Abstract
Purpose of Review
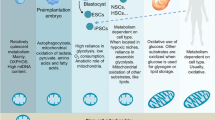
Metabolism is increasingly recognized as a major player in control of stem cell function and fate. How stem cell metabolism is established, maintained, and regulated is a fundamental question of biology and medicine. In this review, we discuss major metabolic programs in stem cells and cancer stem cells, with a focus on key transcription factors that shape the stem cell metabolic phenotype.
Recent Findings
Cancer stem cells primarily use oxidative phosphorylation for energy generation, in contrast to normal stem cells, which rely on glycolytic metabolism with the exception of mouse embryonic stem cells. Transcription factors control the metabolic phenotype of stem cells by modulating the expression of enzymes and thus the activity of metabolic pathways. It is evident that HIF1α and PGC1α function as master regulators of glycolytic and mitochondrial metabolism, respectively.
Summary
Transcriptional regulation is a key mechanism for establishing specific metabolic programs in stem cells and cancer stem cells.

Similar content being viewed by others
References
Papers of particular interest, published recently, have been highlighted as: • Of importance,•• Of major importance
De Los AA, Ferrari F, Xi R, Fujiwara Y, Benvenisty N, Deng H, et al. Hallmarks of pluripotency. Nature. 2015;525(7570):469–78. doi:10.1038/nature15515.
Weissman IL, Anderson DJ, Gage F. Stem and progenitor cells: origins, phenotypes, lineage commitments, and transdifferentiations. Annu Rev Cell Dev Biol. 2001;17:387–403. doi:10.1146/annurev.cellbio.17.1.387.
Thomson JA, Itskovitz-Eldor J, Shapiro SS, Waknitz MA, Swiergiel JJ, Marshall VS, et al. Embryonic stem cell lines derived from human blastocysts. Science. 1998;282(5391):1145–7.
Evans MJ, Kaufman MH. Establishment in culture of pluripotential cells from mouse embryos. Nature. 1981;292(5819):154–6.
Martin GR. Isolation of a pluripotent cell line from early mouse embryos cultured in medium conditioned by teratocarcinoma stem cells. Proc Natl Acad Sci U S A. 1981;78(12):7634–8.
Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126(4):663–76. doi:10.1016/j.cell.2006.07.024.
Takahashi K, Tanabe K, Ohnuki M, Narita M, Ichisaka T, Tomoda K, et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 2007;131(5):861–72. doi:10.1016/j.cell.2007.11.019.
Copley MR, Beer PA, Eaves CJ. Hematopoietic stem cell heterogeneity takes center stage. Cell Stem Cell. 2012;10(6):690–7. doi:10.1016/j.stem.2012.05.006.
Bond AM, Ming GL, Song H. Adult mammalian neural stem cells and neurogenesis: five decades later. Cell Stem Cell. 2015;17(4):385–95. doi:10.1016/j.stem.2015.09.003.
Fialkow PJ. Clonal origin of human tumors. Biochim Biophys Acta. 1976;458(3):283–321.
Tannock IF. Principles of cell proliferation: cell kinetics. In: DeVita VT, Hellman S, Rosenberg SA, editors. Cancer: principles and practice of oncology. 3rd ed. Philadelphia: J.B. Lipincott; 1989. p. 3–13.
Heppner GH. Tumor heterogeneity. Cancer Res. 1984;44(6):2259–65.
Reya T, Morrison SJ, Clarke MF, Weissman IL. Stem cells, cancer, and cancer stem cells. Nature. 2001;414(6859):105–11.
Meacham CE, Morrison SJ. Tumour heterogeneity and cancer cell plasticity. Nature. 2013;501(7467):328–37. doi:10.1038/nature12624.
Lapidot T, Sirard C, Vormoor J, Murdoch B, Hoang T, Caceres-Cortes J, et al. A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature. 1994;367(6464):645–8. doi:10.1038/367645a0.
Wang JC, Lapidot T, Cashman JD, Doedens M, Addy L, Sutherland DR, et al. High level engraftment of NOD/SCID mice by primitive normal and leukemic hematopoietic cells from patients with chronic myeloid leukemia in chronic phase. Blood. 1998;91(7):2406–14.
Al-Hajj M, Wicha MS, Benito-Hernandez A, Morrison SJ, Clarke MF. Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci U S A. 2003;100(7):3983–8. doi:10.1073/pnas.0530291100.
