Abstract
Purpose of Review
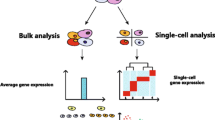
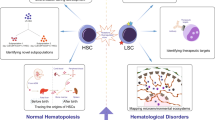
Hematopoietic stem cells (HSCs) are responsible for the lifelong production of blood and immune cells. This review provides an overview of how single-cell (multi)-omic approaches have recently advanced our understanding of healthy hematopoiesis, hematological malignancies, and the stem cell niche.
Recent Findings
Single-cell technologies have revealed tremendous heterogeneity of the HSC compartment, conflicting with the classical view of hematopoiesis. Large-scale single-cell approaches mapping the entire hematopoietic system have enabled an ordering of the observed heterogeneity along meaningful differentiation and cell-state trajectories. These studies provided novel insights into lineage commitment pathways and led to the suggestion of advanced models of hematopoiesis. Single-cell multi-omic technologies, where several entities of individual cells are measured in parallel, have permitted the fine mapping of clonal and developmental differentiation hierarchies, and revealed the molecular consequences of clonal diversification.
Summary
Recent single-cell approaches have changed our perception of healthy hematopoiesis, provided an understanding of hematological malignancies at unprecedented depth, and revealed new insights into the stem cell niche.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The majority of blood and immune cells are constantly replenished by a small population of hematopoietic stem cells (HSCs) located in the bone marrow [1]. Multipotent HSCs give rise to progressively restricted progenitors, finally resulting in the generation of mature blood and immune cells. In the classical model of hematopoiesis, stem and progenitor cell populations have been conceptualized to form discrete cell types resulting in a rigid tree-like model [2, 3]. In this model, HSCs can give rise to additional HSCs in a process called self-renewal, or produce multipotent progenitors (MPPs) that exhibit reduced self-renewal capacity while maintaining multipotency. Subsequently, MPPs diversify into oligopotent progenitors, which display restricted lineage differentiation capacity, and finally produce lineage-restricted progenitors and mature cell types of the blood and immune system (see below).
Basic single-cell technologies, such as flow cytometry, colony forming assays, and single-cell transplantations, have been utilized for decades in the field of hematology [4]. These single-cell approaches have revealed extensive molecular and functional heterogeneity within the hematopoietic stem and progenitor cell (HSPC) compartment with regard to cell cycle activity, lineage bias, metabolic activity, or self-renewal capacity [5,6,7,8]. To account for the observed heterogeneity, a variety of subpopulations that enrich for particular cell states or phenotypes have been introduced [9,10,11,12,13,14,15,16,17,18,19]. Such a compartmentalization of the early hematopoietic system has proven highly valuable, but reinforced the notion that molecular and cellular changes along differentiation pathways and cell states typically occur stepwise. Single-cell analyses revealed that such populations still remain heterogenous [19, 20••, 21,22,23,24,25,26,27, 28••, 29]. The recent development of powerful single-cell omic approaches spanning the entire hematopoietic system has facilitated an ordering of the observed heterogeneity along differentiation trajectories, suggesting more gradual rather than discrete, step-wise transitions between cell states [30,31,32,33,34, 35••, 36••]. Together with functional single-cell and lineage tracing approaches, these studies have enabled a more comprehensive understanding of hematopoiesis, including new insights into lineage commitment hierarchies [5,6,7,8].
Large-scale single-cell approaches have also recently been exploited to study the bone marrow microenvironment responsible for maintaining HSCs and orchestrating their differentiation into mature blood cells [37,38,39,40,41]. These studies have provided fundamental insights into the niche composition and clarified some of the existing controversies in the field. Finally, powerful single-cell multi-omic technologies, where several entities of single cells such transcriptome, mutations, chromatin accessibility, and surface markers are measured in parallel, have recently been developed [42,43,44,45,46,47,48]. These technologies have significantly advanced our understanding of hematological malignancies deriving from HSPCs.
In the following, I discuss how recent single-cell (multi-)omic studies have transformed our understanding of normal and malignant hematopoiesis, as well as the bone marrow niche.
HSC Heterogeneity and Conflicts with the Classical Model of Hematopoiesis
Over the past years, single-cell approaches have revealed extensive molecular and functional heterogeneity within the HSPC compartment, conflicting with the traditional view of hematopoiesis [5,6,7,8]. In this section, I will briefly review the observed heterogeneity in the HSPC compartment, and introduce concepts of lineage-biased HSCs and transcriptional lineage priming. In the following section, I will set the observed heterogeneity in relation to newly introduced models of hematopoiesis.
Functional Heterogeneity of HSPCs
Functional heterogeneity of HSPCs has been recognized for decades [4]. In particular, differences in self-renewal capacities of HSCs have been studied in detail using transplantation of bulk subpopulations or single HSCs [2, 17,18,19, 20••, 21, 25, 27, 49,50,51]. Fluorescence-activated cell sorting (FACS) has been used to purify subpopulations that distinguish HSCs with high self-renewal capacity from MPPs. For example, the SLAM markers CD150 and CD48, the surface markers CD34 and Flt3, or EPCR have been used to identify HSCs with long-term self-renewal capacity, displaying the combined phenotype of Lineage−Sca1+cKit+CD150+CD48−Flt3−CD34−EPCR+ [13, 17,18,19, 52]. Combinations of these markers have also been used to define a plethora of MPPs with different self-renewal capacities, lineage biases, and molecular characteristics [10, 13]. In the classical model, self-renewal capacity is restricted to HSCs. However, elegant single-cell transplantation studies demonstrated that lineage-restricted progenitors present in the phenotypic HSC compartment (here: Lin−Sca-1+cKit+CD34−) might exhibit high self-renewal capacities as well, exceeding those of some stem and multipotent progenitor cells [20••]. In particular, some transplanted phenotypic HSCs showed exclusive reconstitution of platelets, platelets and erythroid cells or platelet, erythroid, and myeloid cells with the absence of other lineages, both in primary and secondary transplantations. This observation challenged the strict mutual dependence of multipotency and self-renewal capacity. However, others have argued that this phenomenon might be restricted to the megakaryocytic lineage (see below) [53].
Moreover, single-cell transplantation and molecular barcoding experiments have demonstrated that the reconstitution patterns of individual HSCs are rarely identical [19, 21, 25, 51, 54,55,56]. In particular, certain lineages are frequently produced more abundantly than others. In this context, the term “lineage bias” has been introduced, referring to the relatively higher production of one or several lineages at the expense of the remaining lineages. The existence of megakaryocyte-, erythrocyte-, pan-myeloid- and pan-lymphoid-biased HSCs has been proposed [16, 18, 19, 20••, 29, 36••, 52, 57, 58]. It is important to note that a heterogenous lineage contribution of a single clone might be the consequence of an intrinsic lineage bias, but might also be imparted by the microenvironment, or simply reflect the consequence of the different kinetics of individual blood lineages [53]. Time-resolved lineage tracing approaches coupled to single-cell transcriptomic readouts will likely unravel the exact nature of HSC lineage biases [56].
