Abstract
Single-stranded telomeric DNA tends to fold into G-quadruplexes made of a stack of G-quartets. Ligands that target G-quadruplex DNA have received considerable attention because of their potential anticancer activity. In this paper, we designed a fluorescence resonance energy transfer system to identify drugs that form G-quadruplexes with human telomeric DNA in a homogeneous medium using gold nanorods (GNRs) as fluorescence quencher. Fluorescein-tagged human telomeric DNA (F-GDNA) adsorbed on the surface of GNRs, which can quench the fluorescence of F-GDNA. In the presence of G-quadruplex-binding drugs, the formation of G-quadruplex further quenched the fluorescence of F-GDNA, which can be utilized to select G-quadruplex-binding ligands. Thirteen natural drugs were studied by this proposed method. G-quadruplex formation induced by drugs was investigated by circular dichroism measurements. Results of research indicated that it offers a simple, sensitive and effective method to identify ligands with potential anticancer activity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Telomeres are essential for the stability and replication of eukaryotic chromosomes [1, 2]. Telomeres generally consist of many tandem repeats of guanine-rich sequence, and a short 3′-overhang single-stranded sequence tends to form a four base-paired planar structure termed G-quadruplex [3, 4]. In vitro, G-quadruplex blocks the binding of telomerase and inhibits telomere extension. Therefore, ligands of G-quadruplex have received great attention because quadruplex-binding ligands have potential applications in genomic studies as well as therapeutic purposes. Methods for practical, rapid, simple, and sensitive selection of G-quadruplex-binding ligands are in great demand since they can be applied to several fields, including genetics, pathology, and especially in designing and screening of antitumor drugs. In recent years, a large effort has been expended in developing techniques for telomere and G-quadruplex research, including X-ray diffraction, nuclear magnetic resonance spectroscopy, fluorescence, circular dichroism (CD), mass spectrometry, electrochemistry and ultraviolet-visible (UV-vis) spectroscopy [5–10]. Due to its sensitivity and flexibility, fluorescence spectroscopy, particularly resonance energy transfer technique (FRET), has been explored as powerful tool to study G-quadruplexes. On the basis of the conformational transition of G-DNA, it is easy to exploit FRET technique to study stability and molecular mechanism of G-DNA with its ligands [11–17]. Simonsson et al. reported a simple method to study G-quadruplex using fluorescence resonance energy transfer methods [18]. In general, the DNA strand was labeled with fluorophore donor and acceptor or fluorophore and quencher at its 5′ and 3′ ends, respectively. They require dual labeling on the same DNA strand. A simple method would be desired to be developed for selecting ligands of G-DNA.
With the development of nanotechnology, a number of nanomaterials have received widespread interest in biotechnological systems for diagnostic application and biological imaging. Owing to their unique optical properties and ease of bioconjugation, gold nanomaterial, such as gold nanoparticles and gold nanorods, is a type of promising nanomaterial playing important roles in the design of chemical and biochemical sensing for DNA hybridization [19–21], metal ions [22, 23], antibodies [24, 25], and for cancer cell imaging etc [26, 27]. GNRs are normally passivated by positively charged surfactants, which give not only high stability but also positively charged surface. Therefore, GNRs could easily adsorb many biomacromolecules that possess negative charges through electrostatic attractions, such as DNA, peptide, and even cells. On the basis of the quenching capability of gold nanorods to the fluorescent dye, Gou et al. took advantage of GNRs-based FRET for monitoring human telomere DNA hybridization and G-quadruplex formation [28].
In this paper, we have designed a simple FRET system for selecting G-quadruplex-binding ligands using GNRs as a fluorescence quencher. The fluorescence intensity of fluorescein-tagged human telomeric DNA (F-GDNA) quenched by GNRs after addition of GNRs into F-GDNA solution. When quadruplex-binding ligands are added, the formation of G-quadruplexes increases the space charge density around the DNA macromolecule, which is expected to strengthen the interaction between the GNRs and the DNA. Therefore, the florescence of DNA probe was further quenched. In this way, G-quadruplex-binding ligands can be selected without labeling drugs. Furthermore, the DNA strand needs only one dye labeled, leading to less laborious and more cost-effective synthesis. Here, 13 natural drugs were studied by this proposed method and experimental results indicated that the FRET approach offers a simple, sensitive, and effective method to identify ligands with potential anticancer activity.
