Abstract
Therapy-related myeloid neoplasms are a life-threatening and often fatal complication, associated with poor prognosis outcomes and with high-risk unfavorable cytogenetic abnormalities including complex karyotype. They occur after the treatment of primary malignancies using chemotherapy and/or radiation therapy. Such therapy is not specific to cancer cells, and also damages the deoxyribonucleic acid (DNA) of normal cells, resulting in unbalanced and balanced translocations. There are eight genetic pathways, whose details are summarized in this review, depending on the cytogenetic abnormalities induced. This abnormality is the major contributor to the development of therapy-related myeloid neoplasms. The etiology of these neoplasms depends on the complex interaction between the nature and dose of the cytotoxic agent, the environment, and the presence of subsequent inherited mutations. This review aims to elaborate upon recent knowledge regarding the etiology, pathogenesis, and genetic pathways of therapy-related myeloid neoplasms. A deeper understanding of their etiology would aid physicians in more careful monitoring of patients during or after cytotoxic therapy for hematological malignancy. Ultimately, this knowledge could influence initial treatment strategies, with the aim of reducing both the incidence and serious complications of neoplasms. Therefore, early detection of DNA lesions is vital. The authors recommend that primary malignancy be treated with targeted therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Why carry out this study? |
Therapy-related myeloid neoplasm is a life-threatening and often fatal complication. |
It is associated with poor prognosis outcomes and with high-risk unfavorable cytogenetic abnormalities including complex karyotype. |
Treating primary hematological disorders with targeted treatment decreases the incidence of therapy-related myeloid neoplasms and increases survival rates among patients. |
What was learned from the study? |
We recommend that primary malignancies be treated with targeted therapy. |
This review document helps to increase our understanding of the pathogenesis, etiology, and consequences of therapy-related leukemia. |
Introduction
Therapy-related myeloid neoplasms (t-MN) are well-recognized hematopoietic stem cell malignant neoplasms which arise as a result of mutational events and are provoked by earlier exposure to chemo- and/or radiotherapy of primary hematological malignancies, solid tumors, and autoimmune disease [1,2,3]. They develop after the occurrence of mutations induced primarily by previous cytotoxic therapy of hematological malignancies [4]. Cytotoxic therapy can lead to other mutations due to its lack of specificity for cancer cells, thus promoting the development of t-MN. t-MN can be divided into three categories: therapy-related acute myeloid leukemia (t-AML), therapy-related myelodysplastic syndrome (t-MDS), and therapy-related myelodysplastic/myeloproliferative neoplasm (t-MDS/MPN) [5].
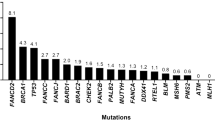
Globally, the incidence of t-MN continues to increase due to the increased prevalence of hematological malignancy. Previous finding have shown an incidence as high as 10–20%. Risk factors such as exposure to alkylating agents, topoisomerase (TOP) II inhibitors, radiation therapy, age, and genetic susceptibility play a contributing role [6]. The side effects of chemotherapy were found to be responsible for a 4.7-fold greater incidence. t-MN is thus becoming a growing healthcare problem worldwide due to the absence of targeted therapy for primary hematological malignancies (Table 1), solid tumors, and autoimmune diseases [7].
t-MN is generally a fatal disease, with life-threatening complications. This is may be due to increased number of blasts in the bone marrow or blood and prolonged cytopenias. The patient is vulnerable to bleeding and various systemic infections. t-MN is characterized by poor prognosis, insidious disease onset with peripheral cytopenias, and high-risk unfavorable cytogenetic abnormalities such as loss of chromosomes 5q and/or 7q and complex karyotype (three or more chromosome abnormalities). Because of this, t-MN is the most serious unpredictable lifelong complication and the greatest barrier to patient cure. Currently, the side effects of cytotoxic therapy represent a significant challenge for patients, as they lead to cardiac disease, chronic pulmonary diseases, permanent bone marrow modification, and direct DNA damage. They also have a direct impact on the economic and social lives of patients [15,16,17].
The aim of this review is to elaborate on the recent knowledge of the etiology, pathogenesis, and genetic pathway of t-MN, focusing specifically on the side effects of traditional therapies. The poor prognosis for patients, unfavorable cytogenetic abnormalities, and therapy that is not targeted to cancer cells results in poor survival. This traditional therapy is not specific to cancerous cells and causes abnormal DNA lesions in normal cells. The early detection of DNA lesions during treatment follow-up is vital for increasing survival time and improving patient outcomes. In addition, early identification of the etiology of t-MN is important to the health professional for preventing additional complications (side effects of therapy). It also guides early therapeutic decision-making for physicians with regard to cytogenetic abnormalities. These issues motivated us to conduct a review of the etiology, pathogenesis, and genetic pathway of t-MN.
This review was conducted on the basis of the relevant literature on the current topic retrieved from electronic databases including PubMed, PubMed Central, Scopus, Google, ScienceDirect, and Google Scholar. Articles on the current issues were searched using keywords and phrases including genetic pathway, t-MN, t-AML, t-MDS, chemotherapy, and radiation therapy, separately and in combination. Articles such as reviews, systematic reviews, and meta-analyses were also considered. The current review includes original, peer-reviewed English-language articles.
Compliance with Ethics Guidelines
This article is based on previously conducted studies and does not contain any studies with human participants or animals performed by any of the authors.
