Abstract
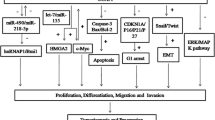
Long noncoding RNA (lncRNA) is a newly identified type of noncoding RNA with a length of more than 200 nucleotides. The latest research shows that lncRNAs play important roles in the occurrence and development of human tumours by acting both as carcinogenic genes and as tumour suppressor genes. LncRNAs plays a role in various biological processes, such as cell growth, apoptosis, migration and invasion. The newly discovered lncRNA DDX11-AS1 is abnormally highly expressed in various malignant tumours, such as hepatocellular carcinoma, colorectal cancer, osteosarcoma, bladder cancer, NSCLC and gastric cancer. DDX11-AS1 mainly regulates the expression of related genes through direct or indirect ways to perform its functions in carcinogenicity. These results indicate that DDX11-AS1 may be a marker or therapeutic target of tumours. This review summarizes the biological function and mechanism of DDX11-AS1 in the process of tumour development.

Similar content being viewed by others
Data availability
Not available.
Abbreviations
- LncRNA:
-
Long noncoding RNA
- DDX11-AS1:
-
DDX11 antisense RNA 1
- EZH2:
-
Enhancer of zeste 2 polycomb repressive complex 2 subunit
- DNMT1:
-
DNA methyltransferase 1
- LATS2:
-
Large tumour suppressor, homologue 2
- CLDN7:
-
Claudin 7
- LAMB3:
-
Laminin subunit beta3
- CDK6:
-
Cyclin-dependent kinase 6
- SPC18:
-
Signalling point code 18
- IRS1:
-
Recombinant insulin receptor substrate 1
- PI3K:
-
Phosphatidylinositol 3-kinase
- AKT/PKB:
-
Protein kinase B
- AFP:
-
Alpha-fetoprotein
- HCC:
-
Hepatocellular carcinoma
- CRC:
-
Colorectal cancer
- GC:
-
Gastric cancer
- NSCLC:
-
Non-small cell lung cancer
- BC:
-
Bladder cancer
- EMT:
-
Epithelial-mesenchymal transition
- Bcl-2:
-
B-cell lymphoma-2
References
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87–108.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2016. CA Cancer J Clin. 2016;66:7–30.
Veneziano D, Di Bella S, Nigita G, Lagana A, Ferro A, Croce CM. Noncoding RNA: current deep sequencing data analysis approaches and challenges. Hum Mutat. 2016;37:1283–98.
Theis M, Paszkowski-Rogacz M, Weisswange I, Chakraborty D, Buchholz F. Targeting human long noncoding transcripts by endoribonuclease-prepared siRNAs. J Biomol Screen. 2015;20:1018–26.
Djebali S, Davis CA, Merkel A, Dobin A, Lassmann T, Mortazavi A, Tanzer A, Lagarde J, Lin W, Schlesinger F, et al. Landscape of transcription in human cells. Nature. 2012;489:101–8.
Lee JT. Epigenetic regulation by long noncoding RNAs. Science. 2012;338:1435–9.
Zhu S, Li W, Liu J, Chen CH, Liao Q, Xu P, Xu H, Xiao T, Cao Z, Peng J, et al. Genome-scale deletion screening of human long non-coding RNAs using a paired-guide RNA CRISPR-Cas9 library. Nat Biotechnol. 2016;34:1279–86.
Zhu J, Kong F, Xing L, Jin Z, Li Z. Prognostic and clinicopathological value of long noncoding RNA XIST in cancer. Clin Chim Acta. 2018;479:43–7.
Han P, Li JW, Zhang BM, Lv JC, Li YM, Gu XY, Yu ZW, Jia YH, Bai XF, Li L, et al. The lncRNA CRNDE promotes colorectal cancer cell proliferation and chemoresistance via miR-181a-5p-mediated regulation of Wnt/beta-catenin signaling. Mol Cancer. 2017;16:9.
