Abstract
Objective
Maternal plasma is a source of circulating placental nucleic acids. This study was designed to detect aberrantly expressed placental mRNA genes circulating in the maternal plasma of pregnancies affected with fetal conotruncal anomalies (CNTRA) and left-ventricular outflow tract (LVOT) obstruction in the second trimester of pregnancy.
Methods
This was a retrospective monocentric study conducted from 1 Jan 2016 to 31 Dec 2016. NanoString technology was used to identify aberrantly expressed genes, comparing 36 women carrying a fetus with CNTRA or LVOT obstruction to 42 controls at 19–24 weeks of gestation. The genes with differential expression were subsequently tested using real-time polymerase chain reaction. Linear discriminant analysis was used to combine all the mRNA species with discriminant ability for CNTRA and LVOT obstruction. A multivariable receiver operating characteristic (ROC) curve having the estimated discriminant score as an explanatory variable was generated for the two affected groups versus controls.
Results
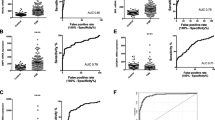
Three genes with differential expression, namely MAPK1, IQGAP1 and Visfatin were found. The ROC curves yielded detection rates of 60 and 62.5% at a false-positive rate of 5% for CNTRA and LVOT, respectively.
Conclusions
These data suggested that molecular screening of CNTRA and LVOT obstruction in the second trimester is feasible. Prospective studies are needed to test the discriminant ability of these genes and to calculate the predictive positive value in the general population.


Similar content being viewed by others
References
Hoffman JI. Congenital heart disease: incidence and inheritance. Pediatr Clin N Am. 1990;37:25–43.
Moller JH, Allen HD, Clark EB, Dajani AS, Golden A, Hayman LL, Lauer RM, Marmer EL, McAnulty JH, Oparil S. Report of the task force on children and youth: American Heart Association. Circulation. 1993;88:2479–86.
Dearani JA, Mavroudis C, Quintessenza J, Deal BJ, Backer CL, Fitzgerald P, et al. Surgical advances in the treatment of adults with congenital heart disease. Curr Opin Pediatr. 2009;21(5):565–72.
Werler M. Additional insights into the etiology of cardiac anomalies. Epidemiology. 2001;12(5):482–4.
Digilio MC, Marino B, Capolino R, et al. Familial recurrence of nonsyndromic congenital heart defects in first degree relatives of patients with deletion 22q11.2. Am J Med Genet A. 2005;134A(2):158–64.
Digilio MC, Marino B. What is new in genetics of congenital heart defects? Front Pediatr. 2016;1(4):120.
Lin AE, Basson CT, Goldmuntz E, Magoulas PL, McDermott DA, McDonald-McGinn DM, McPherson E, Morris CA, Noonan J, Nowak C, Pierpont ME, Pyeritz RE, Rope AF, Zackai E, Pober BR. Adults with genetic syndromes and cardiovascular abnormalities: clinical history and management. Genet Med. 2008;10(7):469–94.
Gong X, Wu X, Ma X, Wu D, Zhang T, He L, Qin S, Li X. Microdeletion and microduplication analysis of chinese conotruncal defects patients with targeted array comparative genomic hybridization. PLoS One. 2013;8(10):e76314.
Digilio MC, Bernardini L, Consoli F, et al. Congenital heart defects in recurrent reciprocal 1q21.1 deletion and duplication syndromes: rare association with pulmonary valve stenosis. Eur J Med Genet. 2013;56(3):144–9.
Wat MJ, Shchelochkov OA, Holder AM, Breman AM, Dagli A, Bacino C, Scaglia F, Zori RT, Cheung SW, Scott DA, Kang SH. Chromosome 8p23.1 deletions as a cause of complex congenital heart defects and diaphragmatic hernia. Am J Med Genet A. 2009;149A(8):1661–77.
Battaglia A, Filippi T, Carey JC. Update on the clinical features and natural history of Wolf–Hirschhorn (4p-) syndrome: experience with 87 patients and recommendations for routine health supervision. Am J Med Genet C Semin Med Genet. 2008;148C(4):246–51.
Restivo A, Piacentini G, Placidi S, Saffirio C, Marino B. Cardiac outflow tract: a review of some embryogenetic aspects of the conotruncal region of the heart. Anat Rec A Discov Mol Cell Evol Biol. 2006;288(9):936–43 (review).
Shinebourne EA, Babu-Narayan SV, Carvalho JS. Tetralogy of Fallot: from fetus to adult. Heart. 2006;92(9):1353–9 (review).
Bonnet D, Coltri A, Butera G, Fermont L, Le Bidois J, Kachaner J, Sidi D. Detection of transposition of the great arteries in fetuses reduces neonatal morbidity and mortality. Circulation. 1999;99(7):916–8.
Kim N, Friedberg MK, Silverman NH. Diagnosis and prognosis of fetuses with double outlet right ventricle. Prenat Diagn. 2006;26(8):740–5.
Duke C, Sharland GK, Jones AM, Simpson JM. Echocardiographic features and outcome of truncus arteriosus diagnosed during fetal life. Am J Cardiol. 2001;88(12):1379–84.
Hornberger LK, Sahn DJ, Kleinman CS, Copel J, Silverman NH. Antenatal diagnosis of coarctation of the aorta: a multicenter experience. J Am Coll Cardiol. 1994;23(2):417–23.