Experimental section
Chemicals
Unless otherwise indicated, all reagents and solvents were purchased in their highest available purity and used without further purification. HAuCl4·4H2O was purchased from Sigma (USA). Daidzein, apigenin, quercetin, luteolin, kaempferol, chrysin, emodin, aloe-emodin, chrysophanol, rhein, berberine, colchicine, and matrine were purchased from Nanjing TCM Institute of Chinese Material Medical (Nanjing, China). Drugs and other reagents were commercially available analytical reagent grade. Fluorescein (FAM)-labeled probe DNA is an oligonucleotide mimicking the human telomeric repeat (5′-FAM-TTAGGGTTAGGGTTAGGGTTAGGG-3′, F-GDNA). F-GDNA and FAM-labeled control DNA (F-DNA, 5′-FAM-GTT CATGCCGCCCATGCTCG-3′) were synthesized by Shanghai Sangon Biotechnology Co. (Shanghai, China) and used without further purification. The oligonucleotide stock solutions (25 μM) were prepared with a TE buffer (10 mM Tris-HCl, 1 mM EDTA, pH 7.2) and kept frozen. Millipore Milli-Q (18 MΩ cm) water was used in all experiments.
Instrumentation
Fluorescence spectra were recorded on a Hitachi F-7000 fluorescence spectrophotometer. CD spectra were measured on a Chirascan Circular Dichroism Spectrometer (Applied Photophysics Ltd, England, UK). UV-visible adsorption spectra were recorded on UV-2450 UV-vis spectrophotometer (Shimadzu Corporation, Kyoto, Japan). Transmission electron microscopy (TEM) images were obtained using a JEM-2100 transmission electron microscope.
Synthesis of gold nanorods
Gold nanorods were synthesized by using a seed-mediated, surfactant-assisted growth method in a two-step procedure [29, 30]. Briefly, colloidal gold seeds were first prepared by mixing aqueous solutions of hexadecylcetyltri-methylammonium bromide (CTAB, 0.1 M, 7.5 mL) and hydrogen tetrachloroaurate(III) hydrate (1%, 0.098 mL). Freshly prepared aqueous solution of sodium borohydride (0.01 M, 0.6 mL) was then added. The colloidal gold seeds solution(0.215 mL) were then injected into an aqueous growth solution of CTAB (0.1 M, 47.6 mL), hydrogen tetrachloroaurate(III) hydrate (1%, 0.788 mL), silver nitrate (0.01 M, 0.3 mL) and freshly prepared ascorbic acid (0.1 M, 0.32 mL). The nanorods were purified by several cycles of suspension in ultrapure water, followed by centrifugation. They were isolated in the precipitate and excess CTAB was removed in the supernatant. Then, it was stored in a refrigerator at 4°C before being used. The nanorods were characterized by absorption spectroscopy.
Fluorescence experiments
First, the fluorescence spectra of FAM-labeled GDNA were recorded on a fluorometer (F-7000, Hitachi) with excitation at 480 nm and emission range from 500 to 600 nm. Then, a certain volume of GNRs solution was added into the FAM-labeled GDNA solution. The fluorescence of this mixture is recorded within 10 min. For the study of drug–GDNA interactions, drug solution with a certain concentration was added into above mixture for fluorescence detection.
CD Spectroscopy
The CD spectra of DNA oligonucleotides were measured for 5 μM DNA total strand concentration using a Chirascan Circular Dichroism Spectrometer (Applied Photophysics Ltd, England, UK). CD spectra were recorded using a quartz cell of 1-mm optical path length and an instrument scanning speed of 100 nm/min with a response time of 2 s at room temperature. CD spectra were obtained by taking the average of three scans made from 200 to 320 nm. All DNA samples at a final concentration of 5 μM were dissolved in Tris-buffer (10 mM, pH 7.2) and heated to 90°C for 5 min, gradually cooled to room temperature, and incubated at 4°C overnight.
UV measurements
Typically, the UV absorb spectra of GNRs were recorded on UV-vis spectrophotometer (Shimadzu Corporation, Kyoto, Japan). Then, a certain quantity of GDNA was added into GNRs suspension solution. After thorough mixing, a suitable ligand was added to the mixture. The solution is vortexed thoroughly and used for the UV-vis absorption spectra measurements experiment approximately 10 min after mixing.