Li P, Ruan X, Yang L, Kiesewetter K, Zhao Y, Luo H, Chen Y, Gucek M, Zhu J, Cao H. A liver-enriched long non-coding RNA, lncLSTR, regulates systemic lipid metabolism in mice. Cell Metab. 2015;21:455–67.
Dang Y, Lan F, Ouyang X, Wang K, Lin Y, Yu Y, Wang L, Wang Y, Huang Q. Expression and clinical significance of long non-coding RNA HNF1A-AS1 in human gastric cancer. World J Surg Oncol. 2015;13:302.
Vance KW, Sansom SN, Lee S, Chalei V, Kong L, Cooper SE, Oliver PL, Ponting CP. The long non-coding RNA Paupar regulates the expression of both local and distal genes. EMBO J. 2014;33:296–311.
Lu D, Luo P, Wang Q, Ye Y, Wang B. lncRNA PVT1 in cancer: a review and meta-analysis. Clin Chim Acta. 2017;474:1–7.
Zhang H, Zhu JK. Emerging roles of RNA processing factors in regulating long non-coding RNAs. RNA Biol. 2014;11:793–7.
St Laurent G, Wahlestedt C, Kapranov P. The Landscape of long noncoding RNA classification. Trends Genet. 2015;31:239–51.
Hadji F, Boulanger MC, Guay SP, Gaudreault N, Amellah S, Mkannez G, Bouchareb R, Marchand JT, Nsaibia MJ, Guauque-Olarte S, et al. Altered DNA methylation of long noncoding RNA H19 in calcific aortic valve disease promotes mineralization by silencing NOTCH1. Circulation. 2016;134:1848–62.
Wu R, Su Y, Wu H, Dai Y, Zhao M, Lu Q. Characters, functions and clinical perspectives of long non-coding RNAs. Mol Genet Genom. 2016;291:1013–33.
Zhou M, Zhang XY, Yu X. Overexpression of the long non-coding RNA SPRY4-IT1 promotes tumor cell proliferation and invasion by activating EZH2 in hepatocellular carcinoma. Biomed Pharmacother. 2017;85:348–54.
Akhade VS, Dighe SN, Kataruka S, Rao MR. Mechanism of Wnt signaling induced down regulation of mrhl long non-coding RNA in mouse spermatogonial cells. Nucleic Acids Res. 2016;44:387–401.
Tsang FH, Au SL, Wei L, Fan DN, Lee JM, Wong CC, Ng IO, Wong CM. Long non-coding RNA HOTTIP is frequently up-regulated in hepatocellular carcinoma and is targeted by tumour suppressive miR-125b. Liver Int. 2015;35:1597–606.
Zhang P, Cao L, Fan P, Mei Y, Wu M. LncRNA-MIF, a c-Myc-activated long non-coding RNA, suppresses glycolysis by promoting Fbxw7-mediated c-Myc degradation. EMBO Rep. 2016;17:1204–20.
Lopez-Pajares V. Long non-coding RNA regulation of gene expression during differentiation. Pflugers Arch. 2016;468:971–81.
Song J, Ahn C, Chun CH, Jin EJ. A long non-coding RNA, GAS5, plays a critical role in the regulation of miR-21 during osteoarthritis. J Orthop Res. 2014;32:1628–35.
Li J, Tian H, Yang J, Gong Z. Long noncoding RNAs regulate cell growth, proliferation, and apoptosis. DNA Cell Biol. 2016;35:459–70.
Wu Y, Liu H, Shi X, Yao Y, Yang W, Song Y. The long non-coding RNA HNF1A-AS1 regulates proliferation and metastasis in lung adenocarcinoma. Oncotarget. 2015;6:9160–72.
Cai B, Ma W, Bi C, Yang F, Zhang L, Han Z, Huang Q, Ding F, Li Y, Yan G, et al. Long noncoding RNA H19 mediates melatonin inhibition of premature senescence of c-kit(+) cardiac progenitor cells by promoting miR-675. J Pineal Res. 2016;61:82–95.
Rao A, Rajkumar T, Mani S. Perspectives of long non-coding RNAs in cancer. Mol Biol Rep. 2017;44:203–18.