Moerman P, Dumoulin M, Lauweryns J, Van der Hauwaert LG. Interrupted right aortic arch in DiGeorge syndrome. Br Heart J. 1987;58(3):274–8.
Sharland GK, Chita SK, Fagg NL, Anderson RH, Tynan M, Cook AC, Allan LD. Left ventricular dysfunction in the fetus: relation to aortic valve anomalies and endocardial fibroelastosis. Br Heart J. 1991;66(6):419–24.
Better DJ, Apfel HD, Zidere V, Allan LD. Pattern of pulmonary venous blood flow in the hypoplastic left heart syndrome in the fetus. Heart. 1999;81(6):646–9.
Arcelli D, Farina A, Cappuzzello C, et al. Identification of circulating placental mRNA in maternal blood of pregnancies affected with fetal congenital heart diseases at the second trimester of pregnancy: implications for early molecular screening. Prenat Diagn. 2010;30:229–34.
Curti A, Lapucci C, Berto S, Prandstraller D, Perolo A, Rizzo N, Farina A. Maternal plasma mRNA species in fetal heart defects: a potential for molecular screening. Prenat Diagn. 2016;36(8):738–43.
Zhu S, Cao L, Zhu J, et al. Identification of maternal serum microRNAs as novel non-invasive biomarkers for prenatal detection of fetal congenital heart defects. Clin Chim Acta. 2013;424:66–72.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) method. Methods. 2001;25(4):402–8.
Whitehead CL, Walker SP, Tong S. Measuring circulating placental RNAs to non-invasively assess the placental transcriptome and to predict pregnancy complications. Prenat Diagn. 2016;36(11):997–1008.
Saba-El-Leil MK, Vella FD, Vernay B, Voisin L, Chen L, Labrecque N, Ang SL, Meloche S. An essential function of the mitogen-activated protein kinase Erk2 in mouse trophoblast development. EMBO Rep. 2003;4(10):964–8.
Wu WJ, Ma GC, Lin YS, Yeang CH, Ni YH, Li WC, Tsai HD, Shur-Fen Gau S, Chen M. Detection of 22q11.2 microduplication by cell-free DNA screening and chromosomal microarray in fetus with multiple anomalies. Ultrasound Obstet Gynecol. 2016;48(4):530–2.
Sbroggiò M, Bertero A, Velasco S, Fusella F, De Blasio E, Bahou WF, Silengo L, Turco E, Brancaccio M, Tarone G. ERK1/2 activation in heart is controlled by melusin, focal adhesion kinase and the scaffold protein IQGAP1. J Cell Sci. 2011;124(Pt 20):3515–24.
Sbroggiò M, Carnevale D, Bertero A, Cifelli G, De Blasio E, Mascio G, Hirsch E, Bahou WF, Turco E, Silengo L, Brancaccio M, Lembo G, Tarone G. IQGAP1 regulates ERK1/2 and AKT signalling in the heart and sustains functional remodelling upon pressure overload. Cardiovasc Res. 2011;91(3):456–64.
Dou Q, Peng Y, Zhou B, Zhang K, Lin J, Dai X, Zhang L, Rao L. Association of nicotinamide phosphoribosyltransferase (NAMPT) gene polymorphisms and of serum NAMPT levels with dilated cardiomyopathy in a Chinese population. Int J Mol Sci. 2015;16(9):22299–318.
Bobbert P, Kühl U, Poller W, Rauch U, Schultheiss HP, Skurk C. Nicotinamide phosphoribosyltransferase/pre-B-cell colony enhancing factor/visfatin plasma levels and clinical outcome in patients with dilated cardiomyopathy. J Card Fail. 2015;21(4):330–8.
Astern JM, Collier AC, Kendal-Wright CE. Pre-B cell colony enhancing factor (PBEF/NAMPT/Visfatin) and vascular endothelial growth factor (VEGF) cooperate to increase the permeability of the human placental amnion. Placenta. 2013;34(1):42–9.
Rasiah SV, Publicover M, Ewer AK, Khan KS, Kilby MD, Zamora J. A systematic review of the accuracy of first-trimester ultrasound examination for detecting major congenital heart disease. Ultrasound Obstet Gynecol. 2006;28(1):110–6.
Friedberg MK, Silverman NH, Moon-Grady AJ, Tong E, Nourse J, Sorenson B, Lee J, Hornberger LK. Prenatal detection of congenital heart disease. J Pediatr. 2009;155(1):26–31 (31.e1).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
E. Contro, L. Stefani, S. Berto, C. Lapucci, D. Arcelli, D. Prandstraller, A. Perolo, N. Rizzo, A. Farina declare that they have no conflict of interest.
Funding
This study was funded by RFO University of Bologna, Italy to AF and “Corsi di preparazione al parto” University of Bologna, Italy to NR.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Contro, E., Stefani, L., Berto, S. et al. Circulating mRNA in Maternal Plasma at the Second Trimester of Pregnancy: A Possible Screening Tool for Cardiac Conotruncal and Left Ventricular Outflow Tract Abnormalities. Mol Diagn Ther 21, 653–661 (2017). https://doi.org/10.1007/s40291-017-0295-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40291-017-0295-7