Background
MicroRNAs (miRNAs) are a significant type of non-coding RNAs, which usually were encoded by endogenous genes with about ~22 nt nucleotides. Accumulating biological experiments have shown that miRNAs have close associations with various human diseases. Although traditional experimental methods achieve great successes in miRNA-disease interaction identification, these methods also have some limitations. Therefore, it is necessary to develop computational method to predict miRNA-disease interactions.
Methods
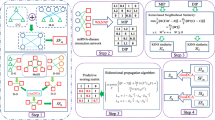
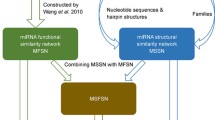
Here, we propose a computational framework (MDVSI) to predict interactions between miRNAs and diseases by integrating miRNA topological similarity and functional similarity. Firstly, the CosRA index is utilized to measure miRNA similarity based on network topological feature. Then, in order to enhance the reliability of miRNA similarity, the functional similarity and CosRA similarity are integrated based on linear weight method. Further, the potential miRNA-disease associations are predicted by using recommendation method. In addition, in order to overcome limitation of recommendation method, for new disease, a new strategy is proposed to predict potential interactions between miRNAs and new disease based on disease functional similarity.
Results
To evaluate the performance of different methods, we conduct ten-fold cross validation and de novo test in experiment and compare MDVSI with two the-state-of-art methods. The experimental result shows that MDVSI achieves an AUC of 0.91, which is at least 0.012 higher than other compared methods.
Conclusion
In summary, we propose a computational framework (MDSVI) for miRNA-disease interaction prediction. The experiment results demonstrate that it outperforms other the-state-of-the-art methods. Case study shows that it can effectively identify potential miRNA-disease interactions.

Article PDF
Similar content being viewed by others
Change history
14 September 2020
The original version of this article unfortunately contained a mistake in the name of the last author. The corrected name is Yi-Ping Phoebe Chen.
References
Ambros, V. (2004) The functions of animal microRNAs. Nature, 431, 350–355
Akhtar, M. M., Micolucci, L., Islam, M. S., Olivieri, F. and Procopio, A. D. (2016) Bioinformatic tools for microRNA dissection. Nucleic Acids Res., 44, 24–44
Iorio, M. V., Ferracin, M., Liu, C. G., Veronese, A., Spizzo, R., Sabbioni, S., Magri, E., Pedriali, M., Fabbri, M., Campiglio, M., et al. (2005) MicroRNA gene expression deregulation in human breast cancer. Cancer Res., 65, 7065–7070
Ashmawy, A. M., Elgeshy, K. M., Abdel Salam, E. T., Ghareeb, M., Kobaisi, M. H., Amin, H. A. A., Sharawy, S. K. and Abdel Wahab, A. H. A. (2017) Crosstalk between liver-related micro-RNAs and Wnt/β-catenin pathway in hepatocellular carcinoma patients. Arab J. Gastroenterol., 18, 144–150
Kim, Y. K., Yu, J., Han, T. S., Park, S. Y., Namkoong, B., Kim, D. H., Hur, K., Yoo, M. W., Lee, H. J., Yang, H. K., et al. (2009) Functional links between clustered microRNAs: suppression of cell-cycle inhibitors by microRNA clusters in gastric cancer. Nucleic Acids Res., 37, 1672–1681
Moss, E. G., Lee, R. C. and Ambros, V. (1997) The cold shock domain protein LIN-28 controls developmental timing in C. elegans and is regulated by the lin-4 RNA. Cell, 88, 637–646
Lan, W., Chen, Q., Li, T., Yuan, C., Mann, S. and Chen, B. (2014) Identification of important positions within miRNAs by integrating sequential and structural features. Curr. Protein Pept. Sci., 15, 591–597
Chen, Q., Lan, W., Wang, J. X. (2013) Mining featured patterns of miRNA interaction based on sequence and structure similarity. IEEE/ACM Trans. Comput. Biol. Bioinform., 10, 415–422
Luo, H., Lan, W., Chen, Q., Wang, Z., Liu, Z., Yue, X. and Zhu, L. (2018) Inferring microRNA-environmental factor interactions based on multiple biological information fusion. Molecules, 23, 2439
Lan, W., Wang, J. X., Li, M., Liu, J. and Pan, Y. (2015) Predicting microRNA-disease associations by integrating multiple biological information. In: EEE International Conference on Bioinformatics and Biomedicine (BIBM), pp. 183–188
Peng, W., Lan, W., Zhong, J., Wang, J. X. and Pan, Y. (2017) A novel method of predicting microRNA-disease associations based on microRNA, disease, gene and environment factor networks. Methods, 124, 69–77
Jiang, Q., Hao, Y., Wang, G., Juan, L., Zhang, T., Teng, M., Liu, Y. and Wang, Y. (2010) Prioritization of disease microRNAs through a human phenome-microRNAome network. BMC Syst. Biol., 4, S2
Xuan, P., Han, K., Guo, M., Guo, Y., Li, J., Ding, J., Liu, Y., Dai, Q., Li, J., Teng, Z., et al. (2013) Prediction of microRNAs associated with human diseases based on weighted k most similar neighbors. PLoS One, 8, e70204
Mørk, S., Pletscher-Frankild, S., Palleja Caro, A., Gorodkin, J. and Jensen, L. J. (2014) Protein-driven inference of miRNA-disease associations. Bioinformatics, 30, 392–397