Malik B, Feng FY. Long noncoding RNAs in prostate cancer: overview and clinical implications. Asian J Androl. 2016;18:568–74.
Shi M, Zhang XY, Yu H, Xiang SH, Xu L, Wei J, Wu Q, Jia R, Wang YG, Lu XJ. DDX11-AS1 as potential therapy targets for human hepatocellular carcinoma. Oncotarget. 2017;8:44195–202.
Li Y, Zhuang W, Huang M, Li X. Long noncoding RNA DDX11-AS1 epigenetically represses LATS2 by interacting with EZH2 and DNMT1 in hepatocellular carcinoma. Biochem Biophys Res Commun. 2019;514:1051–7.
Zhang H, Lin J, Chen J, Gu W, Mao Y, Wang H, Zhang Y, Liu W. DDX11-AS1 contributes to osteosarcoma progression via stabilizing DDX11. Life Sci. 2020;254:117392.
Chen D, Chen J, Gao J, Zhang Y, Ma Y, Wei W, Wei Y. LncRNA DDX11-AS1 promotes bladder cancer occurrence via protecting LAMB3 from downregulation by sponging miR-2355-5p. Cancer Biother Radiopharm. 2020;5:319–28.
Li Q, Wang S, Wu Z, Liu Y. DDX11-AS1exacerbates bladder cancer progression by enhancing CDK6 expression via suppressing miR-499b-5p. Biomed Pharmacother. 2020;127:110164.
Ren Z, Liu X, Si Y, Yang D. Long non-coding RNA DDX11-AS1 facilitates gastric cancer progression by regulating miR-873-5p/SPC18 axis. Artif Cells Nanomed Biotechnol. 2020;48:572–83.
Song W, Qian Y, Zhang MH, Wang H, Wen X, Yang XZ, Dai WJ. The long non-coding RNA DDX11-AS1 facilitates cell progression and oxaliplatin resistance via regulating miR-326/IRS1 axis in gastric cancer. Eur Rev Med Pharmacol Sci. 2020;24:3049–61.
Liu H, Zhang Z, Wu N, Guo H, Zhang H, Fan D, Nie Y, Liu Y. Integrative analysis of dysregulated lncrna-associated ceRNA network reveals functional lncRNAs in gastric cancer. Genes (Basel). 2018;9:303.
Feng X, Yang S, Zhou S, Deng S, Xie Y. Long non-coding RNA DDX11-AS1 promotes non-small cell lung cancer development via regulating PI3K/AKT signalling. Clin Exp Pharmacol Physiol. 2020. https://doi.org/10.1111/1440-1681.13325.
Liao HT, Huang JW, Lan T, Wang JJ, Zhu B, Yuan KF, Zeng Y. Identification of the aberrantly expressed LncRNAs in hepatocellular carcinoma: a bioinformatics analysis based on RNA-sequencing. Sci Rep. 2018;8:5395.
Tian JB, Cao L, Dong GL. Long noncoding RNA DDX11-AS1 induced by YY1 accelerates colorectal cancer progression through targeting miR-873/CLDN7 axis. Eur Rev Med Pharmacol Sci. 2019;23:5714–29.
George J, Patel T. Noncoding RNA as therapeutic targets for hepatocellular carcinoma. Semin Liver Dis. 2015;35:63–74.
Huang JL, Zheng L, Hu YW, Wang Q. Characteristics of long non-coding RNA and its relation to hepatocellular carcinoma. Carcinogenesis. 2014;35:507–14.
Invernizzi F, Vigano M, Grossi G, Lampertico P. The prognosis and management of inactive HBV carriers. Liver Int. 2016;36(Suppl 1):100–4.
Favoriti P, Carbone G, Greco M, Pirozzi F, Pirozzi RE, Corcione F. Worldwide burden of colorectal cancer: a review. Updates Surg. 2016;68:7–11.
