal properties of a pathogenesis-related protein from wheat kernel. Protein J 15(1):35–44. https://doi.org/10.1007/BF01886809
Christensen AB, Cho BH, Næsby M, Gregersen PL, Brandt J, Madriz-Ordeñana K et al (2002) The molecular characterization of two barley proteins establishes the novel PR-17 family of pathogenesis-related proteins. Mol Plant Pathol 3(3):135–144. https://doi.org/10.1046/j.1364-3703.2002.00105.x
Collinge DB, Kragh KM, Mikkelsen JD, Nielsen KK, Rasmussen U, Vad K (1993) Plant chitinases, 3rd edn. IRL Press, Oxford, pp 31–40
de Gerhardt LBA, Sachetto-Martins G, Contarini MG, Sandroni M, de Ferreira RP, de Lima VM et al (1997) Arabidopsis thaliana class IV chitinase is early induced during the interaction with Xanthomonas campestris. FEBS Lett 419(1):69–75. https://doi.org/10.1016/S0014-5793(97)01332-X
Dempsey D, Amp A, Maris A, Silva H, Klessig DF (1998) Engineering disease and pest resistance in plants. Trends Microbiol 6(2):54–61. https://doi.org/10.1016/S0966-842X(97)01186-4
Dietrich RA, Delaney TP, Uknes SJ, Ward ER, Ryals JA, Dangl JL (1994) Arabidopsis mutants simulating disease resistance response. Cell 77(4):565–577. https://doi.org/10.1016/0092-8674(94)90218-6
Dietz K-J, Sauter A, Wichert K, Messdaghi D, Hartung W (2000) Extracellular β-glucosidase activity in barley involved in the hydrolysis of ABA glucose conjugate in leaves. J Exp Bot 51(346):937–944
Ellwood SR, Syme RA, Moffat CS, Oliver RP (2012) Evolution of three Pyrenophora cereal pathogens: recent divergence, speciation and evolution of non-coding DNA. Fungal Genet Biol 49(10):825–829. https://doi.org/10.1016/j.fgb.2012.07.003
Ellwood SR, Piscetek V, Mair WJ, Lawrence JA, Lopez-Ruiz FJ, Rawlinson C (2019) Genetic variation of Pyrenophora teres f. teres isolates in Western Australia and emergence of a Cyp51A fungicide resistance mutation. Plant Pathol 68(1):135–142. https://doi.org/10.1111/ppa.12924
Gamir J, Darwiche R, Hof P, Choudhary V, Stumpe M, Schneiter R et al (2017) The sterol-binding activity of PATHOGENESIS-RELATED PROTEIN 1 reveals the mode of action of an antimicrobial protein. Plant J 89(3):502–509. https://doi.org/10.1111/tpj.13398
Glazebrook J (2005) Contrasting mechanisms of defense against biotrophic and necrotrophic pathogens. Annu Rev Phytopathol 43(1):205–227. https://doi.org/10.1146/annurev.phyto.43.040204.135923
Gorjanovic S, Spillner E, Beljanski M, Gorjanovic R, Pavlovic M, Gojgic-Cvijanovic G (2005) Malting barley grain non-specific lipid-transfer protein (ns-LTP): importance for grain protection. J Inst Brew 111(2):99–104. https://doi.org/10.1002/j.2050-0416.2005.tb00654.x
Gorjanović S (2009) Review: biological and technological functions of barley seed pathogenesis-related proteins (PRs). J Inst Brew 115(4):334–360. https://doi.org/10.1002/j.2050-0416.2009.tb00389.x
Hammond-Kosack KE, Rudd JJ (2008) Plant resistance signalling hijacked by a necrotrophic fungal pathogen. Plant Signal Behav 3(11):993–995. https://doi.org/10.4161/psb.6292
Hanfrey C, Fife M, Buchanan-Wollaston V (1996) Leaf senescence in Brassica napus: expression of genes encoding pathogenesis-related proteins. Int J Mol Biol Mol Genet Biochem 30(3):597–609. https://doi.org/10.1007/BF00049334
Ismail I, Godfrey D, Able A (2014) Proteomic analysis reveals the potential involvement of xylanase from Pyrenophora teres f. teres in net form net blotch disease of barley. J Aust Plant Pathol Soc 43(6):715–726. https://doi.org/10.1007/s13313-014-0314-7
