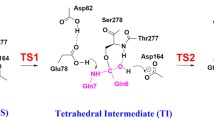
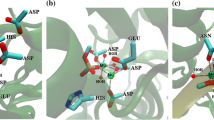
Abstract
Several molecular simulation methods were integrated to investigate the detailed binding process of allophanate to allophanate hydrolase and predict their stable complex structure. The optimal enzyme-substrate complex conformation demonstrates that along with Arg307 and Tyr299, Gly124 is also one of the key anchor residues in the stable complex. The energetic calculation suggests the existence of an intermediate state in the enzyme-substrate binding process. The further atomic-level investigation illuminates that Tyr299, Arg307 and Ser172 can stabilize the substrate in the intermediate state. By this token, the residues Arg307 and Tyr299 function in both binding process and getting stable state.
Similar content being viewed by others
References
Chebrou H., Bigey F., Arnaud A., Galzy P., BBA-Protein Struct. M., 1996, 1298, 285
Shin S., Yun Y. S., Koo H. M., Kim Y. S., Choi K. Y., Oh B. H., J. Biol. Chem., 2003, 278, 24937
Shin S., Lee T. H., Ha N. C., Koo H. M., Kim S. Y., Lee H. S., Kim Y. S., Oh B. H., EMBO J., 2002, 21, 2509
Kim Y. S., Kang S. W., J. Biol. Chem., 1994, 269, 8014
Deutsch D. G., Ueda N., Yamamoto S., Prostag. Leukotr. Ess., 2002, 66, 201
Wu J., Bu W., Sheppard K., Kitabatake M., Kwon S. T., Söll D., Smith J. L., J. Mol. Biol., 2009, 391, 703
Ohtaki A., Murata K., Sato Y., Noguchi K., Miyatake H., Dohmae N., Yamada K., Yohda M., Odaka M., BBA-Proteins Proteom., 2010, 1804, 184
Yasuhira K., Shibata N., Mongami G., Uedo Y., Atsumi Y., Kawashima Y., Hibino A., Tanaka Y., Lee Y. H., Kato D. I., Takeo M., Higuchi Y., Negoro S., J. Biol. Chem., 2010, 285, 1239
Fan C., Chou C. Y., Tong L., Xiang S., J. Biol. Chem., 2012, 287, 9389
Cheng G., Shapir N., Sadowsky M. J., Wackett L. P., Appl. Environ. Microb., 2005, 71, 4437
Kanamori T., Kanou N., Kusakabe S., Atomi H., Imanaka T., FEMS Microbiol. Lett., 2005, 245, 61
Shapir N., Sadowsky M. J., Wackett L. P., J. Bacteriol., 2005, 187, 3731
Strope P. K., Nickerson K. W., Harris S. D., Moriyama E. N., BMC Evolutionary Biology, 2011, 11, 80
Kanamori T., Kanou N., Atomi H., Imanaka T., J. Bacteriol., 2004, 186, 2532
Balotra S., Newman J., French N. G., Briggs L. J., Peat T. S., Scott C., Acta Crystallogr F, 2014, 70, 310
Lin Y., St. Maurice M., Biochemistry, 2013, 52, 690
Fan C., Li Z., Yin H., Xiang S., J. Biol. Chem., 2013, 288, 21422
Balotra S., Newman J., Cowieson N. P., French N. G., Campbell P. M., Briggs L. J., Warden A. C., Easton C. J., Peat T. S., Scott C., Appl. Environ. Microb., 2015, 81, 470
Dong L., Yi Z., Wu Z., Wang H., Zhang A., Chem. J. Chinese Universities, 2015, 36(3), 516
Zhang J. L., Zheng Q. C., Li Z. Q., Zhang H. X., PLoS ONE, 2013, 8, e53811
Zhang J. L., Zheng Q. C., Chu W. T., Zhang H. X., Curr. Comput. Aid. Drug., 2013, 9, 532
Zhang J., Zheng Q., Zhang H., J. Phys. Chem. B, 2010, 114, 7383