Kong J, Sun W, Li C, Wan L, Wang S, Wu Y, Xu E, Zhang H, Lai M. Long non-coding RNA LINC01133 inhibits epithelial-mesenchymal transition and metastasis in colorectal cancer by interacting with SRSF6. Cancer Lett. 2016;380:476–84.
Zhou Y, Chen S, Cheng S, Wei Q, Fathy AH, Shan T. The prognostic value of high LncRNA AFAP1-AS1 expression in various cancers: a systematic review and meta-analysis containing 21 studies. Clin Chim Acta. 2018;481:147–53.
Gong J, Cui Z, Li L, Ma Q, Wang Q, Gao Y, Sun H. MicroRNA-25 promotes gastric cancer proliferation, invasion, and migration by directly targeting F-box and WD-40 Domain Protein 7, FBXW7. Tumour Biol. 2015;36:7831–40.
Li Z, Yu X, Wang Y, Shen J, Wu WK, Liang J, Feng F. By downregulating TIAM1 expression, microRNA-329 suppresses gastric cancer invasion and growth. Oncotarget. 2015;6:17559–69.
Gong Y, Ren J, Liu K, Tang LM. Tumor suppressor role of miR-133a in gastric cancer by repressing IGF1R. World J Gastroenterol. 2015;21:2949–58.
Li Z, Lei H, Luo M, Wang Y, Dong L, Ma Y, Liu C, Song W, Wang F, Zhang J, et al. DNA methylation downregulated mir-10b acts as a tumor suppressor in gastric cancer. Gastric Cancer. 2015;18:43–544.
Jou E, Rajdev L. Current and emerging therapies in unresectable and recurrent gastric cancer. World J Gastroenterol. 2016;22:4812–23.
Wang W, Shen XB, Jia W, Huang DB, Wang Y, Pan YY. The p53/miR-193a/EGFR feedback loop function as a driving force for non-small cell lung carcinoma tumorigenesis. Ther Adv Med Oncol. 2019;11:1758835919850665.
Szpechcinski A, Florczuk M, Duk K, Zdral A, Rudzinski S, Bryl M, Czyzewicz G, Rudzinski P, Kupis W, Wojda E, et al. The expression of circulating miR-504 in plasma is associated with EGFR mutation status in non-small-cell lung carcinoma patients. Cell Mol Life Sci. 2019;76:3641–56.
Osmani L, Askin F, Gabrielson E, Li QK. Current WHO guidelines and the critical role of immunohistochemical markers in the subclassification of non-small cell lung carcinoma (NSCLC): moving from targeted therapy to immunotherapy. Semin Cancer Biol. 2018;52:103–9.
Wang Y, Li Y, He H, Wang F. Circular RNA circ-PRMT5 facilitates non-small cell lung cancer proliferation through upregulating EZH2 via sponging miR-377/382/498. Gene. 2019;720:144099.
Pottgen C, Eberhardt W, Stamatis G, Stuschke M. Definitive radiochemotherapy versus surgery within multimodality treatment in stage III non-small cell lung cancer (NSCLC): a cumulative meta-analysis of the randomized evidence. Oncotarget. 2017;8:41670–8.
Kang DH, Kim JO, Jung SS, Park HS, Chung C, Park D, Lee JE. Efficacy of vinorelbine monotherapy as third- or further-line therapy in patients with advanced non-small-cell lung cancer. Oncology. 2019;97:356–64.
Ploeg M, Aben KK, Kiemeney LA. The present and future burden of urinary bladder cancer in the world. World J Urol. 2009;27:289–93.
Cumberbatch MGK, Jubber I, Black PC, Esperto F, Figueroa JD, Kamat AM, Kiemeney L, Lotan Y, Pang K, Silverman DT, et al. Epidemiology of bladder cancer: a systematic review and contemporary update of risk factors in 2018. Eur Urol. 2018;74:784–95.
Sylvester RJ, van der Meijden AP, Oosterlinck W, Witjes JA, Bouffioux C, Denis L, Newling DW, Kurth K. Predicting recurrence and progression in individual patients with stage Ta T1 bladder cancer using EORTC risk tables: a combined analysis of 2596 patients from seven EORTC trials. Eur Urol. 2006;49:466–5.