Molecular Heterogeneity
In accordance with the existence of lineage-biased HSCs, the expression of transcriptional programs associated with lineage committed progenitors has been observed in individual cells of the phenotypic HSC compartment using single-cell transcriptomic profiling [17, 29, 30, 32,33,34, 35••, 59]. For example, von Willebrand factor and several other genes associated with the megakaryocytic lineage are expressed in a subset of HSCs [60•].This phenomenon has been termed “transcriptional lineage priming”. The use of indexed FACS surface markers common to both single-cell RNAseq data and single-cell ex vivo culture data has allowed to quantitatively link the direction of transcriptional lineage priming to functional properties [35••]. These analyses have revealed that the degree and direction of transcriptional lineage priming is quantitatively linked to the functional bias, with HSCs displaying a certain type of transcriptional priming exhibiting a higher likelihood of commitment into the respective lineage [35••]. In line with heterogeneity in gene expression programs, a study measuring chromatin accessibility using single-cell assays for transposase-accessible chromatin by sequencing (ATACseq) revealed cellular variations among HSPCs at the regulatory level consistent with lineage biases towards different hematopoietic branches [61]. DNA methylation and chromatin modifications have so far been mainly studied at the bulk level in highly purified HSPC compartments [13, 62,63,64,65]. These analyses suggest that extensive heterogeneity might already be imprinted at the epigenetic level in HSC subsets. To address this point mechanistically, an elegant study made use of a mouse model in which individual HSC clones express unique combinations of fluorescent tags [66]. This permitted the simultaneous tracking of clonal fates at the functional level, as well as the mapping of transcriptional and epigenomic programs underlying the clonal heterogeneity. The study revealed that HSC function highly corresponds to its underlying epigenetic configuration under homeostatic and stress conditions [66]. In the future, combined readouts of transcriptome and epigenomic marks from single cells might be instrumental for a detailed rewiring of the cause-effect relationship between epigenetic remodeling and transcription in mediating HSC fate decisions [48, 67].
Cellular Heterogeneity
HSCs display pronounced heterogeneity in cellular processes, such as cell metabolism, autophagy, and cell cycle activity. The most potent HSCs are thought to depend on glycolysis, exhibit low mitochondrial respiration, and, consequently, produce low levels of reactive oxygen species (ROS) [68]. Importantly, a fraction of HSCs are maintained in a long-term quiescent state, termed dormancy, which is associated with a high self-renewal potential [14, 69]. Single-cell analyses in mouse and human have indicated that dormancy is associated with a biosynthetic inactive state and low translational activity [35••, 70]. Similar to the state of dormancy, high autophagy levels in HSCs have been associated with self-renewal capacity, a low metabolic state and quiescence [71, 72]. It is tempting to speculate that the states of dormancy and autophagy are directly linked.
In case of emergency, such as viral infections or blood loss, dormant HSCs are recruited into the cell cycle to accelerate blood production [14, 73,74,75,76]. HSC activation and differentiation is associated with increased metabolic activity, a switch to oxidative phosphorylation, elevated ROS production, and loss of self-renewal activity (Fig. 1).
Hematopoietic stem cell commitment model. a Overview. b Self-renewal capacity and indication of direct lineage commitment to the megakaryocytic lineage. c Indication of transcriptional lineage priming. d Indication of cellular activity. Mk megakaryocyte, Ery erythrocytes, Eosino eosinophils, Baso basophils, Neutro neutrophils, Mono monocytes, cDC conventional dendritic cells, pDCs plasmacytoid dendritic cells, NK cells natural killer cells
Together, extensive molecular, cellular, and functional heterogeneity is observed in the HSPC compartment. While a large extent of this heterogeneity can be linked to the exit of dormancy, change of metabolic states, and preparation for lineage commitment, additional sources of heterogeneity remain to be investigated.
Hematopoietic Landscapes Inferred from Single-Cell Omic Approaches
Large-scale single-cell transcriptomics spanning the entire hematopoietic system of the bone marrow have provided important insights into the organization of hematopoiesis, and enabled the translation of the observed heterogeneity into meaningful differentiation and cell-state trajectories (Fig. 1). While HSC and MPP populations have been conceptualized to form discrete cell types in the classical tree model of hematopoiesis, large-scale single-cell transcriptomic studies in mouse, zebrafish, and human have revealed that HSCs do not acquire transcriptionally stable, discrete cell stages upon lineage commitment, but rather gradually pass through a continuum of differentiation states [32,33,34, 35••, 54, 77]. In trajectory analyses of single-cell transcriptome data, HSCs initially upregulate transcriptional programs associated with lineage priming towards the megakaryocyte/erythrocyte/eosinophil/basophil versus the neutrophil/lymphoid direction, followed by branching into separate lineages and activation of lineage-specific transcriptional programs [32, 35••, 56, 78]. This pattern is observed not only in adult hematopoiesis, but also during embryonic hematopoiesis at the fetal liver stage [31]. Integration of single-cell culture or lineage tracing data with single-cell transcriptomic data suggested that early transcriptional lineage priming (megakaryocyte/erythrocyte/eosinophil/basophil versus the neutrophil/lymphoid priming) is associated with an increased likelihood of lineage commitment, whereas subsequent activation of lineage-specific transcriptional programs is linked to definitive lineage commitment [35••, 54, 56]. Importantly, lineage commitment occurs earlier than previously anticipated as revealed by single-cell molecular and functional analyses of progenitors downstream of HSCs [21, 22, 28••, 35••, 79]. These studies demonstrated that the majority of cells suggested to be oligopotent by the classical model of hematopoiesis (CMPs, MEPs, GMPs) have already acquired transcriptional lineage programs associated with functional lineage commitment [21, 22, 28, 35••, 79]. While oligopotent HSPCs and transcriptional mixed-lineage states do exist, they are much less frequent than suggested by the classical model [20,21,22, 35••, 80].
Highly similar findings that include the continuous nature of cell state transitions and branching hierarchies were made by measuring chromatin accessibility, instead of single-cell transcriptomes in large-scale single-cell ATACseq studies [48, 61]. To account for the continuous nature of hematopoiesis, continuum-based models have been suggested that are capable of explaining single-cell data much more accurately [5, 7, 35••, 61, 80]. In this framework, quiescent and metabolically inactive HSCs are maintained in a flat valley at the top of the hierarchy [35••, 70]. Upon cell cycle induction, HSCs become biosynthetically active and subsequently acquire transcriptional lineage priming in a gradual manner. While in the early phases combinatorial lineage priming might be acquired, unidirectional priming becomes more prevalent over time, increasing the likelihood of future lineage commitment. Initially, barriers between lineages are small, allowing some plasticity. However, rapidly, transcriptional lineage programs associated with lineage commitment and cell-type manifestation are engaged.