Singh SK, Hawkins C, Clarke ID, Squire JA, Bayani J, Hide T, et al. Identification of human brain tumour initiating cells. Nature. 2004;432(7015):396–401.
Dalerba P, Dylla SJ, Park IK, Liu R, Wang X, Cho RW, et al. Phenotypic characterization of human colorectal cancer stem cells. Proc Natl Acad Sci U S A. 2007;104(24):10158–63. doi:10.1073/pnas.0703478104.
O'Brien CA, Pollett A, Gallinger S, Dick JE. A human colon cancer cell capable of initiating tumour growth in immunodeficient mice. Nature. 2007;445(7123):106–10. doi:10.1038/nature05372.
Ricci-Vitiani L, Lombardi DG, Pilozzi E, Biffoni M, Todaro M, Peschle C, et al. Identification and expansion of human colon-cancer-initiating cells. Nature. 2007;445(7123):111–5. doi:10.1038/nature05384.
Li C, Heidt DG, Dalerba P, Burant CF, Zhang L, Adsay V, et al. Identification of pancreatic cancer stem cells. Cancer Res. 2007;67(3):1030–7. doi:10.1158/0008-5472.CAN-06-2030.
Curley MD, Therrien VA, Cummings CL, Sergent PA, Koulouris CR, Friel AM, et al. CD133 expression defines a tumor initiating cell population in primary human ovarian cancer. Stem Cells. 2009;27(12):2875–83. doi:10.1002/stem.236.
Zhang S, Balch C, Chan MW, Lai HC, Matei D, Schilder JM, et al. Identification and characterization of ovarian cancer-initiating cells from primary human tumors. Cancer Res. 2008;68(11):4311–20. doi:10.1158/0008-5472.CAN-08-0364.
Kleinsmith LJ, Pierce Jr GB. Multipotentiality of single embryonal carcinoma cells. Cancer Res. 1964;24:1544–51.
Martin GR. Teratocarcinomas and mammalian embryogenesis. Science. 1980;209(4458):768–76.
Nichols J, Smith A. Naive and primed pluripotent states. Cell Stem Cell. 2009;4(6):487–92. doi:10.1016/j.stem.2009.05.015.
• Ito K, Suda T. Metabolic requirements for the maintenance of self-renewing stem cells. Nat Rev Mol Cell Biol. 2014;15(4):243–56. doi:10.1038/nrm3772. This review provides a thorough overview of metabolic pathways for the maintenance of adult tissue stem cells.
Chandel NS, Jasper H, Ho TT, Passegue E. Metabolic regulation of stem cell function in tissue homeostasis and organismal ageing. Nat Cell Biol. 2016;18(8):823–32. doi:10.1038/ncb3385.
Shyh-Chang N, Daley GQ, Cantley LC. Stem cell metabolism in tissue development and aging. Development. 2013;140(12):2535–47. doi:10.1242/dev.091777.
Wu J, Ocampo A, Izpisua Belmonte JC. Cellular metabolism and induced pluripotency. Cell. 2016;166(6):1371–85. doi:10.1016/j.cell.2016.08.008.
• Peiris-Pagès M, Martinez-Outschoorn UE, Pestell RG, Sotgia F, Lisanti MP. Cancer stem cell metabolism. Breast Cancer Res. 2016;18(1):1–10. doi:10.1186/s13058-016-0712-6. This review provides the most updated information on metabolic phenotypes of cancer stem cells.
Viale A, Draetta GF. Sugar? No thank you, just a deep breath of oxygen for cancer stem cells. Cell Metab. 2015;22(4):543–5. doi:10.1016/j.cmet.2015.09.020.
O'Neill LAJ, Kishton RJ, Rathmell J. A guide to immunometabolism for immunologists. Nat Rev Immunol. 2016;16(9):553–65. doi:10.1038/nri.2016.70.
Warburg O. On respiratory impairment in cancer cells. Science. 1956;124:269–70.
MacIver NJ, Michalek RD, Rathmell JC. Metabolic regulation of T lymphocytes. Annu Rev Immunol. 2013;31:259–83. doi:10.1146/annurev-immunol-032712-095956.
Agathocleous M, Harris WA. Metabolism in physiological cell proliferation and differentiation. Trends Cell Biol. 2013;23(10):484–92. doi:10.1016/j.tcb.2013.05.004.