Chen J, Wang L, Tang Y, Gong G, Liu L, Chen M, Chen Z, Cui Y, Li C, Cheng X, et al. Maspin enhances cisplatin chemosensitivity in bladder cancer T24 and 5637 cells and correlates with prognosis of muscle-invasive bladder cancer patients receiving cisplatin based neoadjuvant chemotherapy. J Exp Clin Cancer Res. 2016;35:2.
Fernandez-Gomez J, Madero R, Solsona E, Unda M, Martinez-Pineiro L, Gonzalez M, Portillo J, Ojea A, Pertusa C, Rodriguez-Molina J, et al. Predicting nonmuscle invasive bladder cancer recurrence and progression in patients treated with bacillus Calmette-Guerin: the CUETO scoring model. J Urol. 2009;182:2195–203.
Hollenbeck BK, Dunn RL, Ye Z, Hollingsworth JM, Lee CT, Birkmeyer JD. Racial differences in treatment and outcomes among patients with early stage bladder cancer. Cancer. 2010;116:50–6.
Molina JR, Quevedo JF, Furth AF, Richardson RL, Zincke H, Burch PA. Predictors of survival from urachal cancer: a Mayo Clinic study of 49 cases. Cancer. 2007;110:2434–40.
Cepeda MLA, Sosa AJ, Mora G. Telangiectatic osteosarcoma in an infant. Bol Med Hosp Infant Mex. 2017;74:60–4.
Mirabello L, Troisi RJ, Savage SA. Osteosarcoma incidence and survival rates from 1973 to 2004: data from the Surveillance, Epidemiology, and End Results Program. Cancer. 2009;115:1531–43.
Savage SA, Mirabello L. Using epidemiology and genomics to understand osteosarcoma etiology. Sarcoma. 2011;2011:548151.
Yu W, Tang L, Lin F, Yao Y, Shen Z, Zhou X. High-intensity focused ultrasound: noninvasive treatment for local unresectable recurrence of osteosarcoma. Surg Oncol. 2015;24:9–15.
Bacci G, Ferrari S, Bertoni F, Ruggieri P, Picci P, Longhi A, Casadei R, Fabbri N, Forni C, Versari M, Campanacci M. Long-term outcome for patients with nonmetastatic osteosarcoma of the extremity treated at the istituto ortopedico rizzoli according to the istituto ortopedico rizzoli/osteosarcoma-2 protocol: an updated report. J Clin Oncol. 2000;18:4016–27.
Bielack SS, Kempf-Bielack B, Delling G, Exner GU, Flege S, Helmke K, Kotz R, Salzer-Kuntschik M, Werner M, Winkelmann W, et al. Prognostic factors in high-grade osteosarcoma of the extremities or trunk: an analysis of 1,702 patients treated on neoadjuvant cooperative osteosarcoma study group protocols. J Clin Oncol. 2002;20:776–90.
Ottaviani G, Jaffe N. The epidemiology of osteosarcoma. Cancer Treat Res. 2009;152:3–13.
Friebele JC, Peck J, Pan X, Abdel-Rasoul M, Mayerson JL. Osteosarcoma: a meta-analysis and review of the literature. Am J Orthop (Belle Mead NJ). 2015;44:547–53.
Funding
This work was supported by the Anhui Science and Technology Major Project (No 17030801020) and the Chen Feihu Central Support for Local-Provincial Translational Medicine Project in 2017 (2017zhyx31).
Author information
Authors and Affiliations
Contributions
FYB planned and conducted the experiments, analysed the data and wrote the text. WMM conducted the experiments and analysed the data. HS participated in the design of the experiment and contributed to discussing the data and text. PXQ and CFH conceived the study, participated in its design and coordination as well as providing critical review of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Feng, Y., Wu, M., Hu, S. et al. LncRNA DDX11-AS1: a novel oncogene in human cancer. Human Cell 33, 946–953 (2020). https://doi.org/10.1007/s13577-020-00409-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13577-020-00409-8