Lineage Commitment Hierarchies
According to the classical model of hematopoiesis, the first bifurcation point into common lymphoid progenitor (CLPs) and common myeloid progenitor (CMPs) separates lymphoid lineages (B, T, and NK cell lineages) from all myeloid, megakaryocyte, and erythrocyte lineages, followed by further sub-branching [2, 3, 11, 12]. This hierarchy was first revised when it was noted that megakaryocytes and erythrocytes branch off from lymphoid and myeloid progenitors, marked by a population termed lymphoid primed multipotent progenitor (LMPP) in mice [17] or multilymphoid progenitor (MLP) in human [81]. However, in these analyses, all myeloid lineages were investigated together and not individually. When myeloid lineages were investigated separately using molecular and functional single-cell approaches, it was noted that eosinophil and basophil lineages share common precursors with the megakaryocyte/erythrocytic lineages, whereas the neutrophil, dendritic cell, and monocyte lineages initially co-develop with lymphoid branches, followed by further sub-branching [79, 82, 83] (Fig. 1). This is in line with trajectory analyses from large-scale single-cell transcriptome data of HSPCs (see above).
Megakaryocyte Lineage Commitment—a Special Case?
A decade ago, it was already noted that despite their apparent phenotypic differences, HSCs and the megakaryocytic lineage share many common features [84]. These include commonly used transcription factors (e.g., Runx-1, Gata2, Evi-1, Tal-1), common surface receptors (e.g., c-mpl), and specific signaling pathways [84]. Later, it was shown that the phenotypic HSC compartment contains (i) megakaryocyte-biased HSCs that reside at the apex of the differentiation hierarchy [60•] and (ii) stem-like megakaryocyte progenitors that share many features with HSCs, exhibit high self-renewal capacity, but are committed to the megakaryocytic lineage [20••, 29]. An elegant study recently addressed the relationship between HSCs and the megakaryocytic lineage in unperturbed, native hematopoiesis [36]. For this purpose, the authors made use of transposon-based tagging of HSPCs combined with single-cell transcriptomic readouts to trace lineage relationships. This analysis revealed megakaryocyte commitment to be the predominant native fate outcome of HSCs (Fig. 1b). The close connection between HSCs and the megakaryocytic lineage might serve as an emergency pathway. For example, dramatic losses of platelets that might occur during infections induce rapid proliferation and differentiation of megakaryocyte-primed cells of the HSC compartment [29, 60•]. On the molecular level, stem-like megakaryocyte progenitors express Mk transcripts, but protein synthesis is suppressed during homeostasis. Upon inflammatory signaling initiation, translation of Mk transcripts is induced, mediating an emergency differentiation program that rapidly restores platelet levels [29].
Single-Cell Multi-omic Approaches in Hematological Disease
Leukemias frequently develop from healthy stem progenitor or precursor cells by a stepwise acquisition of genetic lesions, resulting in a differentiation block and accumulation of leukemia cells in the bone marrow. Understanding the complex clonal hierarchies in cancer is of great importance, since small sub-clones present at diagnosis might expand, and ultimately cause therapy resistance and relapse. While single-cell mutational profiling provides highly detailed maps of clonal hierarchies, combined single-cell transcriptomic and genomic assays offer the possibility of determining the molecular consequences of clonal evolution. In recent years, several methods that permit combined transcriptomic and genomic readouts have been introduced [42,43,44,45,46,47, 85], where mutations are called either indirectly from cDNA of scRNAseq data or separately from genomic DNA. Such approaches have been applied to distinct hematological malignancies, including acute myeloid leukemia (AML) and myeloproliferative neoplasms (MPNs). Chronic myeloid leukemia (CML) is an MPN typically driven by the oncogenic fusion of BCR and ABL1. In an elegant single-cell multi-omic study of CML, the Mead group combined high sensitivity BCR-ABL detection with whole transcriptome scRNAseq [43]. This allowed the discrimination of mutated CML cells at high precision and sensitivity, enabled the prediction of therapy response to tyrosine kinase inhibitors (TKIs), and led to the characterization of a quiescent CML stem cell population persisting throughout TKI therapy. In BCR-ABL-negative MPNs, the CALR gene has recently been described to be recurrently mutated [86]. To determine the effects of CALR mutations in MPNs, combined CALR genotyping and transcriptome analyses of thousands of single-cells has recently been performed using a novel high-throughput, droplet-based approach, termed genotyping of transcriptomes (GoT) [45]. This revealed that CALR mutations impart different effects on the transcriptome, dependent on the cellular state and disease type. For example, in patients with essential thrombocythemia, an MPN associated with an overproduction of platelets, CALR mutations resulted specifically in a pronounced cell cycle activation of mutant megakaryocyte progenitors [45].
To understand complex sub-clonal structures, protocols that allow the simultaneous mapping of many mutations in scRNAseq experiments have been developed and applied to bone marrow samples of MPN and AML patients [42, 44, 45, 47]. In AML, such studies have enabled the fine mapping of clonal and developmental differentiation hierarchies, revealed gene expression programs associated with pre-leukemic (e.g., TET2, SRSF2) and leukemic (e.g., CEBPA) mutations, and suggested that AMLs with monocyte-like phenotypes preferentially display immune modularity activity and T cell suppression [42, 44]. Importantly, computational pipelines for calling mutations directly from standard droplet-based scRNAseq approaches have been developed [85], facilitating multi-omic analyses without additional technical adaptations, albeit with much higher dropout rates if compared with specialized methods. Moreover, the possibility of tracking genetic clones by measuring mitochondrial mutations in single-cell genomic assays has been demonstrated, offering an elegant and efficient approach of tracing lineage relationships independent of genomic mutations [87].
Beyond mutations, a recent singe-cell multi-omic study combined simultaneous measurements of transcriptomes, chromatin accessibility, and surface markers of single cells [48]. Using this approach, the authors generated maps of healthy hematopoiesis and mixed-phenotype acute leukemia. Integrative analyses identified putative transcription factors which regulate leukemia-specific expression programs.
Together, these single-cell multi-omic studies provide a framework for future integrative analyses and set the starting point for an in-depth understanding of hematological malignancies.
The Hematopoietic Stem Cell Niche at a Single-Cell Resolution
The bone marrow (BM) microenvironment, also referred to as the niche, plays a fundamental role in the maintenance and differentiation of HSCs [88, 89]. A multitude of niche cell types have been implicated in regulating HSC maintenance, including leptin receptor (Lepr) expressing cells, CXCL12 abundant reticular (CAR) cells, Nestin, and Ng2 expressing cells, as well as others [90,91,92,93,94,95,96]. The most intensively studied factors required for HSC maintenance in the BM are CXCLl12 and SCF [90,91,92, 94,95,96]. Conventional approaches to study the niche typically make use of genetic labelling based on a single marker gene, in combination with genetic deletion of molecular factors, cell type ablation, or imaging. These approaches have provided fundamental insights into the contribution of niche cell types and cytokines required for HSC maintenance. However, the use of single genes for cell type definition has frequently resulted in the labelling of heterogenous cell populations, and has therefore led to some controversies with fundamental questions remaining unaddressed.
