Abstract
Purpose
Highly aggressive adult soft tissue sarcomas (STS), i.e., leiomyosarcomas (LMS) and undifferentiated pleomorphic sarcomas (UPS), present complex genomic anomalies and overall 5-year survival rates of 20 to 40%. Here, we aimed to identify new biomarkers that may be employed to improve the treatment of non-translocation STS patients. We validated 12 miRNAs implicated in tumor development using primary STS samples and selected miR-152 for further analysis in STS-derived cell lines.
Methods
59 primary STS samples (27 LMS and 32 UPS) and 10 matched normal control tissues were included in the study, as well as 3 STS-derived cell lines (HT1080, SW872 and SKLMS1) and a normal control mesenchymal cell line (hMSC). miRNA expression analyses were performed using a TaqMan microRNA Array platform and qRT-PCR (miR-152), respectively. The expression levels of the putative miR-152 targets MET and KIT were assessed using qRT-PCR and immunohistochemistry on tissue microarrays, respectively. In addition, various functional analyses were performed before and after miR-152 transfection into SKLMS1 cells.
Results
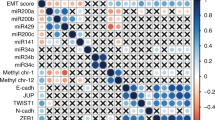
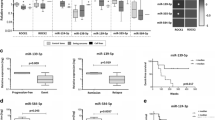
We found that 12 pre-selected miRNAs were down-regulated in primary STS tumor samples compared to its normal control samples. A statistically significant miR-152 down-regulation was found to be accompanied by high MET and KIT mRNA levels in both the primary samples and the STS-derived cell lines tested. miR-152 transfection in SKLMS1 cells led to a reduction in KIT and MET mRNA and protein levels which, in turn, was associated with a transient down-regulation of the PI3K/AKT pathway, a transient decrease in cell growth, and a transient increase in both apoptotic and S-phase cells.
Conclusions
Our data indicate that over-expression of MET and KIT in primary STS samples and its derived cell lines is associated with miR-152 down-regulation. This shift may play a role in STS development and, thus, may be used to identify patients at risk. The effect of MET down-regulation on downstream signaling pathways, such as the PI3K/AKT pathway, may provide a basis for the future design of novel STS treatment strategies.






Similar content being viewed by others
References
P. Rutkowski, I. Lugowska, Follow-up in soft tissue sarcomas. Memory 7, 92–96 (2014)
M. Miettinen, J. F. Fetsch, Evaluation of biological potential of smooth muscle tumours. Histopathology 48, 97–105 (2006)
P. Picci, M. Manfrini, N. Fabbri, M. Gambarotti, D. Vanel, Atlas of Muscoloskeletal tumors and Tumorlike lesion (Springer, New York, 2014), pp. 311–313
C. D. M. Fletcher, J. A. Bridge, P. Hogendoorn, F. Mertens, WHO classification of tumours of soft tissue and bone, 4thEdition edn. (IARC Press, Lyon, 2013)
A. F. Nascimenàkto, C. P. Raut, Diagnosis and management of pleomorphic sarcomas (so-called "MFH") in adults. J. Surg. Oncol. 97, 330–339 (2008)
P. Dei Tos, Classification of pleomorphic sarcomas: where are we now? Histopathology 48, 51–62 (2006)
L. J. Helman, P. Meltzer, Mechanisms of sarcoma development. Nat. Rev. Cancer 3, 685–694 (2003)