Etiology of Therapy-Related Myeloid Neoplasms
The etiology of t-MN depends on the complex interaction between the nature and dose of the chemotherapeutic agent, radiation intensity, genetic factors, and the environment. Patients with inherited mutations who are additionally exposed to chemo- and/or radiation therapies are susceptible to further induced secondary mutations. These subsequent mutations confer a competitive advantage for abnormal cell clonal proliferation over normal hematopoiesis, resulting in the development of genetic instability and factors that favor leukemia-initiating cells [18,19,20].
Alkylating Agents
Alkylating agents are a large group of chemotherapeutic drugs, and play an important role in the treatment of several types of cancers [21]. However, they are not specific to cancer cells, and cause the loss of the long arm of chromosomes 5q and 7q, resulting in unbalanced chromosomal aberrations [22]. These agents have a long latency period (on average 5 years), presenting as mostly t-MDS, with poor prognosis [20]. Latency is defined as the time from the first cytotoxic exposure to the first bone marrow examination for the diagnosis of t-MN [15].
There are several different alkylating agents now used for the treatment of many types of cancer. These include melphalan, cyclophosphamide, cisplatin, busulfan, and chlorambucil, all of which are leukemogenic [7].
Mechanism of Action of Alkylating Agents
Alkylating agents destroy cancer cells by the transfer of alkyl groups such as –CH3 or –CH2–CH3 to oxygen or nitrogen atoms of DNA bases. This results in a highly mutagenic DNA base lesion [16]. These agents kill cancer cells by two mechanisms: either by DNA methylation (mono-functional alkylating agent) or by cross-link formation of the DNA strand (bi-functional alkylating agent). Both mechanisms covalently modify the DNA structure, causing a double-strand break (DSB) [20]. Mono-functional alkylating agents destroy cancer cells by the addition of a methyl group to the DNA molecule. DNA methylation is the process of adding a methyl group to the DNA molecule without changing the DNA sequence. This modifies the function of the genes and affects gene expression. The most widely characterized DNA methylation in humans is 5-methylcytosine. Spontaneous deamination of 5-methylcytosine yields thymine. The outcome is a thymine mismatch pair with guanine, which is recognized by DNA mismatch repair (MMR). However, MMR does not remove the methylated base, and becomes a permanent mutation. This results in DSBs, leading to resistance to killing by methylating agents [7, 17, 21, 23].
In contrast to mono-functional alkylating agents, bi-functional alkylating agents have two reactive sites. They can form DNA intrastrand or interstrand cross-links by attaching two opposing bases in a complementary DNA strand. This cross-linkage prevents the uncoiling of the DNA double helix during replication [7, 24]. During replication, interstrand cross-links stop replication forks, which can result in the formation of DNA DSBs. If left unrepaired, these DSBs can give rise to translocations, inversions, and insertions of other chromosomes producing abnormal cells [18].
Topoisomerase (TOP) II Inhibitors
DNA TOPs are critical enzymes responsible for unknotting and relaxing supercoiled DNA, thus allowing DNA replication to occur. TOP binds covalently to the DNA strand and creates a transient single-strand (type I TOP, cleaves one strand of DNA at a time) and DSB (type II TOP, cleaves both strands of the DNA). However, TOP II inhibitors bind to the enzyme/DNA complex at the strand cleavage stage of the TOP reaction and block the re-ligation step, leading to an accumulation of DSBs [18, 19]. Multiple DSBs lead to cell death or chromosomal breakage, causing initiation of leukemia and transformation to abnormal proliferation of cancerous cells [23]. The TOP II inhibitor induces t-AML with balanced translocations that generally arise within 3 years. It is often associated with mixed lymphocytic leukemia (MLL), runt-related transcription factor 1 (RUNX1), and retinoic acid receptor alpha (RARA) loci at 11q23, 21q22, and 17q21, respectively [25, 26].
The balance between enzyme-mediated DNA cleavage and ligation is critical for cell survival. If the level of TOP II-mediated DNA cleavage is below a threshold level, cells ultimately die. If the level of cleavage becomes too high, it results in DNA breaks [27].
Ionizing Radiation
Exposure of cells to ionizing radiation results in the creation of reactive oxygen species through the radiolysis of water molecules. The most important reactive oxygen species are hydroxyl radicals, superoxide radicals, and hydrogen peroxide. These species are highly reactive molecules that can oxidize or deaminate the DNA bases and increase the frequency of DNA DSBs [28]. Radiation photon energy can also directly induce strand breaks by disrupting the sugar-phosphate backbone of DNA to generate DSBs. These DSBs are highly mutagenic, potentially leading to the development of large-scale chromosomal rearrangements that are often found in t-MN [29].
Antimetabolites
Antimetabolites are a group of cytostatic drugs involved in the progression of t-MNs. These drugs are incorporated into the DNA, thereby interfering with the replication phase and leading to cell cycle arrest and apoptosis. Once placed in the newly synthesized DNA strand, they are prone to methylation and the formation of the highly mutagenic base lesion of 6-thio-methylguanine. The DNA MMR mechanism triggers cell cycle arrest and cell death after antimetabolite treatment. However, DNA MMR is unable to repair the lesions, and cells can tolerate 6-thioguanine, potentially proliferate and undergo leukemic clones for the expansion of t-MN [18].