Recently, single-cell transcriptomic studies from non-hematopoietic cells of the BM have elucidated the niche composition and clarified some of the controversies [37,38,39,40,41]. Notably, such studies permitted the unbiased identification and molecular characterization of known as well as previously unknown BM-resident cell types. For example, previously described CAR cells were shown to consist of subpopulations differing in their osteogenic versus adipogenic transcriptional priming, with the latter highly overlapping with previously described Lepr expressing cells. In line with previous studies, systematic assessments of cytokine synthesis across all BM cell types demonstrated that Lepr-expressing CAR cell populations were the main producers of CXCL12 and SCF, with relatively minor contributions of arterial endothelial cells. Remarkably, among all BM-resident cell types, CAR cell populations devote the largest proportion of their transcriptional activity to cytokine synthesis, suggesting that they act as professional cytokine producing cells [37].
Combining single-cell with laser-capture microdissection-based spatial transcriptomics allowed the systematic localization of BM cell types to distinct niches and enabled the spatial mapping of cytokine synthesis [37]. This approach demonstrated that CAR cell subsets differentially localize to either sinusoidal endothelia in the case of adipo-primed CAR cells, or to arterioles and non-vascular regions in the case of osteo-primed CAR cells. In line with this, in situ measurement of cytokine production demonstrated that the highest production of HSC factors occurs around both sinusoidal and arterial vessels. This suggests that unique niches are established by the differential localization of professional cytokine producing cells to specific regions in the BM. In line with previous studies, these findings point to a particular importance of (peri)-vascular niches for the production of HSC maintenance factors. Interestingly, osteo-primed CAR cells were also found to localize to the trabecular part of the bone, suggesting a potential role in osteogenesis [40].
Together, single-cell studies of the BM niche have provided valuable insights into the HSC niche organization, and clarified the cellular and spatial sources of key HSC factors (see [41] for a detailed review). In the future, imaging mass cytometry and advanced spatial transcriptomic approaches, in combination with conventional approaches, could lead to a deeper understanding of the complex molecular and spatial organization of the bone marrow niche.
Conclusions
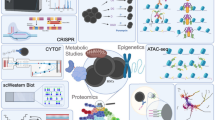
Single-cell approaches have considerably refined our perception of healthy hematopoiesis, hematological malignancies, and the bone marrow niche. In the future, integrative approaches should deliver multi-scale insights into the phenotypes, function, and spatial organization of the hematopoietic system at the single-cell resolution. Descriptive multi-scale phenotyping will include single-cell readouts at the transcriptome, genome, chromatin accessibility, DNA methylation, B cell and T cell receptor sequence, and surface proteome (CITE-seq, AB-seq-related technologies) level [48, 97,98,99], combined with lineage tracing approaches or pooled CRISPR screens in order to bridge descriptive phenotypes with mechanistic or functional data [56, 100,101,102,103].
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Orkin SH, Zon LI. Hematopoiesis: an evolving paradigm for stem cell biology. Cell [Internet]. 2008 Feb 22 [cited 2017 Oct 17];132(4):631–44. Available from: http://linkinghub.elsevier.com/retrieve/pii/S0092867408001256
Seita J, Weissman IL. Hematopoietic stem cell: self-renewal versus differentiation. Wiley Interdiscip Rev Syst Biol Med [Internet]. 2010 Nov [cited 2017 Oct 17];2(6):640–53. Available from: http://www.ncbi.nlm.nih.gov/pubmed/20890962.
Weissman IL, Shizuru JA. The origins of the identification and isolation of hematopoietic stem cells, and their capability to induce donor-specific transplantation tolerance and treat autoimmune diseases. Blood [Internet]. 2008 Nov 1 [cited 2017 Oct 17];112(9):3543–53. Available from: http://www.ncbi.nlm.nih.gov/pubmed/18948588.
Copley MR, Beer PA, Eaves CJ. Hematopoietic stem cell heterogeneity takes center stage. Cell Stem Cell [Internet]. 2012 Jun 14 [cited 2017 Oct 17];10(6):690–7. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22704509.
Laurenti E, Göttgens B. From haematopoietic stem cells to complex differentiation landscapes. Nature Nat Publ Group. 2018;553:418–26.
Jacobsen SEW, Nerlov C. Haematopoiesis in the era of advanced single-cell technologies. Nature Cell Biology Nat Publ Group. 2019;21:2–8.
Haas S, Trumpp A, Milsom MD. Causes and consequences of hematopoietic stem cell heterogeneity. Cell Stem Cell. Cell Press. 2018;22:627–38.
Yamamoto R, Wilkinson AC, Nakauchi H. Changing concepts in hematopoietic stem cells. Science. American Association for the Advancement of Science. 2018;362:895–6.
Wilson A, Oser GM, Jaworski M, Blanco-Bose WE, Laurenti E, Adolphe C, et al. Dormant and self-renewing hematopoietic stem cells and their niches. In: Annals of the New York Academy of Sciences. Blackwell Publishing Inc.; 2007. p. 64–75.
Pietras EM, Reynaud D, Kang Y-A, Carlin D, Calero-Nieto FJ, Leavitt AD, et al. Functionally distinct subsets of lineage-biased multipotent progenitors control blood production in normal and regenerative conditions. Cell Stem Cell [Internet]. 2015 Jul 2 [cited 2017 Oct 15];17(1):35–46. Available from: http://www.ncbi.nlm.nih.gov/pubmed/26095048.
Kondo M, Weissman IL, Akashi K. Identification of clonogenic common lymphoid progenitors in mouse bone marrow. Cell [Internet]. 1997 Nov 28 [cited 2017 Oct 16];91(5):661–72. Available from: http://www.ncbi.nlm.nih.gov/pubmed/9393859.
Akashi K, Traver D, Miyamoto T, Weissman IL. A clonogenic common myeloid progenitor that gives rise to all myeloid lineages. Nature [Internet]. 2000 Mar 9 [cited 2017 Oct 16];404(6774):193–7. Available from: http://www.ncbi.nlm.nih.gov/pubmed/10724173.
Cabezas-Wallscheid N, Klimmeck D, Hansson J, Lipka DB, Reyes A, Wang Q, et al. Identification of regulatory networks in hscs and their immediate progeny via integrated proteome, transcriptome, and dna methylome analysis. Cell Stem Cell [Internet]. 2014 Oct 2 [cited 2017 Oct 15];15(4):507–22. Available from: http://www.ncbi.nlm.nih.gov/pubmed/25158935.
Wilson A, Laurenti E, Oser G, van der Wath RC, Blanco-Bose W, Jaworski M, et al. Hematopoietic stem cells reversibly switch from dormancy to self-renewal during homeostasis and repair. Cell [Internet]. 2008 Dec 12 [cited 2017 Oct 15];135(6):1118–29. Available from: http://www.ncbi.nlm.nih.gov/pubmed/19062086.