T. Niini, L. Lahti, F. Michelacci, S. Ninomiya, C.M. Hattinger, M. Guled, T. Bohling, P. Picci, M. Serra, S. Knuutila, Array comparative genomic hybridization reveals frequent alterations of G1/S checkpoint genes in undifferentiated pleomorphic sarcoma of bone. Genes Chromosom. Cancer 50, 291–306 (2011)
M. Guled, L. Pazzaglia, I. Borze, N. Mosakhani, C. Novello, M. S. Benassi, S. Knuutila, Differentiating soft tissue leiomyosarcoma and undifferentiated pleomorphic sarcoma: a miRNA analysis. Genes Chromosom. Cancer 53, 693–702 (2014)
F. Chibon, P. Lagarde, S. Salas, G. Perot, V. Brouste, F. Tirode, C. Lucchesi, A. de Reynies, A. Kauffmann, B. Bui, P. Terrior, S. Bonvalot, A. LeCesne, D. Vince-Panchere, J. Y. Blay, F. Collin, L. Guillou, A. Leroux, J. M. Coindre, A. Aurias, Validated prediction of clinical outcome in sarcomas and multiple types of cancer on the basis of a gene expression signature related to genome complexity. Nat. Med. 16, 781–787 (2010)
S. L. Romero-Cordoba, I. Salido-Guadarrama, M. Rodriguez-Dorantes, A. Hidalgo-Miranda, miRNA biogenesis: biological impact in the development of cancer. Cancer Biol. Ther. 15, 1444–1455 (2014)
M. Vitiello, A. Tuccoli, L. Poliseno, Long non-coding RNAs in cancer: implications for personalized therapy. Cell. Oncol. 38, 17–28 (2015)
A. Ferraro, Altered primary chromatin structures and their implications in cancer development. Cell. Oncol. 39, 195–210 (2016)
V. Taucher, H. Mangge, J. Haybaeck, Non coding RNAs in pancreatic cancer: challenges and opportunities for clinical application. Cell. Oncol. 39, 295–318 (2016)
G. J. Nuovo, T. D. Schmittgen, Benign metastasizing leiomyoma of the lung: clinicopathologic, immunohistochemical, and micro-RNA analyses. Diagn. Mol. Pathol. 17, 145–150 (2008)
T. Greither, P. Würl, L. Grochola, G. Bond, M. Banche, M. Kappler, C. Lautenschläger, H. J. Holzhausen, S. Wach, A. W. Eckert, H. Taubert, Expression of microRNA 210 associates with poor survival and age of tumor onset of soft-tissue sarcoma patients. Int. J. Cancer 130, 1230–1235 (2012)
E. Gherardi, W. Birchmeier, C. Birchmeier, G. Vande Woude, Targeting MET in cancer: rationale and progress. Nat. Rev. Cancer 12, 89–103 (2012)
A. Esquela-Kerscher, F. J. Slack, Oncomirs - microRNAs with a role in cancer. Nat. Rev. Cancer 6, 259–269 (2006)
M. V. Iorio, C. M. Croce, Causes and consequences of microRNA dysregulation. Cancer J. 8, 215–222 (2012)
S. Subramanian, W. O. Lui, C. H. Lee, I. Espinosa, T. O. Nielsen, M. C. Heinrich, C. L. Corless, A. Z. Fire, M. van de Rijn, MicroRNA expression signature of human sarcomas. Oncogene 27, 2015–2026 (2008)
L. S. Danielson, S. Menendez, C. S. Attolini, M. V. Guijarro, M. Bisogna, J. Wei, N. D. Socci, D. A. Levine, F. Michor, E. Hernando, A differentiation-based microRNA signature identifies leiomyosarcoma as a mesenchymal stem cell-related malignancy. Am. J. Pathol. 177, 908–917 (2010)
C. Zhou, W. Tan, H. Lv, F. Gao, J. Sun, Hypoxia-inducible microRNA-488 regulates apoptosis by targeting Bim in osteosarcoma. Cell. Oncol. 39, 463–471 (2016)
S. Morey, J. C. Ryan, F. M. Van Dolah, Microarray validation: factors influencing correlation between oligonucleotide microarrays and real-time PCR. Biol. Proc. Online 8, 175–193 (2006)