Zhou W, Choi M, Margineantu D, Margaretha L, Hesson J, Cavanaugh C, et al. HIF1α induced switch from bivalent to exclusively glycolytic metabolism during ESC-to-EpiSC/hESC transition. EMBO J. 2012;31(9):2103–16. doi:10.1038/emboj.2012.71.
Tesar PJ, Chenoweth JG, Brook FA, Davies TJ, Evans EP, Mack DL, et al. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature. 2007;448(7150):196–9. doi:10.1038/nature05972.
Zhang J, Khvorostov I, Hong JS, Oktay Y, Vergnes L, Nuebel E, et al. UCP2 regulates energy metabolism and differentiation potential of human pluripotent stem cells. EMBO J. 2011;30(24):4860–73. doi:10.1038/emboj.2011.401.
Ware CB, Nelson AM, Mecham B, Hesson J, Zhou W, Jonlin EC, et al. Derivation of naive human embryonic stem cells. Proc Natl Acad Sci U S A. 2014;111(12):4484–9. doi:10.1073/pnas.1319738111.
Takashima Y, Guo G, Loos R, Nichols J, Ficz G, Krueger F, et al. Resetting transcription factor control circuitry toward ground-state pluripotency in human. Cell. 2014;158(6):1254–69. doi:10.1016/j.cell.2014.08.029.
Huang K, Maruyama T, Fan G. The naive state of human pluripotent stem cells: a synthesis of stem cell and preimplantation embryo transcriptome analyses. Cell Stem Cell. 2014;15(4):410–5. doi:10.1016/j.stem.2014.09.014.
Folmes CD, Nelson TJ, Martinez-Fernandez A, Arrell DK, Lindor JZ, Dzeja PP, et al. Somatic oxidative bioenergetics transitions into pluripotency-dependent glycolysis to facilitate nuclear reprogramming. Cell Metab. 2011;14(2):264–71. doi:10.1016/j.cmet.2011.06.011.
Panopoulos AD, Yanes O, Ruiz S, Kida YS, Diep D, Tautenhahn R, et al. The metabolome of induced pluripotent stem cells reveals metabolic changes occurring in somatic cell reprogramming. Cell Res. 2012;22(1):168–77. doi:10.1038/cr.2011.177.
Hawkins KE, Joy S, Delhove JM, Kotiadis VN, Fernandez E, Fitzpatrick LM, et al. NRF2 orchestrates the metabolic shift during induced pluripotent stem cell reprogramming. Cell Rep. 2016;14(8):1883–91. doi:10.1016/j.celrep.2016.02.003.
Kida Yasuyuki S, Kawamura T, Wei Z, Sogo T, Jacinto S, Shigeno A, et al. ERRs mediate a metabolic switch required for somatic cell reprogramming to pluripotency. Cell Stem Cell. 2015;16(5):547–55. doi:10.1016/j.stem.2015.03.001.
Cheung TH, Rando TA. Molecular regulation of stem cell quiescence. Nat Rev Mol Cell Biol. 2013;14(6):329–40. doi:10.1038/nrm3591.
Cavallucci V, Fidaleo M, Pani G. Neural stem cells and nutrients: poised between quiescence and exhaustion. Trends Endocrinol Metab. 2016; doi:10.1016/j.tem.2016.06.007.
DeBerardinis RJ, Lum JJ, Hatzivassiliou G, Thompson CB. The biology of cancer: metabolic reprogramming fuels cell growth and proliferation. Cell Metab. 2008;7(1):11–20. doi:10.1016/j.cmet.2007.10.002.
Vander Heiden MG, Cantley LC, Thompson CB. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science. 2009;324(5930):1029–33. doi:10.1126/science.1160809.
Cairns RA, Harris IS, Mak TW. Regulation of cancer cell metabolism. Nat Rev Cancer. 2011;11(2):85–95. doi:10.1038/nrc2981.
• Pavlova NN, Thompson CB. The emerging hallmarks of cancer metabolism. Cell Metab. 2016;23(1):27–47. doi:10.1016/j.cmet.2015.12.006. This comprehensive review catalogs cancer-related metabolic changes with disscussion of their contributions to tumorigenesis.