Shin JY, Hu W, Naramura M, Park CY. High c-Kit expression identifies hematopoietic stem cells with impaired self-renewal and megakaryocytic bias. J Exp Med [Internet]. 2014 Feb 10 [cited 2017 Oct 15];211(2):217–31. Available from: https://doi.org/10.1084/jem.20131128
Beerman I, Bhattacharya D, Zandi S, Sigvardsson M, Weissman IL, Bryder D, et al. Functionally distinct hematopoietic stem cells modulate hematopoietic lineage potential during aging by a mechanism of clonal expansion. Proc Natl Acad Sci [Internet]. 2010 Mar 23 [cited 2017 Oct 15];107(12):5465–70. Available from: http://www.ncbi.nlm.nih.gov/pubmed/20304793.
Adolfsson J, Månsson R, Buza-Vidas N, Hultquist A, Liuba K, Jensen CT, et al. Identification of Flt3+ lympho-myeloid stem cells lacking erythro-megakaryocytic potential. Cell [Internet]. 2005 Apr 22 [cited 2017 Oct 15];121(2):295–306. Available from: http://www.ncbi.nlm.nih.gov/pubmed/15851035.
Kiel MJ, Yilmaz ÖH, Iwashita T, Yilmaz OH, Terhorst C, Morrison SJ. SLAM Family receptors distinguish hematopoietic stem and progenitor cells and reveal endothelial niches for stem cells. Cell [Internet]. 2005 Jul 1 [cited 2017 Oct 15];121(7):1109–21. Available from: http://www.ncbi.nlm.nih.gov/pubmed/15989959.
Morita Y, Ema H, Nakauchi H. Heterogeneity and hierarchy within the most primitive hematopoietic stem cell compartment. J Exp Med [Internet]. 2010 Jun 7 [cited 2017 Oct 15];207(6):1173–82. Available from: http://www.ncbi.nlm.nih.gov/pubmed/20421392.
•• Yamamoto R, Morita Y, Ooehara J, Hamanaka S, Onodera M, Rudolph KL, et al. Clonal analysis unveils self-renewing lineage-restricted progenitors generated directly from hematopoietic stem cells. Cell [Internet]. 2013 Aug 29 [cited 2017 Oct 15];154(5):1112–26. Available from: http://www.ncbi.nlm.nih.gov/pubmed/23993099. This study utilized single-cell transplantations to demonstrate that lineage-restricted progenitors with high self-renewal capacity exist.
Notta F, Zandi S, Takayama N, Dobson S, Gan OI, Wilson G, et al. Distinct routes of lineage development reshape the human blood hierarchy across ontogeny. Science (80- ) [Internet]. 2016 Jan 8 [cited 2017 Oct 15];351(6269):aab2116–aab2116. Available from: http://www.ncbi.nlm.nih.gov/pubmed/26541609.
Perié L, Duffy KR, Kok L, de Boer RJ, Schumacher TN. The branching point in erythro-myeloid differentiation. Cell [Internet]. 2015 Dec 17 [cited 2017 Oct 15];163(7):1655–62. Available from: http://www.ncbi.nlm.nih.gov/pubmed/26687356.
Kowalczyk MS, Tirosh I, Heckl D, Rao TN, Dixit A, Haas BJ, et al. Single-cell RNA-seq reveals changes in cell cycle and differentiation programs upon aging of hematopoietic stem cells. Genome Res [Internet]. 2015 Dec [cited 2017 Oct 15];25(12):1860–72. Available from: http://www.ncbi.nlm.nih.gov/pubmed/26430063.
Guo G, Luc S, Marco E, Lin T-W, Peng C, Kerenyi MA, et al. Mapping cellular hierarchy by single-cell analysis of the cell surface repertoire. Cell Stem Cell [Internet]. 2013 Oct 3 [cited 2017 Oct 15];13(4):492–505. Available from: http://linkinghub.elsevier.com/retrieve/pii/S1934590913003627.
Muller-Sieburg CE, Cho RH, Karlsson L, Huang J-F, Sieburg HB. Myeloid-biased hematopoietic stem cells have extensive self-renewal capacity but generate diminished lymphoid progeny with impaired IL-7 responsiveness. Blood [Internet]. 2004 Jun 1 [cited 2017 Oct 15];103(11):4111–8. Available from: http://www.ncbi.nlm.nih.gov/pubmed/14976059.
Wilson NK, Kent DG, Buettner F, Shehata M, Macaulay IC, Calero-Nieto FJ, et al. Combined single-cell functional and gene expression analysis resolves heterogeneity within stem cell populations. Cell Stem Cell [Internet]. 2015 Jun 4 [cited 2017 Oct 15];16(6):712–24. Available from: http://www.ncbi.nlm.nih.gov/pubmed/26004780.
Dykstra B, Kent D, Bowie M, McCaffrey L, Hamilton M, Lyons K, et al. Long-term propagation of distinct hematopoietic differentiation programs in vivo. Cell Stem Cell [Internet]. 2007 Aug 16 [cited 2017 Oct 15];1(2):218–29. Available from: http://www.ncbi.nlm.nih.gov/pubmed/18371352.
Paul F, Arkin Y, Giladi A, Jaitin DA, Kenigsberg E, Keren-Shaul H, et al. Transcriptional heterogeneity and lineage commitment in myeloid progenitors. Cell [Internet]. 2015 Dec 17 [cited 2017 Oct 15];163(7):1663–77. Available from: http://www.ncbi.nlm.nih.gov/pubmed/26627738. This study utilized single-cell transcriptomics to demonstrate that the common myeloid progenitor (CMP) compartment predominantly consists of lineage-committed progenitors.
Haas S, Hansson J, Klimmeck D, Loeffler D, Velten L, Uckelmann H, et al. Inflammation-induced emergency megakaryopoiesis driven by hematopoietic stem cell-like megakaryocyte progenitors. Cell Stem Cell [Internet]. 2015 Oct 1 [cited 2018 Jan 3];17(4):422–34. Available from: http://www.ncbi.nlm.nih.gov/pubmed/26299573.
Giladi A, Paul F, Herzog Y, Lubling Y, Weiner A, Yofe I, et al. Single-cell characterization of haematopoietic progenitors and their trajectories in homeostasis and perturbed haematopoiesis. Nat Cell Biol. 2018;20(7):836–46.
Popescu D-M, Botting RA, Stephenson E, Green K, Webb S, Jardine L, et al. Decoding human fetal liver haematopoiesis. Nature [Internet]. 2019 [cited 2020 Feb 1];574(7778):365–71. Available from: http://www.ncbi.nlm.nih.gov/pubmed/31597962.
Tusi BK, Wolock SL, Weinreb C, Hwang Y, Hidalgo D, Zilionis R, et al. Population snapshots predict early haematopoietic and erythroid hierarchies. Nature. 2018;555(7694):54–60.
Pellin D, Loperfido M, Baricordi C, Wolock SL, Montepeloso A, Weinberg OK, et al. A comprehensive single cell transcriptional landscape of human hematopoietic progenitors. Nat Commun [Internet]. 2019 [cited 2020 Feb 1];10(1):2395. Available from: http://www.ncbi.nlm.nih.gov/pubmed/31160568.
Nestorowa S, Hamey FK, Pijuan Sala B, Diamanti E, Shepherd M, Laurenti E, et al. A single-cell resolution map of mouse hematopoietic stem and progenitor cell differentiation. Blood [Internet]. 2016 Aug 25 [cited 2017 Oct 15];128(8):e20–31. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27365425.