Results and discussion
FRET between F-GDNA and GNRs
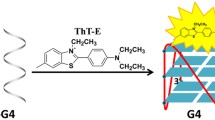
In this paper, we have designed a GNR-based FRET assay for selecting quadruplex-binding ligands using human telomeric repeat sequence as recognition probe. Figure 1 shows the sensing scheme of this approach. Briefly, when a dye-labeled olignucleotide single-stranded probes sticks to the GNRs, the attendant proximity of the dye to the gold leads to fluorescence quenching of the dye. In the presence of quadruplex-binding ligands, the formation of G-quadruplexes strengthened the interaction between probe DNA and GNRs, resulting in the decrease in fluorescence intensity. The feasibility of this FRET protocol was first investigated by adding emodin into the mixed solution of FAM-labeled GDNA and GNRs. Curve a in Fig. 2a shows the fluorescence spectra of 10 nM FAM-labeled telomere DNA (5′-FAM-TTAGGGTTAGGGTTAGGGTTAGGG-3′, F-GDNA) in TE buffer at pH 7.2. It was found that the fluorescence intensity is obviously decreased by ∼27.5% after mixed with gold nanorods (curve b), which indicated that FRET occurred between fluorescent dye and GNRs. That is, as GNRs was added to the F-GDNA solution, F-GDNA was adsorbed onto the surface of the positively charged GNRs because of the electrostatic interaction between the anionic backbone phosphates of the oligonucleotides and the cationic surfactant bilayer around the GNRs [28]. Upon addition of the emodin, the fluorescence intensity of F-GDNA/GNRs mixture was in an obvious decline. The observed change induced by drug addition can be ascribed to the strengthened interaction between FGDNA and GNRs due to the formation of G-quadruplex in the presence of emodin because G-quadruplex DNA has higher charge density than the ssDNA [24].
Fluorescence emission spectra of FAM-labeled GDNA at different experiment conditions. a a FGDNA in TE, b FGDNA/GNRs, c–g FGDNA/GNRs + emodin, the concentration of emodin is 5, 10, 50, 100, and 500 nM, respectively. b a FGDNA, b FGDNA/GNRs, c–g FGDNA/GNRs + matrine, the concentration of matrine is 5, 10, 50, 100, and 500 nM, respectively. Fluorescence intensity was recorded at 520 nm with an excitation wavelength of 480 nm
To exclude other possibilities that lead to the fluorescence reduction, the effect of ligand on fluorescence intensity of F-GDNA was first studied in the TE buffer (Figure S1, Supporting Information (SI)). Fluorescence intensity of F-GDNA has little change with the increasing concentration of emodin in the absence of GNRs. Figure S1 indicated that fluorescence decrease resulted from the electrostatic interaction between FGDNA and GNRs.
The FRET system used GNRs as fluorescent quenchers by noncovalent adsorption of DNA probes onto the surface of GNRs. A control experiment with a negative G-quadruplex binding ligand was performed to prove the specific recognition interaction between F-GDNA and emodin. According to our previous work, matrine was chosen as a negative G-quadruplex binding ligand. It is apparent from Fig. 2b that the fluorescent signal remained steady after the addition of matrine to the F-GDNA/GNRs solution. Control F-DNA was designed to further verify the specific molecular recognition between F-GDNA and emodin. The fluorescence intensity of F-DNA obviously decreased after mixed with GNRs. However, there is little difference in fluorescence intensity of FDNA/GNRs solution before and after adding emodin into the FDNA/GNRs (Figure S2, SI). All these control experiments indicated that the fluorescence reduction resulted from the G-quadruplex formation in the presence of emodin, which enhanced the electrostatic interaction between DNA and GNRs.
To further validate the feasibility of this FRET system, the relationship between ligand concentration and fluorescence reduction was studied. From curve c to g as shown in Fig. 2a, the fluorescence of F-GDNA/GNRs solution continuously decreased with the concentration of emodin ranged from 5 to 500 nM, respectively. Upon the addition of emodin, the fluorescence of the F-GDNA/GNRs solution decreases significantly, indicating that ligand of GDNA can modulate the energy transfer between fluorophore and GNRs. That is, G-quadruplex formation has been on the increase with the rise of emodin concentration, leading to stronger electrostatic interaction between DNA and GNRs. However, the fluorescent signal has little change with the increasing concentration of matrine (Fig. 2b). The result further verified that the fluorescence reduction of F-GDNA/GNRs solution in the presence of emodin was attributable to specific binding between F-GDNA and emodin. Therefore, this FRET assay is feasible to select the quadruplex-binding ligands.