Genetic Pathways of Therapy-Related Myeloid Neoplasms
Based on abnormal cytogenetic characteristics produced through cytotoxic therapy, eight different genetic pathways involved in t-MN have been proposed by different authors (Table 2) [30, 31]. The cytotoxic therapy results in unbalanced chromosomal translocation, balanced chromosome aberrations, and normal karyotype [32].
Pathway I
This pathway is characterized by the loss of the long arm of chromosome 7 (−7q) but with normal chromosome 5q. The loss of this chromosome further causes extra chromosome aberrations such as balanced t(3; 21). This chromosome abnormality is found mostly in t-AML and t-MDS after chemotherapy treatment. It is also characterized by mutations of the rat sarcoma (RAS) pathway [33, 34], overexpression of Fm-like tyrosine kinases (FLT3), impaired differentiation, and poor prognosis. Mutations in RAS genes can lead to the production of permanently activated RAS proteins and their various signaling pathways. As a result, this can cause unintended and overactive signaling inside the cell, even in the absence of incoming signals, leading to uncontrolled proliferation of abnormal cells [35].
This pathway is also closely associated with earlier therapy with alkylating agents as well as point mutations of the RUNX1 gene [31, 36]. Mutation of the RAS oncogene affects the function of neuroblastoma rat sarcoma (NRAS), and Kirsten rat sarcoma (KRAS), leading to aberrant proliferative signaling through activation of the extracellular signal-regulated kinase pathway [37].
Pathway II
This is initiated by deletion of chromosome 5 (that contains myeloid tumor suppressor gene) with or without abnormalities of chromosome 7, due to previous cytotoxic therapy [38]. It is also associated with mutations of TP53 [39,40,41] duplication of chromosome 8, impaired differentiation, and increased expression of cell cycle regulatory proteins due to parallel loss of the tumor suppressor gene, and results in poor outcomes [42]. The function of tumor suppressor protein is to regulate mitosis and the G2 checkpoint cell cycle, control transcription, and regulate translation. This loss of the tumor protein enables the rapid expression and proliferation of cancer cells [35].
Pathway III
This pathway is characterized by balanced translocations to chromosome band 11q23 with chimeric rearrangement between the MLL gene and one of its many partner genes. This aberration is caused by the cleavage of the MLL gene associated with treatment by TOP II inhibitors [42]. TOP II plays a role in replication, transcription, chromosome condensation, and segregation. The dimeric enzyme cleaves DNA at a pair of phosphodiester bonds at four base pairs apart and generates a staggered DSB cleavage complex. This enzyme remains covalently bound to the ends of the DSB via a 5′-phosphotyrosyl linkage. This cleavage results in the formation of gene fusion between MLL and other partner genes at the transcription promoter region. There are more than 50 different partner genes involved in inappropriate ligation in various human leukemias, especially t-MN [43, 44].
Pathway IV
This pathway also involves balanced translocations of chromosome band 21q22 or 16q22 due to TOP II inhibitors, leading to chimeric rearrangements between RUNX1 and core-binding factor-beta (CBFB). It has the best prognosis for t-MN. The RUNX1 and CBFB genes encode two proteins, which as heterodimers make up the CBFB complex, an important transcription factor in normal hematopoiesis. RUNX1 is the cleavage site for the TOP II enzyme and fuses near internal transcription start regions. The fusion gene inhibits transcription genes that are essential for normal hematopoietic proliferation and differentiation of cells [45, 46].
Pathway V
This pathway is found in patients who present with therapy-related promyelocytic leukemia (PML) with the t(15; 17) resulting in the PML-RARA fusion and mutation of FML3. The FLT3 genes encode membrane-bound class III receptor of tyrosine kinase (RTKs) and are important mediators of mitogenic signal transduction. Two types of FLT3 mutations have been identified: internal tandem duplications (ITD) of the juxtamembrane domain and activating loop mutations in the cytoplasm tyrosine kinase (TK) domain. This pathway is associated with a good prognosis as compared with the previous pathways [47].
Pathway VI
This pathway occurs due to balanced translocations of 11p15 associated with the TOP II inhibitor. The fusion gene nucleoporin 98 (NUP98) and homebox gene (HOX) occur in the breakpoint region of inversion [18] (p15q22). Nucleoporin functions as a site for the mediating transport of RNA and protein between the cytoplasm and nucleus. The HOX family genes encode evolutionarily conserved transcription factors containing the 61-amino acid home domain. The genes seem to be the master control of transcription factors. They play a major role in the early and late stages of development and hematopoietic stem cell differentiation into mature cells. The HOX genes in hematopoietic cells show lineage and differentiation stage-specific expression patterns and seem to regulate hematopoiesis. More recently, the NUP98–HOX fusion gene has been cloned from the breakpoint region of t(2;11)(q31;p15) associated with childhood t-AML [48, 49].
Pathway VII
This t-MN pathway is characterized by normal karyotype, ITD of FLT3 and MLL, and nucleophosmin gene (NPM1) mutations. NPM1 is involved during ribosome biogenesis and centromere duplication. It also modulates the activity of the TP53 and tumor suppressors in cell division [35].
Pathway VIII
This pathway is uncharacteristic; sometimes unique chromosome aberrations are grouped and other chromosomal abnormalities [36].