Frisch M. J., Trucks G. W., Schlegel H. B., Scuseria G. E., Robb M. A., Cheeseman J. R., Scalmani G., Barone V., Mennucci B., Petersson G. A., Nakatsuji H., Caricato M., Li X., Hratchian H. P., Izmaylov A. F., Bloino J., Zheng G., Sonnenberg J. L., Hada M., Ehara M., Toyota K., Fukuda R., Hasegawa J., Ishida M., Nakajima T., Honda Y., Kitao O., Nakai H., Vreven T., Montgomery Jr. J. A., Peralta J. E., Ogliaro F., Bearpark M. J., Heyd J., Brothers E. N., Kudin K. N., Staroverov V. N., Kobayashi R., Normand J., Raghavachari K., Rendell A. P., Burant J. C., Iyengar S. S., Tomasi J., Cossi M., Rega N., Millam N. J., Klene M., Knox J. E., Cross J. B., Bakken V., Adamo C., Jaramillo J., Gomperts R., Stratmann R. E., Yazyev O., Austin A. J., Cammi R., Pomelli C., Ochterski J. W., Martin R. L., Morokuma K., Zakrzewski V. G., Voth G. A., Salvador P., Dannenberg J. J., Dapprich S., Daniels A. D., Farkas Ö., Foresman J. B., Ortiz J. V., Cioslowski J., Fox D. J., Gaussian 09, Gaussian Inc., Wallingford CT, 2009
Wu G., Robertson D. H., Brooks C. L. I., Vieth M., J. Comput. Chem., 2003, 24, 1549
Discovery Studio, Release 2.5, Accelrys Software Inc., San Diego, 2013
Humphrey W., Dalke A., Schulten K., J. Mol. Graph., 1996, 14, 33
Phillips J. C., Braun R., Wang W., Gumbart J., Tajkhorshid E., Villa E., Chipot C., Skeel R. D., Kalé L., Schulten K., J. Comput. Chem., 2005, 26, 1781
MacKerell A. D., Feig M., Brooks C. L., J. Am. Chem. Soc., 2003, 126, 698
Vanommeslaeghe K., Raman E. P., MacKerell A. D., J. Chem. Inf. Model., 2012, 52, 3155
Vanommeslaeghe K., MacKerell A. D., J. Chem. Inf. Model., 2012, 52, 3144
Becke A. D., J. Chem. Phys., 1993, 98, 5648
Jorgensen W. L., Chandrasekhar J., Madura J. D., Impey R. W., Klein M. L., J. Chem. Phys., 1983, 79, 926
Darden T., York D., Pedersen L., J. Chem. Phys., 1993, 98, 10089
Feller S. E., Zhang Y., Pastor R. W., Brooks B. R., J. Chem. Phys., 1995, 103, 4613
Chovancova E., Pavelka A., Benes P., Strnad O., Brezovsky J., Kozlikova B., Gora A., Sustr V., Klvana M., Medek P., Biedermannova L., Sochor J., Damborsky J., PLoS Comput. Biol., 2012, 8, e1002708
Zhang J. L., Zheng Q. C., Zhang H. X., Chem. Phys. Lett., 2010, 484, 338
Hénin J., Chipot C., J. Chem. Phys., 2004, 121, 2904
Rodriguez-Gomez D., Darve E., Pohorille A., J. Chem. Phys., 2004, 1 20, 3563
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supported by the International Postdoctoral Exchange Fellowship Program(No.20130037), the China Postdoctoral Science Foundation(Nos.2013T60320, 2013M541289), and the National Natural Science Foundation of China(Nos.21203072, 21303068).
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Zhang, Z., Zhang, J., Zheng, Q. et al. Theoretical investigation on binding process of allophanate to allophanate hydrolase. Chem. Res. Chin. Univ. 31, 1023–1028 (2015). https://doi.org/10.1007/s40242-015-5108-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40242-015-5108-0