Identification of quadruplex formation
CD is a sensitive technology able to study the configuration inversion of DNA and can be used to conveniently monitor quadruplex formation. To understand whether G-quadruplex formed in the presence of selected ligands, we measured CD spectra. As can be seen from Fig. 3, the CD spectra of human telomeric DNA in the absence of emodin are characterized by a positive peaks at 255 nm and a negative peak at 234 nm (curve a). With the addition of emodin, we observed a concentration-dependent enhancement of the positive peak around 295 nm. At the same time, the positive peaks at 255 nm decreased and a negative peak at about 265 nm start to appear, which indicated G-quadruplex formation. That is, the human telomeric DNA underwent a conformational change and formed the G-quadruplex structure. These spectral changes were not observed when matrine was added to Human24 DNA (Figure S3 of the Supporting Information) [11, 31]. The result is in accordance with FRET measurements. These control experiments indicated that the proposed method is a reliable method to select G-quadruplex-binding ligands.
Mechanism investigation
It is known that the UV adsorption spectra of GNRs displays two bands assigned to the transversal and longitudinal modes of electronic oscillations, and the locations depends on the aspect ratio. As shown in Fig. 4, the two UV absorb bands for our prepared GNRs are located around 518 and 783 nm. When DNA was added to the GNRs suspension, they were adsorbed onto the surface of the positively charged GNRs because of the electrostatic interactions between the anionic backbone phosphates of oligonucleotides and the cationic surfactant bilayer around the nanorods. However, the electrostatic interaction is not strong enough to change the suspension state of GNRs. So, there is little change in the UV absorb spectra. It is worth noticed that the longitudinal adsorption obviously decreased after adding emodin to the mixture of GDNA and GNRs suspension. The decrease of longitudinal adsorption indicated the aggregation of GNRs in solution [19]. These variations of adsorption features resulted from G-quadruplex formation, which strengthened the electrostatic interaction between DNA and GNRs. That is, the strong electrostatic interaction induces the aggregation of GNRs. The sensing mechanism has been further verified by TEM images. From images a and b in Fig. 5 we can see, GNRs are dispersed in the aqueous medium in the absence and presence of GDNA. However, GNRs obviously aggregated after that emodin was added into the suspension (image c), which is accordance with the UV-visible adsorption measurements. This is consistent with observations with fluorescence measurements.
FRET selection of G-quadruplex-binding ligands
A large number of ligands reportedly interact with GDNA to form G-quadruplexes. Folding of telomeric DNA into G-quadruplexes inhibits telomerase by locking the single-stranded telomeric substrate into an inactive conformation that is no longer recognized, nor elongated, by the enzyme [32]. So identification of quadruplex-binding ligands is of great theoretical and practical importance. Here, 13 traditional Chinese medicine monomers were investigated as potential ligands using the proposed method. Daidzein, apigenin, quercetin, luteolin, kaempferol and chrysin belong to flavonoids. Emodin, aloe-emodin, chrysophanol, and rhein belong to anthraquinone. Berberine, colchicine, and matrine belong to alkaloid. They possess anti-allergic, anti-inflammatory, anti-microbial, and anticancer activities. In Fig. 6, the influences of different drugs on fluorescence intensity were demonstrated by comparing the ratios of fluorescence intensity (F/F 0) of FGDNA/GNRs mixture, where F and F 0 are the fluorescence intensity of FGDNA/GNRs in the presence and absence of these drugs, respectively. As shown in Fig. 6, the fluorescence intensity of FGDNA/GNRs further decreased after incubating with daidzein, apigenin, quercetin, luteolin, kaempferol, chrysin, emodin, aloe-emodin, chrysophanol, rhein, and berberine. However, the fluorescence intensity hardly changed after interacting with colchicine and matrine. That is, these natural drugs that enhanced the FRET efficiency between GNRs and FGDNA are G-quadruplex-binding ligands.