Pathogenesis of Therapy-Related Myeloid Neoplasm
T-MN results from a complex interaction between individual predisposition and exposure to cytotoxic agents. It is determined by subsequent acquisition of somatic mutations and epigenetic modifications. Somatic mutations are critical genes that may precede and favor leukemia development [9]. Exposure to chemotherapeutic agents results in DNA damage, genetic mutations, mitochondrial dysfunction, and alteration of cellular metabolism and deregulation of normal hematopoietic progenitor cells [50, 51].
Various studies show that both class I and class II mutations are required for the pathogenesis of t-MN. Class I mutations are characterized by constitutive activation of receptor tyrosine kinases (TK) that result in abnormal cell proliferation and survival. It includes mutation of the RTK, FLT3, and the intracellular non-receptor TK, together with KRAS, NRAS, and extracellular signal transduction pathway proteins [52]. Class II mutations, on the contrary, inactivate hematopoietic transcription factors and result in the loss of cell differentiation that occurs in t-MDS/AML [22, 30].
TK is the major target for the treatment of cancers. However, its function is deregulated by three mechanisms. The first mechanism is the fusion of a receptor or non-receptor TK with a partner protein by balanced chromosomal translocation formed due to cytotoxic therapy. This translocation leads to constitutive oligomerization of the TK in the absence of ligand binding or physiologic activating signals, thereby promoting autophosphorylation and activation. This leads to the high proliferation of cells occurring in the bone marrow [53].
The second mechanism involves mutation FLT3, small deletions, and point mutations in the kinase domain that disrupt autoregulation of the kinase. The third mechanism relates to the overexpression of the receptor TK. Increased TK activity can result from a decrease in factors that limit TK activity, such as impaired tyrosine phosphatase activity and decreased expression of TK inhibitor proteins. Aberrant TK activation can increase the survival, proliferation, and cytotoxic drug resistance of malignant cells. In general, the pathogenesis of t-MN can be summarized by three mechanisms: direct initiation of fusion genes, induction of genetic instability, and selection of pre-existing mutated cells in the bone marrow during cytotoxic treatment [54, 55].
Direct Initiation of Fusion Genes
The oncogene gene is provoked in a susceptible target cell during cytotoxic therapy. This leads to clonal outgrowth of the transformed cell mediated by TOP II inhibitors. These inhibitors interfere with the re-ligation step during DNA replication and generate single-stranded DNA for the cancer cells. This mechanism reduces the progression and proliferation of cancer cells [17, 19]. However, these inhibitors are not specific for cancer cells, and also cause DSBs of the normal cell DNA. These DSBs are relatively stable and mobile, and form balanced chromosomal translocations that easily occur in close-linked genes. These rearrangements mainly lead to the activation of a proto-oncogene that disrupts the normal balance of the oncogene and tumor suppressor genes. This imbalance of gene control also provides a selective growth advantage for the proliferation of cancer cells during carcinogenesis that initiate the development of t-MN [56].
Patients are exposed to TOP II inhibitors to treat the primary malignancy, but this inhibitor causes breaks in the normal DNA. The double-strand DNA breaks give rise to the formation of balanced translocations including MLL at 11q23, NUP98 at 11p15, RUNX1 at 21q22, and RARA at 17q21 [57]. These changes characteristically result in a dominant loss of function of the transcription factor (class II mutations), with defects in differentiation and increased self-renewal capacity [53].
The balanced translocation between band 11q23 and the MLL gene most frequently occurs in t-MN associated with TOP II inhibitors. The MLL gene has three domains: the methyltransferase domain (contains transcription repressor protein), which is involved in the epigenetic regulation of transcription by methylation, and the activation domain and set domain for recruiting chromatin re-modeling complexes to specific chromosomal regions. This gene function is directly bound to the DNA at the promoter region and regulates the target genes. Translocations involving band 11q23 usually lead to a breakage in the MLL gene. The 5′ part of the MLL gene is retained on the derivative of chromosome 11, where it is fused with the 3′ part of the partner gene. Therefore, the active fusion gene (5′ MLL-3′partner) is almost always located in the derivative of chromosome 11 [49, 58].
The breakpoints within the MLL gene cluster in the 8.5 kb region, called the breakpoint cluster region, located between exons 5 and 11 in the TOP II inhibitor binding sites. The inhibitors inhibit the ligase enzyme function of the TOP II enzyme, leaving DNA free ends that can repair through non-homologous recombination between the MLL and the partner genes. The fusion gene occurs in the methyltransferase domain that lacks set and activation domains. For this reason, histone methylation at the promoter region cannot occur. MLL contains enzyme-like histone methyltransferase that is critical for regulating gene expression during hematopoiesis. The fusion of MLL with other proteins results in oncogenic gene products that trigger faulty self-renewal of stem cells, leading to leukemia [59, 60].
The other balanced translocation that occurs due to the TOP II inhibitor is t(15;17), which results in the fusion of the PML gene at 15q22 with the RARA at 17q21. The TOP II inhibitor enhances the double-strand cleavage of the normal homologs of the PML and RARA genes into two sections, the PML (UPN1, UPN4) and RARA breakpoints (UPN2, UPN5). They share breakpoints between the PML and RARA genes and form a reciprocal translocation, PML–RARA and RARA–PML. This fusion induces further development of t-MN [61].