Velten L, Haas SF, Raffel S, Blaszkiewicz S, Islam S, Hennig BP, et al. Human haematopoietic stem cell lineage commitment is a continuous process. Nat Cell Biol [Internet]. 2017 mar 20 [cited 2017 Oct 15];19(4):271–81. Available from: http://www.ncbi.nlm.nih.gov/pubmed/28319093. This manuscript demonstrates that human hematopoisis is a continuous process.
Rodriguez-Fraticelli AE, Wolock SL, Weinreb CS, Panero R, Patel SH, Jankovic M, et al. Clonal analysis of lineage fate in native haematopoiesis. Nature [Internet]. 2018 Jan 3 [cited 2018 mar 9];553(7687):212–6. Available from: http://www.ncbi.nlm.nih.gov/pubmed/29323290. This manuscript reveals that megakaryocyte commitment is the predominant native fate outcome of HSCs.
Baccin C, Al-Sabah J, Velten L, Helbling PM, Grünschläger F, Hernández-Malmierca P, et al. Combined single-cell and spatial transcriptomics reveal the molecular, cellular and spatial bone marrow niche organization. Nat Cell Biol. 2020;22(1):38–48.
Baryawno N, Przybylski D, Kowalczyk MS, Kfoury Y, Severe N, Gustafsson K, et al. A cellular taxonomy of the bone marrow stroma in homeostasis and leukemia. Cell. 2019;177(7):1915–1932.e16.
Wolock SL, Krishnan I, Tenen DE, Matkins V, Camacho V, Patel S, et al. Mapping distinct bone marrow niche populations and their differentiation paths. Cell Rep. 2019;28(2):302–311.e5.
Tikhonova AN, Dolgalev I, Hu H, Sivaraj KK, Hoxha E, Cuesta-Domínguez Á, et al. The bone marrow microenvironment at single-cell resolution. Nature. 2019 May 9;569(7755):222–8.
Al-Sabah J, Baccin C, Haas S. Single-cell and spatial transcriptomics approaches of the bone marrow microenvironment. Curr Opin Oncol. 2019;1.
Velten L, Story BA, Hernandez-Malmierca P, Milbank J, Paulsen M, Lutz C, et al. MutaSeq reveals the transcriptomic consequences of clonal evolution in acute myeloid leukemia. bioRxiv. 2018;21:500108.
Giustacchini A, Thongjuea S, Barkas N, Woll PS, Povinelli BJ, Booth CAG, et al. Single-cell transcriptomics uncovers distinct molecular signatures of stem cells in chronic myeloid leukemia. Nat Med. 2017 Jun 1;23(6):692–702.
van Galen P, Hovestadt V, Wadsworth MH, Hughes TK, Griffin GK, Battaglia S, et al. Single-cell RNA-Seq reveals AML hierarchies relevant to disease progression and immunity. Cell. 2019;176(6):1265–1281.e24.
Nam AS, Kim KT, Chaligne R, Izzo F, Ang C, Taylor J, et al. Somatic mutations and cell identity linked by genotyping of Transcriptomes. Nature. 2019;571(7765):355–60.
Macaulay IC, Haerty W, Kumar P, Li YI, Hu TX, Teng MJ, et al. G&T-seq: parallel sequencing of single-cell genomes and transcriptomes. Nat Methods. 2015;12(6):519–22.
Rodriguez-Meira A, Buck G, Clark SA, Povinelli BJ, Alcolea V, Louka E, et al. Unravelling intratumoral heterogeneity through high-sensitivity single-cell mutational analysis and parallel RNA sequencing. Mol Cell. 2019;73(6):1292–1305.e8.
Granja JM, Klemm S, McGinnis LM, Kathiria AS, Mezger A, Corces MR, et al. Single-cell multiomic analysis identifies regulatory programs in mixed-phenotype acute leukemia. Nature Biotechnology.Nature Research. 2019;37:1458–65.
Doulatov S, Notta F, Laurenti E, Dick JE. Hematopoiesis: a human perspective. Cell Stem Cell [Internet]. 2012 Feb 3 [cited 2018 Mar 10];10(2):120–36. Available from: https://www.sciencedirect.com/science/article/pii/S1934590912000082#bib109.
Notta F, Doulatov S, Laurenti E, Poeppl A, Jurisica I, Dick JE. Isolation of single human hematopoietic stem cells capable of long-term multilineage engraftment. Science (80- ) [Internet]. 2011 Jul 8 [cited 2018 mar 10];333(6039):218–21. Available from: http://www.ncbi.nlm.nih.gov/pubmed/21737740.
Müller-Sieburg CE, Cho RH, Thoman M, Adkins B, Sieburg HB. Deterministic regulation of hematopoietic stem cell self-renewal and differentiation. Blood [Internet]. 2002 Aug 15 [cited 2017 Oct 15];100(4):1302–9. Available from: http://www.ncbi.nlm.nih.gov/pubmed/12149211.
Benz C, Copley MR, Kent DG, Wohrer S, Cortes A, Aghaeepour N, et al. Hematopoietic stem cell subtypes expand differentially during development and display distinct lymphopoietic programs. Cell Stem Cell [Internet]. 2012 Mar 2 [cited 2017 Oct 15];10(3):273–83. Available from: http://linkinghub.elsevier.com/retrieve/pii/S1934590912000653
Carrelha J, Meng Y, Kettyle LM, Luis TC, Norfo R, Alcolea V, et al. Hierarchically related lineage-restricted fates of multipotent haematopoietic stem cells. Nature. 2018;554(7690):106–11.
Karamitros D, Stoilova B, Aboukhalil Z, Hamey F, Reinisch A, Samitsch M, et al. Single-cell analysis reveals the continuum of human lympho-myeloid progenitor cells. Nat Immunol [Internet]. 2018 Jan 21 [cited 2018 mar 9];19(1):85–97. Available from: http://www.ncbi.nlm.nih.gov/pubmed/29167569.
Naik SH, Perié L, Swart E, Gerlach C, van Rooij N, de Boer RJ, et al. Diverse and heritable lineage imprinting of early haematopoietic progenitors. Nature [Internet]. 2013 Apr 3 [cited 2017 Oct 15];496(7444):229–32. Available from: http://www.ncbi.nlm.nih.gov/pubmed/23552896.
Weinreb C, Rodriguez-Fraticelli A, Camargo FD, Klein AM. Lineage tracing on transcriptional landscapes links state to fate during differentiation. Science [Internet]. 2020 Jan 23 [cited 2020 Feb 1]; Available from: http://www.ncbi.nlm.nih.gov/pubmed/31974159.
Challen GA, Boles NC, Chambers SM, Goodell MA. Distinct hematopoietic stem cell subtypes are differentially regulated by TGF-β1. Cell Stem Cell [Internet]. 2010 Mar 5 [cited 2017 Oct 15];6(3):265–78. Available from: http://www.ncbi.nlm.nih.gov/pubmed/20207229.