Ciavardelli D, Rossi C, Barcaroli D, Volpe S, Consalvo A, Zucchelli M, et al. Breast cancer stem cells rely on fermentative glycolysis and are sensitive to 2-deoxyglucose treatment. Cell Death Dis. 2014;5:e1336. doi:10.1038/cddis.2014.285.
Liao J, Qian F, Tchabo N, Mhawech-Fauceglia P, Beck A, Qian Z, et al. Ovarian cancer spheroid cells with stem cell-like properties contribute to tumor generation, metastasis and chemotherapy resistance through hypoxia-resistant metabolism. PLoS One. 2014;9(1):e84941. doi:10.1371/journal.pone.0084941.
Emmink BL, Verheem A, Van Houdt WJ, Steller EJA, Govaert KM, Pham TV, et al. The secretome of colon cancer stem cells contains drug-metabolizing enzymes. J Proteome. 2013;91:84–96. doi:10.1016/j.jprot.2013.06.027.
Viale A, Corti D, Draetta GF. Tumors and mitochondrial respiration: a neglected connection. Cancer Res. 2015;75(18):3687–91. doi:10.1158/0008-5472.can-15-0491.
Vlashi E, Pajonk F. The metabolic state of cancer stem cells—a valid target for cancer therapy? Free Radic Biol Med. 2015;79:264–8. doi:10.1016/j.freeradbiomed.2014.10.732.
Dando I, Dalla Pozza E, Biondani G, Cordani M, Palmieri M, Donadelli M. The metabolic landscape of cancer stem cells. IUBMB Life. 2015;67(9):687–93. doi:10.1002/iub.1426.
Janiszewska M, Suvà ML, Riggi N, Houtkooper RH, Auwerx J, Clément-Schatlo V, et al. Imp2 controls oxidative phosphorylation and is crucial for preserving glioblastoma cancer stem cells. Genes Dev. 2012;26(17):1926–44. doi:10.1101/gad.188292.112.
Vlashi E, Lagadec C, Vergnes L, Matsutani T, Masui K, Poulou M, et al. Metabolic state of glioma stem cells and nontumorigenic cells. Proc Natl Acad Sci. 2011;108(38):16062–7. doi:10.1073/pnas.1106704108.
Lagadinou Eleni D, Sach A, Callahan K, Rossi Randall M, Neering Sarah J, Minhajuddin M, et al. BCL-2 inhibition targets oxidative phosphorylation and selectively eradicates quiescent human leukemia stem cells. Cell Stem Cell. 2013;12(3):329–41. doi:10.1016/j.stem.2012.12.013.
Pasto A, Bellio C, Pilotto G, Ciminale V, Silic-Benussi M, Guzzo G, et al. Cancer stem cells from epithelial ovarian cancer patients privilege oxidative phosphorylation, and resist glucose deprivation. Oncotarget. 2014;5(12):4305–19. doi:10.18632/oncotarget.2010.
Farnie G, Sotgia F, Lisanti MP. High mitochondrial mass identifies a sub-population of stem-like cancer cells that are chemo-resistant. Oncotarget. 2015;6(31):30472–86. doi:10.18632/oncotarget.5401.
Wolf DA. Is reliance on mitochondrial respiration a “chink in the armor” of therapy-resistant cancer? Cancer Cell. 2014;26(6):788–95. doi:10.1016/j.ccell.2014.10.001.
Roesch A, Fukunaga-Kalabis M, Schmidt EC, Zabierowski SE, Brafford PA, Vultur A, et al. A temporarily distinct subpopulation of slow-cycling melanoma cells is required for continuous tumor growth. Cell. 2010;141(4):583–94. doi:10.1016/j.cell.2010.04.020.
Roesch A, Vultur A, Bogeski I, Wang H, Zimmermann Katharina M, Speicher D, et al. Overcoming intrinsic multidrug resistance in melanoma by blocking the mitochondrial respiratory chain of slow-cycling JARID1B high cells. Cancer Cell. 2013;23(6):811–25. doi:10.1016/j.ccr.2013.05.003.