Jin Kim Y, Kook Hwang B (2000) Pepper gene encoding a basic pathogenesis-related 1 protein is pathogen and ethylene inducible. Physiol Plant 108(1):51–60. https://doi.org/10.1034/j.1399-3054.2000.108001051.x
Jung HW, Kim W, Hwang BK (2003) Three pathogen-inducible genes encoding lipid transfer protein from pepper are differentially activated by pathogens, abiotic, and environmental stresses. Plant Cell Environ 26(6):915–928. https://doi.org/10.1046/j.1365-3040.2003.01024.x
Kader J-C (1997) Lipid-transfer proteins: a puzzling family of plant proteins. Trends Plant Sci 2(2):66–70. https://doi.org/10.1016/S1360-1385(97)82565-4
Kiefer I, Slusarenko A (2003) The pattern of systemic acquired resistance induction with the Arabidopsis rosette in relation to the pattern of translocation. Plant Physiol 132(2):840–847. https://doi.org/10.1104/pp.103.021709
Kosiada T (2008) Influence of temperature and daylight length on barley infection by Pyrenophora teres. J Plant Prot Res 48(1):9–15. https://doi.org/10.2478/v10045-008-0002-0
Krasny L, Bland P, Kogata N, Wai P, Howard BA, Natrajan RC et al (2018) SWATH mass spectrometry as a tool for quantitative profiling of the matrisome. J Proteomics 189:11–22. https://doi.org/10.1016/j.jprot.2018.02.026
Krupinska K, Haussühl K, Schäfer A, van Der Kooij TAW, Leckband G, Lörz H et al (2002) A novel nucleus-targeted protein is expressed in barley leaves during senescence and pathogen infection. Plant Physiol 130(3):1172–1180. https://doi.org/10.1104/pp.008565
Kunkel BN, Brooks DM (2002) Cross talk between signaling pathways in pathogen defense. Curr Opin Plant Biol 5(4):325–331. https://doi.org/10.1016/S1369-5266(02)00275-3
Lancashire PD, Bleiholder H, Boom TVD, Langelüddeke P, Stauss R, Weber E et al (1991) A uniform decimal code for growth stages of crops and weeds. Ann Appl Biol 119(3):561–601. https://doi.org/10.1111/j.1744-7348.1991.tb04895.x
Lemarié S, Robert-Seilaniantz A, Lariagon C, Lemoine J, Marnet N, Jubault M et al (2015) Both the jasmonic acid and the salicylic acid pathways contribute to resistance to the biotrophic clubroot agent Plasmodiophora brassicae in Arabidopsis. Plant Cell Physiol 56(11):2158–2168. https://doi.org/10.1093/pcp/pcv127
Leon-Reyes A, Van der Does D, De Lange ES, Delker C, Wasternack C, Van Wees SCM et al (2010) Salicylate-mediated suppression of jasmonate-responsive gene expression in Arabidopsis is targeted downstream of the jasmonate biosynthesis pathway. Planta 232(6):1423–1432. https://doi.org/10.1007/s00425-010-1265-z
Lightfoot D, Able A (2010) Growth of Pyrenophora teres in planta during barley net blotch disease. Austr Plant Pathol 39(6):499–507. https://doi.org/10.1071/AP10121
Liljeroth E, Marttila S, Bothmer R (2005) Immunolocalization of defence-related proteins in the floral organs of barley (Hordeum vulgare L). J Phytopathol 153(11–12):702–709. https://doi.org/10.1111/j.1439-0434.2005.01043.x
Liu Z, Ellwood Simon R, Oliver Richard P, Friesen Timothy L (2010) Pyrenophora teres: profile of an increasingly damaging barley pathogen. Mol Plant Pathol 12(1):1–19. https://doi.org/10.1111/j.1364-3703.2010.00649.x
Liu Z, Gao Y, Kim YM, Faris JD, Shelver WL, Wit PJGM et al (2016) SnTox1, a Parastagonospora nodorum necrotrophic effector, is a dual-function protein that facilitates infection while protecting from wheat-produced chitinases. New Phytol 211(3):1052–1064. https://doi.org/10.1111/nph.13959