Induction of Genetic Instability
The second mechanism is related to the cytotoxic therapy itself. The induction of radiation, chemotherapy, and/or combination therapy cause endogenous DSBs of the DNA. The DNA lesions can alter the primary structure of the double helix, thereby affecting transcription and replication of the normal process [62]. The chemotherapy/radiation kills or reduces the progression of the cancer cells. Its target is DNA but it is not specific for the cancer cells of the DNA, and thus causes the loss of the whole long arm of chromosomes 5q and 7q. The abnormalities of these chromosomes are often associated with a complex karyotype, including recurrent and non-recurrent chromosome aberrations. These abnormalities should be undergoing DNA repair mechanisms and result in the activation of ataxia-telangiectasia mutation (ATM) gene signaling for the DNA repair process [63].
The ATM gene is normally responsible for the activation of the P53 protein and upregulation of genes involved in apoptosis, playing a central role in the repair of these DNA lesions [63]. However, the function of the ATM gene is dysregulated by chemo- and/or radiation therapy and is unable to undergo autophosphorylation and activate the P53 protein [64].
The p53 gene has a critical role in DNA damage response signaling, affecting cell cycle, cell death, and DNA repair pathways. However, the effects of cytotoxic therapy on p53 cause the occurrence of cell proliferation without repair and resistance to apoptosis. Abnormal p53 activity leads to reduced ability to repair DNA damage, resulting in genetic instability and increased susceptibility to leukemogenesis of t-MN [65, 66].
Selection of Pre-existing Mutated Hematopoietic Cell Clones
The third mechanism of t-MN pathogenesis involves the proliferation of pre-existing mutated cell clones in the bone marrow during treatment of the primary malignancy. Chemotherapy and/or radiation therapy may exert selective pressure on hematopoietic stem cells such that certain mutant populations (known as clones) have a selective advantage under cytotoxic conditions [67]. These mutant clones serve as premalignant cells with a growth advantage during cytotoxic therapy for the development of t-MN. One of the most common pre-existing mutations in the marrow is TP53. A previous study indicates that TP53 mutant cells can exist at low frequencies in the bone marrow prior to chemotherapy and then rapidly proliferate during or after cytotoxic treatment, thus playing a major role in the progression to t-MN [68].
Clonal hematopoiesis has recently been associated with mutations in protein phosphatase Mn2+/Mg2+-dependent 1D (PPM1D), which is part of the DNA damage response pathway. PPM1D is part of a regulatory feedback loop of p53. It activates p53 and induces expression of PPM1D, which then both directly and indirectly dephosphorylates p53, leading to downregulation of p53-mediated apoptosis. Mutations in PPM1D are typically nonsense or frameshift mutations in the sixth exons that produce a C-terminal truncated protein. PPM1D mutations have been frequently observed specifically in patients with t-MN. Somatic mutations of PPMID and TP53 are resistant to chemotherapy and increase preferentially after treatment and accumulate in the bone marrow without differentiation, which leads to further development of t-MNs [67, 69].
Conclusion
t-MNs are a serious, long-term, and life-threatening complication that develop as a result of previous cytotoxic therapy of primary hematological and non-hematological malignancies. The therapy itself causes cytogenetic abnormalities including loss of the long arm of chromosome 7q and/or 5q and different balanced chromosomal aberrations. The pathogenesis of t-MNs is summarized by three mechanisms: direct induction of an oncogene, genetic instability, and selection of a pre-existing mutated cell in the marrow. It also associated with eight genetic pathways summarized by unbalanced chromosome aberrations (pathways I and II), balanced rearrangements (pathways III–VI), and normal karyotype (pathways VII and VIII). We conclude that in order to reduce the side effects of chemo- and/or radiation therapy and the risk of t-MN and to increase patient survival, the primary malignancy should be treated with targeted therapy.
References
Arber DA, Orazi A, Hasserjian R, Thiele J, Borowitz MJ, Le Beau MM, et al. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood. 2016;127(20):2391–405.
Godley LA, Njiaju UO, Green M, Weiner H, Lin S, Odenike O, et al. Treatment of therapy-related myeloid neoplasms with high-dose cytarabine/mitoxantrone followed by hematopoietic stem cell transplant. Leuk Lymphoma. 2010;51(6):995–1006.
Yee-Loong Tang D, Wai-Kit Chia M, Yap W, Shuib Salwati D. Dismal outcome of therapy-related myeloid neoplasm associated with complex aberrant karyotypes and monosomal karyotype: a case report. Malays J Pathol. 2016;38(3):315.
Larson RA. Etiology and management of therapy-related myeloid leukemia. ASH Educ Program Book. 2007;2007(1):453–9.
Bueso-Ramos CE, Kanagal-Shamanna R, Routbort MJ, Hanson CA. Therapy-related myeloid neoplasms. Am J Clin Pathol. 2015;144(2):207–18.
Metafuni E, Chiusolo P, Laurenti L, Sorà F, Giammarco S, Bacigalupo A, et al. Allogeneic hematopoietic stem cell transplantation in therapy-related myeloid neoplasms (t-MN) of the adult: monocentric observational study and review of the literature. Mediterr J Hematol Infect Dis. 2018; 10(1):e2018005.
Joannides M, Grimwade D. Molecular biology of therapy-related leukaemias. Clin Transl Oncol. 2010;12(1):8–14.