•• Viale A, Pettazzoni P, Lyssiotis CA, Ying H, Sanchez N, Marchesini M, et al. Oncogene ablation-resistant pancreatic cancer cells depend on mitochondrial function. Nature. 2014;514(7524):628–32. doi:10.1038/nature13611. This study provides experimental evidence for targeting oxidative phosphorylation as a therapeutic strategy for eliminating pancreatic cancer stem cells
•• Sancho P, Burgos-Ramos E, Tavera A, Bou Kheir T, Jagust P, Schoenhals M, et al. MYC/PGC-1α balance determines the metabolic phenotype and plasticity of pancreatic cancer stem cells. Cell Metab. 2015;22(4):590–605. doi:10.1016/j.cmet.2015.08.015. This study describes a molecular mechanism for the metabolic plasticity of pancreatic cancer stem cells.
Semenza GL. Hypoxia-inducible factors in physiology and medicine. Cell. 2012;148(3):399–408. doi:10.1016/j.cell.2012.01.021.
Palomäki S, Pietilä M, Laitinen S, Pesälä J, Sormunen R, Lehenkari P, et al. HIF-1α is upregulated in human mesenchymal stem cells. Stem Cells. 2013;31(9):1902–9. doi:10.1002/stem.1435.
Suda T, Takubo K, Semenza GL. Metabolic regulation of hematopoietic stem cells in the hypoxic niche. Cell Stem Cell. 2011;9(4):298–310. doi:10.1016/j.stem.2011.09.010.
Hu CJ, Wang LY, Chodosh LA, Keith B, Simon MC. Differential roles of hypoxia-inducible factor 1alpha (HIF-1alpha) and HIF-2alpha in hypoxic gene regulation. Mol Cell Biol. 2003;23(24):9361–74.
Gordan JD, Thompson CB, Simon MC. HIF and c-Myc: sibling rivals for control of cancer cell metabolism and proliferation. Cancer Cell. 2007;12(2):108–13. doi:10.1016/j.ccr.2007.07.006.
Teslaa T, Teitell MA. Pluripotent stem cell energy metabolism: an update. EMBO J. 2015;34(2):138–53. doi:10.15252/embj.201490446.
Takubo K, Goda N, Yamada W, Iriuchishima H, Ikeda E, Kubota Y, et al. Regulation of the HIF-1alpha level is essential for hematopoietic stem cells. Cell Stem Cell. 2010;7(3):391–402. doi:10.1016/j.stem.2010.06.020.
Takubo K, Nagamatsu G, Kobayashi CI, Nakamura-Ishizu A, Kobayashi H, Ikeda E, et al. Regulation of glycolysis by Pdk functions as a metabolic checkpoint for cell cycle quiescence in hematopoietic stem cells. Cell Stem Cell. 2013;12(1):49–61. doi:10.1016/j.stem.2012.10.011.
Mathieu J, Zhou W, Xing Y, Sperber H, Ferreccio A, Agoston Z, et al. Hypoxia-inducible factors have distinct and stage-specific roles during reprogramming of human cells to pluripotency. Cell Stem Cell. 2014;14(5):592–605. doi:10.1016/j.stem.2014.02.012.
Prigione A, Rohwer N, Hoffmann S, Mlody B, Drews K, Bukowiecki R, et al. HIF1alpha modulates cell fate reprogramming through early glycolytic shift and upregulation of PDK1-3 and PKM2. Stem Cells. 2014;32(2):364–76. doi:10.1002/stem.1552.
Argiropoulos B, Yung E, Humphries RK. Unraveling the crucial roles of Meis1 in leukemogenesis and normal hematopoiesis. Genes Dev. 2007;21(22):2845–9. doi:10.1101/gad.1619407.
Simsek T, Kocabas F, Zheng J, Deberardinis RJ, Mahmoud AI, Olson EN, et al. The distinct metabolic profile of hematopoietic stem cells reflects their location in a hypoxic niche. Cell Stem Cell. 2010;7(3):380–90. doi:10.1016/j.stem.2010.07.011.
Kocabas F, Zheng J, Thet S, Copeland NG, Jenkins NA, DeBerardinis RJ, et al. Meis1 regulates the metabolic phenotype and oxidant defense of hematopoietic stem cells. Blood. 2012;120(25):4963–72. doi:10.1182/blood-2012-05-432260.
Dang CV. MYC on the path to cancer. Cell. 2012;149(1):22–35. doi:10.1016/j.cell.2012.03.003.
Dang CV. MYC, metabolism, cell growth, and tumorigenesis. Cold Spring Harbor perspectives in medicine. 2013;3(8):a014217. doi:10.1101/cshperspect.a014217.