Van Loon LC, Van Kammen A (1970) Polyacrylamide disc electrophoresis of the soluble leaf proteins from Nicotiana tabacum var. ‘Samsun’ and ‘Samsun NN’: II. Changes in protein constitution after infection with tobacco mosaic virus. Virology 40(2):199–211. https://doi.org/10.1016/0042-6822(70)90395-8
Manning VA, Ciuffetti LM (2005) Localization of Ptr ToxA produced by Pyrenophora tritici-repentis reveals protein import into wheat mesophyll cells. Plant Cell 17(11):3203–3212. https://doi.org/10.1105/tpc.105.035063
Mills ENC, Tatham AS (2003) Allergens. In: Caballero B (ed) Encyclopedia of food sciences and nutrition, 2nd edn. Academic Press, Oxford, pp 143–150
Mock HP, Grimm B (1997) Reduction of uroporphyrinogen decarboxylase by antisense RNA expression affects activities of other enzymes involved in tetrapyrrole biosynthesis and leads to light-dependent necrosis. Plant Physiol 113(4):1101. https://doi.org/10.1104/pp.113.4.1101
Mohammadi M, Kazemi H (2002) Changes in peroxidase and polyphenol oxidase activities in susceptible and resistant wheat heads inoculated with Fusarium graminearum and induced resistance. Plant Sci 162(4):491–498. https://doi.org/10.1016/S0168-9452(01)00538-6
Molitor A, Kogel K-H, Waller F, Zajic D, Voll LM, Pons-Kühnemann J et al (2011) Barley leaf transcriptome and metabolite analysis reveals new aspects of compatibility and Piriformospora indica-mediated systemic induced resistance to powdery mildew. Mol Plant Microbe Interact 24(12):1427–1439. https://doi.org/10.1094/MPMI-06-11-0177
Mott GA, Middleton MA, Desveaux D, Guttman DS (2014) Peptides and small molecules of the plant–pathogen apoplastic arena. Front Plant Sci 5:677. https://doi.org/10.3389/fpls.2014.00677
Mouradov A, Mouradova E, Scott K (1994) Gene family encoding basic pathogenesis-related 1 proteins in barley. Int J Mol Biol Mol Genet Biochem 26(1):503–507. https://doi.org/10.1007/BF00039561
Nouchi I, Hayashi K, Hiradate S, Ishikawa S, Fukuoka M, Chen CP, Kobayashi K (2012) Overcoming the difficulties in collecting apoplastic fluid from rice leaves by the infiltration-centrifugation method. Plant Cell Physiol 53:1659–1668. https://doi.org/10.1093/pcp/pcs102
Pekkarinen AI, Sarlin TH, Laitila AT, Haikara AI, Jones BL (2003) Fusarium species synthesize alkaline proteinases in infested barley. J Cereal Sci 37(3):349–356. https://doi.org/10.1006/jcrs.2002.0512
Petre B, Major I, Rouhier N, Duplessis S (2011) Genome-wide analysis of eukaryote thaumatin-like proteins (TLPs) with an emphasis on poplar. BMC Plant Biol 11(1):33. https://doi.org/10.1186/1471-2229-11-33
Rasmussen CB, Henriksen A, Abelskov AK, Jensen RB, Rasmussen SK, Hejgaard J et al (1997) Purification, characterization and stability of barley grain peroxidase BP 1, a new type of plant peroxidase. Physiol Plant 100(1):102–110. https://doi.org/10.1111/j.1399-3054.1997.tb03459.x
Regente M, Pinedo M, Elizalde M, de la Canal L (2012) Apoplastic exosome-like vesicles: a new way of protein secretion in plants? Plant Signal Behav 7(5):544–546. https://doi.org/10.4161/psb.19675
Schweizer P, Pokorny J, Abderhalden O, Dudler R (1999) Transient assay system for the functional assessment of defense-related genes in wheat. Mol Plant Microbe Interact 12(8):647–654. https://doi.org/10.1094/MPMI.1999.12.8.647
Smedegård-Petersen V (1971) Pyrenophora teres f. maculata f. nov. and Pyrenophora teres f. teres on barley in Denmark. Aarsskrift Kongelige Veterinear of Landbohojskole. Aug. Bangs Boghandel, Copenhagen, pp 124–144