Espírito Santo A, Chacim S, Ferreira I, Leite L, Moreira C, Pereira D, et al. Effect of therapy-related acute myeloid leukemia on the outcome of patients with acute myeloid leukemia. Oncol Lett. 2016;12(1):262–8.
Fianchi L, Pagano L, Piciocchi A, Candoni A, Gaidano G, Breccia M, et al. Characteristics and outcome of therapy-related myeloid neoplasms: report from the Italian network on secondary leukemias. Am J Hematol. 2015;90(5):E80–5.
Zhou Y, Tang G, Medeiros LJ, McDonnell TJ, Keating MJ, Wierda WG, et al. Therapy-related myeloid neoplasms following fludarabine, cyclophosphamide, and rituximab (FCR) treatment in patients with chronic lymphocytic leukemia/small lymphocytic lymphoma. Mod Pathol. 2012;25(2):237–45.
Imagawa J, Harada Y, Shimomura T, Tanaka H, Okikawa Y, Hyodo H, et al. Clinical and genetic features of therapy-related myeloid neoplasms after chemotherapy for acute promyelocytic leukemia. Blood J Am Soc Hematol. 2010;116(26):6018–22.
Nardi V, Winkfield KM, Ok CY, Niemierko A, Kluk MJ, Attar EC, et al. Acute myeloid leukemia and myelodysplastic syndromes after radiation therapy are similar to de novo disease and differ from other therapy-related myeloid neoplasms. J Clin Oncol. 2012;30(19):2340.
Kayser S, Döhner K, Krauter J, Köhne C-H, Horst HA, Held G, et al. The impact of therapy-related acute myeloid leukemia (AML) on outcome in 2853 adult patients with newly diagnosed AML. Blood J Am Soc Hematol. 2011;117(7):2137–45.
Epperla N, Pham AQ, Burnette BL, Wiseman GA, Habermann TM, Macon WR, et al. Risk of histological transformation and therapy-related myelodysplasia/acute myeloid leukaemia in patients receiving radioimmunotherapy for follicular lymphoma. Br J Haematol. 2017;178(3):427–33.
Churpek JE, Marquez R, Neistadt B, Claussen K, Lee MK, Churpek MM, et al. Inherited mutations in cancer susceptibility genes are common among survivors of breast cancer who develop therapy-related leukemia. Cancer. 2016;122(2):304–11.
Bhatia S. Therapy-related myelodysplasia and acute myeloid leukemia. Semin Oncol. 2013;40(6):666–75.
Heuser M. Therapy-related myeloid neoplasms: does knowing the origin help to guide treatment? Hematol Am Soc Hematol Educ Program. 2016;2016(1):24–32.
Sill H, Olipitz W, Zebisch A, Schulz E, Wölfler A. Therapy-related myeloid neoplasms: pathobiology and clinical characteristics. Br J Pharmacol. 2011;162(4):792–805.
Cowell IG, Austin CA. Mechanism of generation of therapy related leukemia in response to anti-topoisomerase II agents. Int J Environ Res Public Health. 2012;9(6):2075–91.
McNerney ME, Godley LA, Le Beau MM. Therapy-related myeloid neoplasms: when genetics and environment collide. Nat Rev Cancer. 2017;17(9):513.
Casorelli I, Bossa C, Bignami M. DNA damage and repair in human cancer: molecular mechanisms and contribution to therapy-related leukemias. Int J Environ Res Public Health. 2012;9(8):2636–57.
Qian Z, Joslin JM, Tennant TR, Reshmi SC, Young DJ, Stoddart A, et al. Cytogenetic and genetic pathways in therapy-related acute myeloid leukemia. Chem Biol Interact. 2010;184(1–2):50–7.
Leone G, Fianchi L, Pagano L, Voso MT. Incidence and susceptibility to therapy-related myeloid neoplasms. Chem Biol Interact. 2010;184(1–2):39–45.
Helleday T, Petermann E, Lundin C, Hodgson B, Sharma RA. DNA repair pathways as targets for cancer therapy. Nat Rev Cancer. 2008;8(3):193–204.
Hashimoto A, Takada K, Horiguchi H, Sato T, Iyama S, Murase K, et al. Combination chemotherapy of azacitidine and cetuximab for therapy-related acute myeloid leukemia following oxaliplatin for metastatic colorectal cancer. Case Rep Oncol. 2014;7(2):316–22.
Larson RA. Therapy-related myeloid neoplasms. Haematologica. 2009;94(4):454–9.
Pendleton M, Lindsey RH, Felix CA, Grimwade D, Osheroff N. Topoisomerase II and leukemia. Ann N Y Acad Sci. 2014;1310(1):98–110.
Rassool FV, Gaymes TJ, Omidvar N, Brady N, Beurlet S, Pla M, et al. Reactive oxygen species, DNA damage, and error-prone repair: a model for genomic instability with progression in myeloid leukemia? Cancer Res. 2007;67:8762–71.
Klymenko SV, Bink K, Trott KR, Bebeshko VG, Bazyka DA, Dmytrenko IV, et al. MLL gene alterations in radiation-associated acute myeloid leukemia. Exp Oncol. 2005;27:71–5.
Czader M, Orazi A. Therapy-related myeloid neoplasms. Am J Clin Pathol. 2009;132(3):410–25.