Grover A, Sanjuan-Pla A, Thongjuea S, Carrelha J, Giustacchini A, Gambardella A, et al. Single-cell RNA sequencing reveals molecular and functional platelet bias of aged haematopoietic stem cells. Nat Commun [Internet]. 2016 Mar 24 [cited 2017 Oct 15];7:11075. Available from: https://doi.org/10.1038/ncomms11075
Månsson R, Hultquist A, Luc S, Yang L, Anderson K, Kharazi S, et al. Molecular evidence for hierarchical transcriptional lineage priming in fetal and adult stem cells and multipotent progenitors. Immunity [Internet]. 2007 Apr [cited 2017 Oct 15];26(4):407–19. Available from: http://www.ncbi.nlm.nih.gov/pubmed/17433729.
Sanjuan-Pla A, Macaulay IC, Jensen CT, Woll PS, Luis TC, Mead A, et al. Platelet-biased stem cells reside at the apex of the haematopoietic stem-cell hierarchy. Nature [Internet]. 2013 Aug 11 [cited 2017 Oct 15];502(7470):232–6. Available from: http://www.ncbi.nlm.nih.gov/pubmed/23934107. This study exploited lineage-tracing and single-cell approaches to reveal the existence of platelet-biased HSCs.
Buenrostro JD, Corces MR, Lareau CA, Wu B, Schep AN, Aryee MJ, et al. Integrated single-cell analysis maps the continuous regulatory landscape of human hematopoietic differentiation. Cell. 2018;173(6):1535–1548.e16.
Beerman I, Rossi DJ. Epigenetic control of stem cell potential during homeostasis, aging, and disease. Cell Stem Cell [Internet]. 2015 Jun [cited 2017 Oct 15];16(6):613–25. Available from: http://linkinghub.elsevier.com/retrieve/pii/S1934590915002180
Farlik M, Halbritter F, Müller F, Choudry FA, Ebert P, Klughammer J, et al. DNA methylation dynamics of human hematopoietic stem cell differentiation. Cell Stem Cell [Internet]. 2016 Dec 1 [cited 2017 Oct 15];19(6):808–22. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27867036.
Lipka DB, Wang Q, Cabezas-Wallscheid N, Klimmeck D, Weichenhan D, Herrmann C, et al. Identification of DNA methylation changes at cis-regulatory elements during early steps of HSC differentiation using tagmentation-based whole genome bisulfite sequencing. Cell Cycle [Internet]. 2014 [cited 2017 Oct 15];13(22):3476–87. Available from: http://www.ncbi.nlm.nih.gov/pubmed/25483069.
Sun D, Luo M, Jeong M, Rodriguez B, Xia Z, Hannah R, et al. Epigenomic profiling of young and aged HSCs reveals concerted changes during aging that reinforce self-renewal. Cell Stem Cell [Internet]. 2014 May 1 [cited 2017 Oct 15];14(5):673–88. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24792119.
Yu VWC, Yusuf RZ, Oki T, Wu J, Saez B, Wang X, et al. Epigenetic memory underlies cell-autonomous heterogeneous behavior of hematopoietic stem cells. Cell [Internet]. 2016 Nov 17 [cited 2017 Oct 15];167(5):1310-1322.e17. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27863245.
Clark SJ, Argelaguet R, Kapourani CA, Stubbs TM, Lee HJ, Alda-Catalinas C, et al. ScNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells e. Nat Commun. 2018;1:9(1).
Filippi MD, Ghaffari S. Mitochondria in the maintenance of hematopoietic stem cells: new perspectives and opportunities. Blood. American Society of Hematology. 2019;133:1943–52.
Bernitz JM, Kim HS, MacArthur B, Sieburg H, Moore K. Hematopoietic stem cells count and remember self-renewal divisions. Cell [Internet]. 2016 Nov 17 [cited 2017 Oct 15];167(5):1296-1309.e10. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27839867.
Cabezas-Wallscheid N, Buettner F, Sommerkamp P, Klimmeck D, Ladel L, Thalheimer FB, et al. Vitamin A-retinoic acid signaling regulates hematopoietic stem cell dormancy. Cell [Internet]. 2017 May 18 [cited 2017 Oct 15];169(5):807–823.e19. Available from: http://linkinghub.elsevier.com/retrieve/pii/S0092867417304646.
Ho TT, Warr MR, Adelman ER, Lansinger OM, Flach J, Verovskaya E V, et al. Autophagy maintains the metabolism and function of young and old stem cells. Nature [Internet]. 2017 Mar 9 [cited 2017 Oct 16];543(7644):205–10. Available from: https://doi.org/10.1038/nature21388
Warr MR, Kohli L, Passegué E. Born to survive: autophagy in hematopoietic stem cell maintenance. Cell Cycle [Internet]. 2013 Jul 1 [cited 2017 Oct 16];12(13):1979–80. Available from: http://www.ncbi.nlm.nih.gov/pubmed/23759583.
Essers MAG, Offner S, Blanco-Bose WE, Waibler Z, Kalinke U, Duchosal MA, et al. IFNα activates dormant haematopoietic stem cells in vivo. Nature [Internet]. 2009 Apr 16 [cited 2018 Jan 3];458(7240):904–8. Available from: http://www.ncbi.nlm.nih.gov/pubmed/19212321.
Baldridge MT, King KY, Boles NC, Weksberg DC, Goodell MA. Quiescent haematopoietic stem cells are activated by IFN-γ in response to chronic infection. Nature [Internet]. 2010 Jun 10 [cited 2018 Jan 3];465(7299):793–7. Available from: http://www.ncbi.nlm.nih.gov/pubmed/20535209.
King KY, Goodell MA. Inflammatory modulation of HSCs: viewing the HSC as a foundation for the immune response. Nat Rev Immunol [Internet]. 2011 Sep 9 [cited 2017 Oct 15];11(10):685–92. Available from: http://www.ncbi.nlm.nih.gov/pubmed/21904387.
Hirche C, Frenz T, Haas SF, Döring M, Borst K, Tegtmeyer P-K, et al. Systemic Virus Infections Differentially Modulate Cell Cycle State and Functionality of long-term hematopoietic stem cells in vivo. Cell Rep [Internet]. 2017 Jun 13 [cited 2018 Jan 3];19(11):2345–56. Available from: http://www.ncbi.nlm.nih.gov/pubmed/28614719.
Macaulay IC, Svensson V, Labalette C, Ferreira L, Hamey F, Voet T, et al. Single-cell RNA-sequencing reveals a continuous spectrum of differentiation in hematopoietic cells. Cell Rep [Internet]. 2016 Feb 2 [cited 2017 Oct 16];14(4):966–77. Available from: http://www.ncbi.nlm.nih.gov/pubmed/26804912.
Zheng S, Papalexi E, Butler A, Stephenson W, Satija R. Molecular transitions in early progenitors during human cord blood hematopoiesis. Mol Syst Biol. 2018;14(3).