Bar chart of fluorescence response of drugs to F-GDNA/GNRs, where F 0 and F are the F-GDNA/GNRs intensities in the absence and presence of drugs. Black bars the fluorescence response of F-GDNA/GNRs in the absence of drug. Red bars FRET efficiency of different drugs measured in buffer solution. Histograms and error bars represent average values and standard deviations, respectively. Relative standard deviations are all less than 5%
Conclusion
In this paper, we have developed a fluorescence method for selection of G-quadruplex-binding ligands by GNRs-based FRET assay without expensive reagents and complicated modification. Thirteen natural drugs were studied by the proposed method, indicating that daidzein, apigenin, quercetin, luteolin, kaempferol, chrysin, emodin, aloe-emodin, chrysophanol, rhein, and berberine can induce the human telomeric DNA to form G-quadruplex structure. The sensing mechanism has been verified by the CD, UV, and TEM measurements. The major advantages of this assay are its speed, simplicity, excellent specificity, and convenience. This method provides a useful method to rapid screening antitumor drugs and is of great theoretical and practical importance.
References
Neidle S, Parkinson GN (2003) The structure of telomeric DNA. Curr Opin Struct Biol 13:275–283
Chang CC, Wu JY, Chien CW, Wu WS, Liu H, Kang CC, Yu LJ, Chang TC (2003) A fluorescent carbazole derivative: high sensitivity for quadruplex DNA. Anal Chem 75:6177–6183
Davis JT (2004) G-quartets 40 years later: from 5′-GMP to molecular biology and supramolecular chemistry. Angew Chem Int Ed 43:668–698
Burge S, Parkinson GN, Hazel P, Todd AK, Neidle S (2006) Quadruplex DNA: sequence, topology and structure. Nucleic Acids Res 34:5402–5415
Yu HQ, Miyoshi D, Sugimoto N (2006) Characterization of structure and stability of long telomeric DNA G-quadruplexes. J Am Chem Soc 128:15461–15468
Phan AT, Kuryavyi V, Ma JB, Faure A, Andréola ML, Patel DJ (2005) An interlocked dimeric parallel-stranded DNA quadruplex: a potent inhibitor of HIV-1 integrase. PANS 102:634–639
Kan ZY, Yao Y, Wang P, Li XH, Hao YH, Tan Z (2006) Molecular crowding induces telomere G-quadruplex formation under salt-deficient conditions and enhances its competition with duplex formation. Angew Chem Int Ed 45:1629–1632
Schultze P, Hud NV, Smith FW, Feigon J (1999) The effect of sodium, potassium and ammonium ions on the conformation of the dimeric quadruplex formed by the Oxytricha nova telomere repeat oligonucleotide D(G4T4G4). Nucleic Acids Res 27:3018–3028
Ho HA, Leclerc M (2004) Optical sensors based on hybrid aptamer/conjugated polymer complexes. J Am Chem Soc 126:1384–1387
Mergny JL, Maurizot JC (2001) Fluorescence resonance energy transfer as a probe for G-quartet formation by a telomeric repeat. ChemBioChem 2:124–132
Jin Y, Li HY, Bai JY (2009) Homogeneous selecting of a quadruplex-binding ligand-based gold nanoparticle fluorescence resonance energy transfer assay. Anal Chem 81:5709–5715
Juskowiak B (2006) Analytical potential of the quadruplex DNA-based FRET probes. Anal Chim Acta 568:171–180
Ueyama H, Takagi M, Takenaka S (2002) A novel potassium sensing in aqueous media with a synthetic oligonucleotide derivative. Fluorescence resonance energy transfer associated with guanine quartet-potassium ion complex formation. J Am Chem Soc 124:14286–14287
He F, Tang YL, Wang S, Li YL, Zhu DB (2005) Fluorescent amplifying recognition for DNA G-quadruplex folding with a cationic conjugated polymer: a platform for homogeneous potassium detection. J Am Chem Soc 127:12343–12346
Bourdoncle A, Torres AE, Gosse C, Lacroix L, Vekhoff P, Saux TL, Jullien L, Mergny JL (2006) Quadruplex-based molecular beacons as tunable DNA probes. J Am Chem Soc 128:11094–11105
Ying LM, Green JJ, Li HT, Klenerman D, Balasubramanian S (2003) Studies on the structure and dynamics of the human telomeric G quadruplex by single-molecule fluorescence resonance energy transfer. Proc Nat Acad Sci USA 100:14629–14634
