Abstract
Purpose
Image registration for internal organs and soft tissues is considered extremely challenging due to organ shifts and tissue deformation caused by patients’ movements such as respiration and repositioning. In this paper, we propose a fast deformable image registration method. The purpose of our work is to greatly improve the registration time while maintaining the registration accuracy.
Methods
In this study, we formulate the deformable image registration problem as a quadratic optimization problem that minimizes strain energy subject to the constraints of 3D curves of blood vessel centerlines and point marks. The proposed method does not require iteration and is local minimum free. By using 2nd order B-splines to model the blood vessels in the moving image and a new transformation model, our method provides a closed-form solution that imitates the manner in which physical soft tissues deform, thus guarantees a physically consistent match.
Results
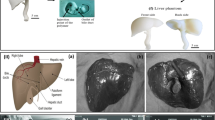
We have demonstrated the effectiveness of our deformable technique in registering MR images of the liver. Validation results show that we can achieve a target registration error (TRE) of 1.29 mm and an average centerline distance error (ACD) of 0.84 ± 0.55 mm.
Conclusions
This technique has the potential to significantly improve registration capabilities and the quality of intraoperative image guidance. To the best of our knowledge, this is the first time that a global analytical solution has been determined for the registration energy function with 3D curve constraints.
Similar content being viewed by others
References
Sotiras A, Davatzikos C, Paragios N. Deformable medical image registration: a survey. IEEE T Med Imaging. 2013; 32(7): 1153–90.
Pluim JPW, Maintz JBA, Viergever MA. Mutual-information based registration of medical images: a survey. IEEE T Med Imaging. 2003; 22(8):986–1004.
Huang X, Ren J, Guiraudon G, Boughner D, Peters TM. Rapid dynamic image registration of the beating heart for diagnosis and surgical navigation. IEEE T Med Imaging. 2009; 28(11):1802–14.
Reinertsen I, Descoteaux M, Siddiqi K, Collins DL. Validation of vessel-based registration for correction of brain shift. Med Image Anal. 2007; 11(4):374–88.
Brock KK. Imaging and image-guided radiation therapy in liver cancer. Semin Radiat Oncol. 2011; 21(4):247–55.
Osorio EMV, Hoogeman MS, Romero AM, Wielopolski P, Zolnay A, Heijmen BJM. Accurate CT/MR vessel-guided nonrigid registration of largely deformed livers. Med Phys. 2012; 39(5):2463–77.
Lange T, Papenberg N, Heldmann S, Modersitzki J, Fischer B, Lamecker H, Schlag PM. 3D ultrasound-CT registration of the liver using combined landmark-intensity information. Int J Comput Assist Radiol Surg. 2009; 4(1):79–88.
Shusharina N, Sharp G. Analytic regularization for landmarkbased image registration. Phys Med Biol. 2012; 57(6):1477–98.
Zitová B, Flusser J. Image registration methods: a survey. Image Vis Comput. 2003; 21(11):977–1000.
Hill DL, Batchelor PG, Holden M, Hawkes DJ. Medical image registration. Phys Med Biol. 2001; 46(3):R1–45.
Maintz JBA, Viergever MA. A survey of medical image registration. Med Image Anal. 1998; 2(1):1–36.
Davis MH, Khotanzad A, Flamig DP, Harms SE. A physicsbased coordinate transformation for 3-D image matching. IEEE T Med Imaging. 1997; 16(3):317–28.
Meinguet J. Multivariate interpolation at arbitrary points made simple. J Appl Math Phys. 1979; 30(2):292–304.
Bookstein FL. Principal warps: thin-plate splines and the decomposition of deformations. IEEE T Pattern Anal Mach Intell. 1989; 11(6):567–85.
Gobbi DG, Comeau RM, Peters TM. Ultrasound/MRI overlay with image warping for neurosurgery. Conf Proc Med Image Comput Comput-Assist Interv. 2000; 1:106–14.
Murphy K, van Ginneken B, Reinhardt JM, Kabus S, Ding K, Deng X, Cao K, Du K, Christensen GE, Garcia V, Vercauteren T, Ayache N, Commowick O, Malandain G, Glocker B, Paragios N, Navab N, Gorbunova V, Sporring J, de Bruijne M, Han X, Heinrich MP, Schnabel JA, Jenkinson M, Lorenz C, Modat M, McClelland JR, Ourselin S, Muenzing SE, Viergever MA, De Nigris D, Collins DL, Arbel T, Peroni M, Li R, Sharp GC, Schmidt-Richberg A, Ehrhardt J, Werner R, Smeets D, Loeckx D, Song G, Tustison N, Avants B, Gee JC, Staring M, Klein S, Stoel BC, Urschler M, Werlberger M, Vandemeulebroucke J, Rit S, Sarrut D, Pluim JP. Evaluation of registration methods on thoracic CT: the EMPIRE10 challenge. IEEE T Med Imaging. 2011; 30(11):1901–20.
Baumhauer M, Feuerstein M, Meinzer HP, Rassweiler J. Navigation in endoscopic soft tissue surgery: perspectives and limitations. J Endourol. 2008; 22(4):751–66.
Huang X, Abdalbari A, Zaheer S, Looi T, Ren J, Drake J. 3D curve constrained deformable registration using a neuro-fuzzy transformation model. Conf Proc IEEE Eng Med Biol Soc. 2012; 1:5294–7.
Commowick O, Arsigny V, Isambert A, Costa J, Dhermain F, Bidault F, Bondiau PY, Ayache N, Malandain G. An efficient locally affine framework for the smooth registration of anatomical structures. Med Image Anal. 2008; 12(4):427–41.
Ciarlet PG. Mathermatical Elasticity, Vol I. North-holland, Amsterdam, Newyork: Oxford; 1988.
Bower AF. Applied Mechanics of Solids. 1st ed. CRC Press; 2009.
Fitzpatrick JM, West JB, Maurer CR Jr. Predicting error in rigidbody point-based registration. IEEE T Med Imaging. 1998; 17(5):694–702.
Huang X, Babyn PS, Looi T, Kim PCW. A novel hybrid model for deformable image registration in abdominal procedures. Proc SPIE 7964 Med Imaging; 2011. doi: 10.1117/12.878068.
3D Slicer. http://www.slicer.org/. Accessed 2 Mar 2016.
VMTK. http://www.vmtk.org/. Accessed 2 Mar 2016.
Huang X, Zaheer S, Abdalbari A, Looi T, Ren J, Drake J. Extraction of liver vessel centerlines under guidance of patientspecific models. Conf Proc IEEE Eng Med Biol Soc. 2012; 1:2347–50.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Huang, X., Ren, J., Abdalbari, A. et al. Vessel-based fast deformable registration with minimal strain energy. Biomed. Eng. Lett. 6, 47–55 (2016). https://doi.org/10.1007/s13534-016-0213-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13534-016-0213-7