Drissen R, Buza-Vidas N, Woll P, Thongjuea S, Gambardella A, Giustacchini A, et al. Distinct myeloid progenitor-differentiation pathways identified through single-cell RNA sequencing. Nat Immunol [Internet]. 2016 Jun [cited 2017 Oct 16];17(6):666–76. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27043410.
Olsson A, Venkatasubramanian M, Chaudhri VK, Aronow BJ, Salomonis N, Singh H, et al. Single-cell analysis of mixed-lineage states leading to a binary cell fate choice. Nature [Internet]. 2016 Aug 31 [cited 2017 Oct 15];537(7622):698–702. Available from: http://www.ncbi.nlm.nih.gov/pubmed/27580035.
Doulatov S, Notta F, Eppert K, Nguyen LT, Ohashi PS, Dick JE. Revised map of the human progenitor hierarchy shows the origin of macrophages and dendritic cells in early lymphoid development. Nat Immunol [Internet]. 2010 Jul 13 [cited 2017 Oct 16];11(7):585–93. Available from: https://doi.org/10.1038/ni.1889
Görgens A, Radtke S, Möllmann M, Cross M, Dürig J, Horn PA, et al. Revision of the human hematopoietic tree: granulocyte subtypes derive from distinct hematopoietic lineages. Cell Rep [Internet]. 2013 May [cited 2017 Oct 16];3(5):1539–52. Available from: http://linkinghub.elsevier.com/retrieve/pii/S2211124713002076
Drissen R, Thongjuea S, Theilgaard-Mönch K, Nerlov C. Identification of two distinct pathways of human myelopoiesis. Sci Immunol. 2019;4(35).
Huang H, Cantor AB. Common features of megakaryocytes and hematopoietic stem cells: what’s the connection? Journal of Cellular Biochemistry. 2009;107:857–64.
Petti AA, Williams SR, Miller CA, Fiddes IT, Srivatsan SN, Chen DY, et al. A general approach for detecting expressed mutations in AML cells using single cell RNA-sequencing. Nat Commun. 2019;10(1).
Nangalia J, Massie CE, Baxter EJ, Nice FL, Gundem G, Wedge DC, et al. Somatic CALR mutations in myeloproliferative neoplasms with nonmutated JAK2. N Engl J Med [Internet]. 2013 Dec 19 [cited 2020 Feb 1];369(25):2391–405. Available from: https://doi.org/10.1056/NEJMoa1312542
Ludwig LS, Lareau CA, Ulirsch JC, Christian E, Muus C, Li LH, et al. Lineage tracing in humans enabled by mitochondrial mutations and single-cell genomics. Cell. 2019 Mar 7;176(6):1325–1339.e22.
Mendelson A, Frenette PS. Hematopoietic stem cell niche maintenance during homeostasis and regeneration. Nat Med [Internet]. 2014 Aug 6 [cited 2017 Oct 15];20(8):833–46. Available from: http://www.ncbi.nlm.nih.gov/pubmed/25100529.
Morrison SJ, Scadden DT. The bone marrow niche for haematopoietic stem cells. Nature [Internet]. 2014 Jan 15 [cited 2017 Oct 15];505(7483):327–34. Available from: http://www.ncbi.nlm.nih.gov/pubmed/24429631.
Zhou BO, Yue R, Murphy MM, Peyer JG, Morrison SJ. Leptin-receptor-expressing mesenchymal stromal cells represent the main source of bone formed by adult bone marrow. Cell Stem Cell. 2014;15(2):154–68.
Kunisaki Y, Bruns I, Scheiermann C, Ahmed J, Pinho S, Zhang D, et al. Arteriolar niches maintain haematopoietic stem cell quiescence. Nature. 2013;502(7473):637–43.
Asada N, Kunisaki Y, Pierce H, Wang Z, Fernandez NF, Birbrair A, et al. Differential cytokine contributions of perivascular haematopoietic stem cell niches. Nat Cell Biol. 2017;19(3):214–23.
Méndez-Ferrer S, Michurina TV, Ferraro F, Mazloom AR, MacArthur BD, Lira SA, et al. Mesenchymal and haematopoietic stem cells form a unique bone marrow niche. Nature. 2010 Aug 12;466(7308):829–34.
Ding L, Morrison SJ. Haematopoietic stem cells and early lymphoid progenitors occupy distinct bone marrow niches. Nature [Internet]. 2013 Mar 14 [cited 2020 Feb 1];495(7440):231–5. Available from: http://www.ncbi.nlm.nih.gov/pubmed/23434755.
Ding L, Saunders TL, Enikolopov G, Morrison SJ. Endothelial and perivascular cells maintain haematopoietic stem cells. Nature. 2012;481(7382):457–62.
Sugiyama T, Kohara H, Noda M, Nagasawa T. Maintenance of the hematopoietic stem cell pool by CXCL12-CXCR4 chemokine signaling in bone marrow stromal cell niches. Immunity [Internet]. 2006 Dec [cited 2017 Oct 15];25(6):977–88. Available from: http://www.ncbi.nlm.nih.gov/pubmed/17174120.
Azizi E, Carr AJ, Plitas G, Cornish AE, Konopacki C, Prabhakaran S, et al. Single-cell map of diverse immune phenotypes in the breast tumor microenvironment. Cell. 2018;174(5):1293–1308.e36.
Stoeckius M, Hafemeister C, Stephenson W, Houck-Loomis B, Chattopadhyay PK, Swerdlow H, et al. Simultaneous epitope and transcriptome measurement in single cells. Nat Methods. 2017;14(9):865–8.
Shahi P, Kim SC, Haliburton JR, Gartner ZJ, Abate AR. Abseq: ultrahigh-throughput single cell protein profiling with droplet microfluidic barcoding. Sci Rep. 2017;14:7.
Jaitin DA, Weiner A, Yofe I, Lara-Astiaso D, Keren-Shaul H, David E, et al. Dissecting immune circuits by linking CRISPR-pooled screens with single-cell RNA-seq. Cell. 2016;167(7):1883–1896.e15.
Yao J, Dai HL. Is pooled CRISPR-screening the dawn of a new era for functional genomics. In: Advances in Experimental Medicine and Biology. Springer New York LLC; 2018. p. 171–6.
Dixit A, Parnas O, Li B, Chen J, Fulco CP, Jerby-Arnon L, et al. Perturb-seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell. 2016;167(7):1853–1866.e17.
Gundry MC, Dever DP, Yudovich D, Bauer DE, Haas S, Wilkinson AC, et al. Technical considerations for the use of CRISPR/Cas9 in hematology research. Experimental Hematology. Elsevier Inc. 2017;54:4–11.
Acknowledgments
I would like to thank Jude Al-Sabah for critically proofreading the manuscript.
Funding
Open Access funding provided by Projekt DEAL.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
None.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Cancer and Stem Cells
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Haas, S. Hematopoietic Stem Cells in Health and Disease—Insights from Single-Cell Multi-omic Approaches. Curr Stem Cell Rep 6, 67–76 (2020). https://doi.org/10.1007/s40778-020-00174-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40778-020-00174-2