Chen, X. and Yan, G. Y. (2014) Semi-supervised learning for potential human microRNA-disease associations inference. Sci. Rep., 4, 5501
Peng, W., Lan, W., Yu, Z., Wang, J., Pan, Y. (2016) A Framework for integrating multiple biological networks to predict microRNAdisease associations. IEEE Trans. Nanobio., 16, 100–107
Yan, C., Wang, J., Ni, P., Lan, W., Wu, F. X., Pan, Y. (2019) DNRLMF-MDA: predicting microRNA-disease associations based on similarities of microRNAs and diseases. IEEE/ACM Trans. Comput. Biol. Bioinform., 16, 233–243
Luo, J. and Xiao, Q. (2017) A novel approach for predicting microRNA-disease associations by unbalanced bi-random walk on heterogeneous network. J. Biomed. Inform., 66, 194–203
Lan, W., Wang, J. X., Li, M., Liu, J., Wu, F.X., Pan, Y. (2018) Predicting microRNA-disease associations based on improved microRNA and disease similarities. IEEE/ACM Trans. Comput. Biol. Bioinform., 15, 1774–1782
Liu, Y., Zeng, X., He, Z., Zou, Q. (2016) Inferring microRNAdisease associations by random walk on a heterogeneous network with multiple data sources. IEEE/ACM Trans. Comput. Biol. Bioinform., 14, 905–915
Zou, Q., Li, J., Hong, Q., Lin, Z., Wu, Y., Shi, H. and Ju, Y. (2015) Prediction of microRNA-disease associations based on social network analysis methods. BioMed Res. Int., 2015, 810514
Aushev, V. N., Zborovskaya, I. B., Laktionov, K. K., Girard, N., Cros, M. P., Herceg, Z. and Krutovskikh, V. (2013) Comparisons of microRNA patterns in plasma before and after tumor removal reveal new biomarkers of lung squamous cell carcinoma. PLoS One, 8, e78649
Chang, J. T., Wang, F., Chapin, W. and Huang, R. S. (2016) Identification of microRNAs as breast cancer prognosis markers through the Cancer Genome Atlas. PLoS One, 11, e0168284
Köhler, S., Bauer, S., Horn, D. and Robinson, P. N. (2008) Walking the interactome for prioritization of candidate disease genes. Am. J. Hum. Genet., 82, 949–958
Lan, W., Li, M., Zhao, K., Liu, J., Wu, F. X., Pan, Y. and Wang, J. (2017) LDAP: a web server for lncRNA-disease association prediction. Bioinformatics, 33, 458–460
Lan, W., Huang, L., Lai, D., and Chen, Q. (2018) Identifying Interactions Between Long Noncoding RNAs and Diseases Based on Computational Methods. In: Computational Systems Biology, pp. 205–221, Springer Nature
Lan, W., Wang, J. X., Li, M., Peng, W. and Wu, F. (2015) Computational approaches for prioritizing candidate disease genes based on PPI networks. Tsinghua Sci. Technol., 20, 500–512
Li, Y., Qiu, C., Tu, J., Geng, B., Yang, J., Jiang, T. and Cui, Q. (2014) HMDD v2.0: a database for experimentally supported human microRNA and disease associations. Nucleic Acids Res., 42, D1070–D1074
Hwang, S., Kim, C. Y., Yang, S., Kim, E., Hart, T., Marcotte, E. M. and Lee, I. (2019) HumanNet v2: human gene networks for disease research. Nucleic Acids Res., 47, D573–D580
Wang, D., Wang, J., Lu, M., Song, F. and Cui, Q. (2010) Inferring the human microRNA functional similarity and functional network based on microRNA-associated diseases. Bioinformatics, 26, 1644–1650
Chen, L. J., Zhang, Z. K., Liu, J. H., Gao, J. and Zhou, T. (2017) A vertex similarity index for better personalized recommendation. Physica A, 466, 607–615
Jiang, H., Wang, J., Li, M., Lan, W., Wu, F., Pan, Y. (2018) miRTRS: a recommendation algorithm for predicting miRNA targets. IEEE/ACM Trans. Comput. Biol. Bioinform., 1
Acknowledgements
The work reported in this paper was partially supported by the National Natural Science Foundation of China (Nos. 61702122, 61751314 and 31560317), the Natural Science Foundation of Guangxi (Nos. 2017GXNSFDA198033 and 2018GXNSFBA281193), the Key Research and Development Plan of Guangxi (No. AB17195055), the Bossco Project of Guangxi University (No. 20190240), the Hunan Provincial Science and Technology Program (No. 2018WK4001) and 111 Project (No. B18059).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors Qingfeng Chen, Zhao Zhe,Wei Lan, Ruchang Zhang, Zhiqiang Wang, Cheng Luo and Yi-Ping Pheobe Chen declare that they have no conflict of interests.
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Author summary: Rapidly increasing evidences show that miRNAs have close relationships with many diseases. Therefore, identifying miRNA-disease interactions may contribute to understand the pathogenesis of diseases. In this paper, we propose a computational framework to predict miRNA-disease interactions based on recommendation method. The experiment results show that our method outperforms other the-state-of-the-art methods.
Rights and permissions
About this article
Cite this article
Chen, Q., Zhe, Z., Lan, W. et al. Identifying MiRNA-disease association based on integrating miRNA topological similarity and functional similarity. Quant Biol 7, 202–209 (2019). https://doi.org/10.1007/s40484-019-0176-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40484-019-0176-7