Kim JW, Zeller KI, Wang Y, Jegga AG, Aronow BJ, O'Donnell KA, et al. Evaluation of myc E-box phylogenetic footprints in glycolytic genes by chromatin immunoprecipitation assays. Mol Cell Biol. 2004;24(13):5923–36. doi:10.1128/MCB.24.13.5923-5936.2004.
Osthus RC, Shim H, Kim S, Li Q, Reddy R, Mukherjee M, et al. Deregulation of glucose transporter 1 and glycolytic gene expression by c-Myc. J Biol Chem. 2000;275(29):21797–800. doi:10.1074/jbc.C000023200.
Zheng X, Boyer L, Jin M, Mertens J, Kim Y, Ma L, et al. Metabolic reprogramming during neuronal differentiation from aerobic glycolysis to neuronal oxidative phosphorylation. eLife. 2016;5:e13374. doi:10.7554/eLife.13374.
Scarpulla RC, Vega RB, Kelly DP. Transcriptional integration of mitochondrial biogenesis. Trends in Endocrinology & Metabolism. 2012;23(9):459–66. doi:10.1016/j.tem.2012.06.006.
Torres-Padilla ME, Chambers I. Transcription factor heterogeneity in pluripotent stem cells: a stochastic advantage. Development. 2014;141(11):2173–81. doi:10.1242/dev.102624.
Chen CL, Uthaya Kumar DB, Punj V, Xu J, Sher L, Tahara SM, et al. NANOG metabolically reprograms tumor-initiating stem-like cells through tumorigenic changes in oxidative phosphorylation and fatty acid metabolism. Cell Metab. 2016;23(1):206–19. doi:10.1016/j.cmet.2015.12.004.
Boiani M, Scholer HR. Regulatory networks in embryo-derived pluripotent stem cells. Nat Rev Mol Cell Biol. 2005;6(11):872–84. doi:10.1038/nrm1744.
Kim H, Jang H, Kim TW, Kang B-H, Lee SE, Jeon YK, et al. Core pluripotency factors directly regulate metabolism in embryonic stem cell to maintain pluripotency. Stem Cells. 2015;33(9):2699–711. doi:10.1002/stem.2073.
Zhang J, Nuebel E, Daley George Q, Koehler Carla M, Teitell MA. Metabolic regulation in pluripotent stem cells during reprogramming and self-renewal. Cell Stem Cell. 2012;11(5):589–95. doi:10.1016/j.stem.2012.10.005.
Haq R, Shoag J, Andreu-Perez P, Yokoyama S, Edelman H, Rowe Glenn C, et al. Oncogenic BRAF regulates oxidative metabolism via PGC1α and MITF. Cancer Cell. 2013;23(3):302–15. doi:10.1016/j.ccr.2013.02.003.
Vazquez F, Lim J-H, Chim H, Bhalla K, Girnun G, Pierce K, et al. PGC1α expression defines a subset of human melanoma tumors with increased mitochondrial capacity and resistance to oxidative stress. Cancer Cell. 2013;23(3):287–301. doi:10.1016/j.ccr.2012.11.020.
Garraway LA, Widlund HR, Rubin MA, Getz G, Berger AJ, Ramaswamy S, et al. Integrative genomic analyses identify MITF as a lineage survival oncogene amplified in malignant melanoma. Nature. 2005;436(7047):117–22.
Wray J, Kalkan T, Smith AG. The ground state of pluripotency. Biochem Soc Trans. 2010;38(4):1027–32. doi:10.1042/BST0381027.
Carbognin E, Betto RM, Soriano ME, Smith AG, Martello G. Stat3 promotes mitochondrial transcription and oxidative respiration during maintenance and induction of naive pluripotency. EMBO J. 2016;35(6):618–34. doi:10.15252/embj.201592629.
Acknowledgments
H.-F.D. is supported by a grant from the US National Institutes of Health (R01CA190429).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
Ahmet Alptekin, Bingwei Ye, and Han-Fei Ding declare that they have no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
This article is part of the Topical Collection on Metabolism and Stem Cells
Rights and permissions
About this article
Cite this article
Alptekin, A., Ye, B. & Ding, HF. Transcriptional Regulation of Stem Cell and Cancer Stem Cell Metabolism. Curr Stem Cell Rep 3, 19–27 (2017). https://doi.org/10.1007/s40778-017-0071-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40778-017-0071-y