Westman M, Pedersen-Bjergaard J, Andersen M, Andersen M. IDH1 and IDH2 mutations in therapy-related myelodysplastic syndrome and acute myeloid leukemia are associated with a normal karyotype and with der (1; 7)(q10; p10). Leukemia. 2013;27(4):957–9.
Ji Z, Zhang L, Peng V, Ren X, McHale C, Smith M. A comparison of the cytogenetic alterations and global DNA hypomethylation induced by the benzene metabolite, hydroquinone, with those induced by melphalan and etoposide. Leukemia. 2010;24(5):986–91.
Stephenson J, Lizhen H, Mufti GJ. Possible co-existence of RAS activation and monosomy 7 in the leukaemic transformation of myelodysplastic syndromes. Leuk Res. 1995;19(10):741–8.
Side L, Teel K, Wang P, Mahgoub N, Larson R, LeBeau M, et al. Activating RAS mutations in therapy-related myeloid disorders associated with deletions of chromosomes 5 and 7. Blood. 1996;88(10):2252.
Stoddart A, McNerney ME, Bartom E, Bergerson R, Young DJ, Qian Z, et al. Genetic pathways leading to therapy-related myeloid neoplasms. Mediterr J Hematol Infect Dis. 2011;3(1):e2011019.
Pedersen-Bjergaard J, Christiansen D, Desta F, Andersen M. Alternative genetic pathways and cooperating genetic abnormalities in the pathogenesis of therapy-related myelodysplasia and acute myeloid leukemia. Leukemia. 2006;20(11):1943–9.
Cleven AH, Nardi V, Ok CY, Goswami M, Dal Cin P, Zheng Z, et al. High p53 protein expression in therapy-related myeloid neoplasms is associated with adverse karyotype and poor outcome. Mod Pathol. 2015;28(4):552.
Joslin JM, Fernald AA, Tennant TR, Davis EM, Kogan SC, Anastasi J, et al. Haploinsufficiency of EGR1, a candidate gene in the del (5q), leads to the development of myeloid disorders. Blood. 2007;110(2):719–26.
Horiike S, Misawa S, Kaneko H, Sasai Y, Kobayashi M, Fujii H, et al. Distinct genetic involvement of the TP53 gene in therapy-related leukemia and myelodysplasia with chromosomal losses of Nos 5 and/or 7 and its possible relationship to replication error phenotype. Leukemia. 1999;13(8):1235.
Christiansen DH, Andersen MK, Pedersen-Bjergaard J. Mutations with loss of heterozygosity of p53 are common in therapy-related myelodysplasia and acute myeloid leukemia after exposure to alkylating agents and significantly associated with deletion or loss of 5q, a complex karyotype, and a poor prognosis. J Clin Oncol. 2001;19(5):1405–13.
Shih AH, Chung SS, Dolezal EK, Zhang S-J, Abdel-Wahab OI, Park CY, et al. Mutational analysis of therapy-related myelodysplastic syndromes and acute myelogenous leukemia. Haematologica. 2013;98(6):908–12.
Mitelman F, Johansson B, Mertens F. The impact of translocations and gene fusions on cancer causation. Nat Rev Cancer. 2007;7(4):233–45.
Cowell IG, Sondka Z, Smith K, Lee KC, Manville CM, Sidorczuk-Lesthuruge M, et al. Model for MLL translocations in therapy-related leukemia involving topoisomerase IIβ-mediated DNA strand breaks and gene proximity. Proc Natl Acad Sci. 2012;109(23):8989–94.
Blanco JG, Edick MJ, Relling MV. Etoposide induces chimeric Mll gene fusions. FASEB J. 2004;18(1):173–5.
Smith KA, Cowell IG, Zhang Y, Sondka Z, Austin CA. The role of topoisomerase II beta on breakage and proximity of RUNX1 to partner alleles RUNX1T1 and EVI1. Genes Chromosom Cancer. 2014;53(2):117–28.
Itzhar N, Dessen P, Toujani S, Auger N, Preudhomme C, Richon C, et al. Chromosomal minimal critical regions in therapy-related leukemia appear different from those of de novo leukemia by high-resolution aCGH. PLoS One. 2011;6(2):e16623.
Godley Lucy A, Larson Richard A. Therapy-related myeloid leukemia. Semin Oncol. 2008;35(4):418–29.
Nishiyama M, Arai Y, Tsunematsu Y, Kobayashi H, Asami K, Yabe M, et al. 11p15 translocations involving the NUP98 gene in childhood therapy-related acute myeloid leukemia/myelodysplastic syndrome. Genes Chromosom Cancer. 1999;26(3):215–20.
De Braekeleer M, Morel F, Le Bris M-J, Herry A, Douet-Guilbert N. The MLL gene and translocations involving chromosomal band 11q23 in acute leukemia. Anticancer Res. 2005;25(3B):1931–44.
Murthy GS, Hamadani M, Dhakal B, Hari P, Atallah E. Incidence and survival of therapy related myeloid neoplasm in United States. Leukemia Res. 2018;71:95–9.
Akhtari M, Bhatt VR, Tandra PK, Krishnamurthy J, Horstman H, Dreessen A, et al. Therapy-related myeloid neoplasms after autologous hematopoietic stem cell transplantation in lymphoma patients. Cancer Biol Ther. 2013;14(12):1077–88.
Desta F, Christiansen D, Andersen M, Pedersen-Bjergaard J. Activating mutations of JAK2 V617F are uncommon in t-MDS and t-AML and are only observed in atypic cases. Leukemia. 2006;20(3):547–8.