Lee JY, Okumus B, Kim DS, Ha T (2005) Extreme conformational diversity in human telomeric DNA. Proc Nat Acad Sci USA 102:18938–18943
Simonsson T, Sjöback R (1999) DNA tetraplex formation studied with fluorescence resonance energy transfer. J Biol Chem 274:17379–17383
He W, Huang CZ, Li YF, Xie JP, Yang RG, Zhou PF, Wang J (2008) One-step label-free optical genosensing system for sequence-specific DNA related to the human immunodeficiency virus based on the measurements of light scattering signals of gold nanorods. Anal Chem 80:8424–8430
Gao Q, Zhang WY, Guo YY, Qi HL, Zhang CX (2011) Highly sensitive impedimetric sensing of DNA hybridization based on the target. Electrochem Commun 13:335–337
Chen CK, Shiang YC, Huang CC, Chang HT (2011) Using self-assembled aptamers and fibrinogen-conjugated gold nanoparticles to detect DNA based on controlled thrombin activity. Biosens Bioelectron 26:3464–3468
Li W, Nie Z, He KY, Xu XH, Li Y, Huang Y, Yao SZ (2011) Simple, rapid and label-free colorimetric assay for Zn2+ based on unmodified gold nanoparticles and specific Zn2+ binding peptide. Chem Commun 47:4412–4414
Wang H, Wang YX, Jin JY, Yang RH (2008) Gold nanoparticle-based colorimetric and “Turn-On” fluorescent probe for mercury(II) ions in aqueous solution. Anal Chem 80:9021–9028
Chang JY, Wu HM, Chen H, Ling YC, Tan WH (2005) Oriented assembly of Au nanorods using biorecognition system. Chem Commun 8:1092–1094
Park HS, Agarwal A, Kotov NA, Lavrentovich OD (2008) Controllable side-by-side and end-to-end assembly of Au nanorods by lyotropic chromonic materials. Langmuir 24:13833–13837
Wang HF, Huff TB, Zweifel DA, He W, Low PS, Wei A, Cheng JX (2005) In vitro and in vivo two-photon luminescence imaging of single gold nanorods. Proc Nat Acad Sci USA 102:15752–15756
Ha SH, Carson A, Agarwal A, Kotov NA, Kim K (2011) Detection and monitoring of the multiple inflammatory responses by photoacoustic molecular imaging using selectively targeted gold nanorods. Biomedical Optics Express 2:645–657
Gou XC, Liu J, Zhang HL (2010) Monitoring human telomere DNA hybridization and G-quadruplex formation using gold nanorods. Anal Chim Acta 668:208–214
Nikoobakht B, El-Sayed MA (2003) Preparation and growth mechanism of gold nanorods (NRs) using seed-mediated growth method. Chem Mater 15:1957–1962
Murphy CJ, Sau TK, Gole AM, Orendorff CJ, Gao JX, Gou LF, Hunyadi SE, Li T (2005) Anisotropic metal nanoparticles: synthesis, assembly, and optical applications. J Phys Chem B 109:13857–13870
Jin Y, Li HY, Liu P (2010) Label-free electrochemical selection of G-quadruplex-binding ligands based on structure switching. Biosens Bioelectron 25:2669–2674
Xu Y, Suzuki Y, Ito K, Komiyama M (2010) Telomeric repeat-containing RNA structure in living cells. Proc Nat Acad Sci USA 107:14579–14584
Acknowledgment
This work was financially supported by the National Natural Science Foundation of China (no. 21075079 and no. 20705017).
Open Access
This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
FAM-labeled GDNA with the addition of emodin. From a to g, the concentration of emodin is 0 nM, 5 nM, 10 nM, 50 nM, 100 nM, and 500 nM, respectively. The concentration of F-GDNA was 10 nM (DOC 116 kb)
ESM 2
FAM-labeled control DNA at different experiment conditions. a FDNA in TE buffer, b FDNA/GNRs, c–g FDNA/GNRs + emodin. The concentration of emodin is 5 nM, 10 nM, 50 nM, 100 nM, 500 nM, and 1 μΜ, respectively. The excitation wavelength was at 480 nm and emission was monitored at 520 nm (DOC 118 kb)
ESM 3
CD spectra of the Human24 DNA (5 μM) in 10 mM Tris-HCl buffer: a without matrine, b with 10 μM matrine, c with 30 μM matrine, d with 60 μM matrine (DOC 136 kb)
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Jin, Y., Chen, G. & Wang, Y. Gold nanorod-based FRET assay for selection of G-quadruplex-binding ligands. Gold Bull 44, 163–169 (2011). https://doi.org/10.1007/s13404-011-0022-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13404-011-0022-9