Speakman J, Pommer EH (1986) A simple method for producing large volumes of Pyrenophora teres spore suspension. Bull Br Mycol Soc 20(2):129–130. https://doi.org/10.1016/S0007-1528(86)80041-4
Sterk P, Booij H, Schellekens GA, Van Kammen A, De Vries SC (1991) Cell-specific expression of the carrot EP2 lipid transfer protein gene. Plant Cell 3(9):907–921. https://doi.org/10.2307/3869154
The UniProt Consortium (2017) UniProt: the universal protein knowledgebase. Nucl Acids Res 45(D1):D158–D169. https://doi.org/10.1093/nar/gkw1099
Uematsu K, Suzuki N, Iwamae T, Inui M, Yukawa H (2012) Increased fructose 1,6-bisphosphate aldolase in plastids enhances growth and photosynthesis of tobacco plants. J Exp Bot 63(8):3001–3009. https://doi.org/10.1093/jxb/ers004
Vleeshouwers VGAA, Oliver RP (2015) Effectors as tools in disease resistance breeding against biotrophic, hemibiotrophic, and necrotrophic plant pathogens. Mol Plant Microbe Interact 2015(1):17. https://doi.org/10.1094/MPMI-10-13-0313-CR.test
Wang X, Tang C, Deng L, Cai G, Liu X, Liu B, Han Q, Buchenauer H, Wei G, Han D, Huang L, Kang Z (2018) Characterization of a pathogenesis-related thaumatin-like protein gene TaPR5 from wheat induced by stripe rust fungus. J Plant Pathol 86(4):295
Witzel K, Shahzad M, Matros A, Mock H-P, Mühling KH (2011) Comparative evaluation of extraction methods for apoplastic proteins from maize leaves. Plant Methods 7:48. https://doi.org/10.1186/1746-4811-7-48
Zellerhoff N, Himmelbach A, Dong W, Bieri S, Schaffrath U, Schweizer P (2010) Nonhost resistance of barley to different fungal pathogens is associated with largely distinct, quantitative transcriptional responses. Plant Physiol 152(4):2053. https://doi.org/10.1104/pp.109.151829
Zhang WJ, Pedersen C, Kwaaitaal M, Gregersen PL, Mørch SM, Hanisch S et al (2012) Interaction of barley powdery mildew effector candidate CSEP0055 with the defence protein PR17c. Mol Plant Pathol 13(9):1110–1119. https://doi.org/10.1111/j.1364-3703.2012.00820.x
Zhang J, Wang F, Liang F, Zhang Y, Ma L, Wang H et al (2018) Functional analysis of a pathogenesis-related thaumatin-like protein gene TaLr35PR5 from wheat induced by leaf rust fungus. BMC Plant Biol 18(1):76. https://doi.org/10.1186/s12870-018-1297-2
Zubarev RA (2013) The challenge of the proteome dynamic range and its implications for in-depth proteomics. Proteomics 13(5):723–726. https://doi.org/10.1002/pmic.201200451
Acknowledgements
This research was funded by a Curtin University student grant with support from the Grain Research and Development Corporation Program CUR00023 P5. The SWATH-MS analysis was facilitated through the Australian Proteome Analysis Facility, supported under the Australian Government’s National Collaborative Research Infrastructure Strategy (NCRIS). Special thanks to Julie Lawrence, Nola D’Souza, Vladimir Piscetek and Robert Syme who were always willing to provide advice and help in the laboratory.
Author information
Authors and Affiliations
Contributions
All authors contributed in the design of the project. Experiments and analysis were performed by KH supported by MJM-G and SE. KZ provided advice on sample stability and proteomic analysis. KH drafted the manuscript, and all authors contributed to the final version of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hassett, K., Ellwood, S.R., Zulak, K.G. et al. Analysis of apoplastic proteins expressed during net form net blotch of barley. J Plant Dis Prot 127, 683–694 (2020). https://doi.org/10.1007/s41348-020-00318-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s41348-020-00318-w