Pedersen-Bjergaard J, Andersen MK, Andersen M, Christiansen D. Genetics of therapy-related myelodysplasia and acute myeloid leukemia. Leukemia. 2008;22(2):240–8.
Christiansen DH, Andersen MK, Desta F, Pedersen-Bjergaard J. Mutations of genes in the receptor tyrosine kinase (RTK)/RAS-BRAF signal transduction pathway in therapy-related myelodysplasia and acute myeloid leukemia. Leukemia. 2005;19(12):2232.
Side LE, Curtiss NP, Teel K, Kratz C, Wang PW, Larson RA, et al. RAS, FLT3, and TP53 mutations in therapy-related myeloid malignancies with abnormalities of chromosomes 5 and 7. Genes Chromosom Cancer. 2004;39(3):217–23.
Zheng J. Oncogenic chromosomal translocations and human cancer. Oncol Rep. 2013;30(5):2011–9.
Pedersen-Bjergaard J. Insights into leukemogenesis from therapy-related leukemia. N Engl J Med. 2005;352(15):1591–4.
Schoch C, Schnittger S, Klaus M, Kern W, Hiddemann W, Haferlach T. AML with 11q23/MLL abnormalities as defined by the WHO classification: incidence, partner chromosomes, FAB subtype, age distribution, and prognostic impact in an unselected series of 1897 cytogenetically analyzed AML cases. Blood. 2003;102(7):2395–402.
Larson RA, Le Beau MM, Ratain MJ, Rowley JD. Balanced translocations involving chromosome bands 11q23 and 21q22 in therapy-related leukemia. Blood. 1992;79(7):1892–3.
Super HJ, McCabe NR, Thirman MJ, Larson RA, Le Beau MM, Pedersen-Bjergaard J. Rearrangements of the MLL gene in therapy-related acute myeloid leukemia in patients previously treated with agents targeting DNA-topoisomerase II. Blood. 1993;82(12):3705–11.
Zhang R, Kim YM, Wang X, Li Y, Pang H, Lee JY. Coexistence of t (15; 17) and t (15; 16; 17) detected by fluorescence in situ hybridization in a patient with acute promyelocytic leukemia: a case report and literature review. Oncol Lett. 2014;8(3):1001–8.
Torgovnick A, Schumacher B. DNA repair mechanisms in cancer development and therapy. Front Genet. 2015;6:157.
Shieh A, Mohamed AA. A case of therapy-related acute myeloid leukemia in a patient with heterozygous mutations in the ataxia telangiectasia mutated gene. J Hematol. 2017;6(4):96–100.
Choi M, Kipps T, Kurzrock R. ATM mutations in cancer: therapeutic implications. Mol Cancer Ther. 2016;15(8):1781–91.
Fabiani E, Falconi G, Fianchi L, Criscuolo M, Ottone T, Cicconi L, et al. Clonal evolution in therapy-related neoplasms. Oncotarget. 2017;8(7):12031.
Schulz E, Kashofer K, Heitzer E, Mhatre KN, Speicher MR, Hoefler G, et al. Preexisting TP53 mutation in therapy-related acute myeloid leukemia. Ann Hematol. 2015;94(3):527–9.
Ok CY, Patel KP, Garcia-Manero G, Routbort MJ, Peng J, Tang G, et al. TP53 mutation characteristics in therapy-related myelodysplastic syndromes and acute myeloid leukemia is similar to de novo diseases. J Hematol Oncol. 2015;8(1):45.
Wong TN, Ramsingh G, Young AL, Miller CA, Touma W, Welch JS, et al. Role of TP53 mutations in the origin and evolution of therapy-related acute myeloid leukaemia. Nature. 2015;518:552–5.
Hsu JI, Dayaram T, Tovy A, De Braekeleer E, Jeong M, Wang F, et al. PPM1D mutations drive clonal hematopoiesis in response to cytotoxic chemotherapy. Cell Stem Cell. 2018;23(5):700–13.
Acknowledgements
Funding
No funding or sponsorship was received for this study or publication of this article.
Authorship
All named authors meet the International Committee of Medical Journal Editors (ICMJE) criteria for authorship for this article, take responsibility for the integrity of the work as a whole, and have given their approval for this version to be published.
Authorship Contributions
TT was a major contributor to the writing of this manuscript. BE and ES reviewed and edited the manuscript. All authors read and approved the final manuscript.
Disclosures
Tegenaw Tiruneh, Bamlaku Enawgaw, and Elias Shiferaw have nothing to disclose.
Compliance with Ethics Guidelines
This article is based on previously conducted studies and does not contain any studies with human participants or animals performed by any of the authors.
Data Availability
All data generated or analyzed during this review are included in this manuscript and published articles.
Open Access
This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
Author information
Authors and Affiliations
Corresponding author
Additional information
Enhanced Digital Features
To view enhanced digital features for this article go to https://doi.org/10.6084/m9.figshare.11907891.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Tiruneh, T., Enawgaw, B. & Shiferaw, E. Genetic Pathway in the Pathogenesis of Therapy-Related Myeloid Neoplasms: A Literature Review. Oncol Ther 8, 45–57 (2020). https://doi.org/10.1007/s40487-020-00111-7
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40487-020-00111-7