X. Zhou, F. Zhao, Z. N. Wang, Y. X. Song, H. Chang, Y. Chiang, H. M. Xu, Altered expression of miR-152 and miR-148a in ovarian cancer is related to cell proliferation. Oncol. Rep. 27, 447–454 (2012)
Z. Lichner, A. Fendler, C. Saleh, A. N. Nasser, D. Boles, S. Al-Haddad, P. Kupchak, M. Dharsee, P. S. Nuin, K. R. Evans, K. Jung, C. Stephan, N. E. Fleshner, G. M. Yousef, MicroRNA signature helps distinguish early from late biochemical failure in prostate cancer. Clin. Chem. 59, 1595–1603 (2013)
C. U. Köhler, O. Bryk, S. Meier, K. Lan, P. Rozynek, T. Brüning, H. U. Käfferlein, Analyses in human urothelial cells identify methylation of miR-152, miR-200b and miR-10a genes as candidate bladder cancer biomarkers. Biochem. Biophys. Res. Commun. 438, 48–53 (2013)
E. Hiroki, J. Akahira, F. Suzuki, S. Nagase, K. Ito, T. Suzuki, H. Sasano, N. Yaegashi, Changes in microRNA expression levels correlate with clinicopathological features and prognoses in endometrial serous adenocarcinomas. Cancer Sci. 101, 241–249 (2010)
T. Tsuruta, K. Kozaki, A. Uesugi, M. Furuta, A. Hirasawa, I. Imoto, N. Susumu, D. Aoki, J. Inazawa, miR-152 is a tumor suppressor microRNA that is silenced by DNA hypermethylation in endometrial cancer. Cancer Res. 71, 6450–6462 (2011)
M. Azizi, L. Teimoori-Toolabi, M. K. Arzanani, K. Azadmanesh, P. Fard-Esfahani, S. Zeinali, MicroRNA-148b and microRNA-152 reactivate tumor suppressor genes through suppression of DNA methyltransferase-1 gene in pancreatic cancer cell lines. Cancer Biol. Ther. 15, 419–427 (2014)
Y. Chen, Y. Song, Z. Wang, Z. Yue, H. Xu, C. Xing, Z. Liu, Altered expression of MiR-148a and MiR-152 in gastrointestinal cancers and its clinical significance. J. Gastrointest. Surg. 14, 1170–1179 (2010)
Y. X. Song, Z. Y. Yue, Z. N. Wang, Y. Y. Xu, Y. Luo, H. M. Xu, X. Zhang, L. Jiang, C. Z. Xing, Y. Zhang, MicroRNA-148b is frequently down-regulated in gastric cancer and acts as a tumor suppressor by inhibiting cell proliferation. Mol. Cancer 10, 1 (2011). doi:10.1186/1476-4598-10-1
D. Z. Liu, B. P. Ander, Y. Tian, B. Stamova, G. C. Jickling, R. R. Davis, F. R. Sharp, Integrated analysis of mRNA and microRNA expression in mature neurons, neural progenitor cells and neuroblastoma cells. Gene 495, 120–127 (2012)
N. G. Wang, D. C. Wang, B. Y. Tan, F. Wang, Z. N. Yuan, Down-regulation of microRNA152 is associated with the diagnosis and prognosis of patients with osteosarcoma. Int. J. Clin. Exp. Pathol. 8, 9314–9319 (2015)
Y. W. Dang, J. Zeng, R. Q. He, M. H. Rong, D. Z. Luo, G. Chen, Effects of miR-152 on cell growth inhibition, motility suppression and apoptosis induction in hepatocellular carcinoma cells. Asian Pac. J. Cancer Prev. 15, 4969–4976 (2014)
L. Li, Y. Y. Chen, S. Q. Li, C. Huang, Y. Z. Qin, Expression of miR-148/152 family as potential biomarkers in non-small-cell lung cancer. Med. Sci. Monit. 23, 1155–1161 (2015)
H. Huang, M. Hu, P. Li, C. Lu, M. Li, Mir-152 inhibits cell proliferation and colony formation of CD133+ liver cancer stem cells by targeting KIT. Tumor Biol. 36, 921–928 (2014)
T. Fukuda, E. Ichimura, T. Shinozaki, T. Sano, K. Kashiwabara, T. Oyama, T. Nakajima, T. Nakamura, Coexpression of HGF and c-met/HGF receptor in human bone and soft tissue tumors. Pathol. Int. 48, 757–762 (1998)
G. Lahat, P. Zhang, Q. S. Zhu, K. Torres, M. Ghadimi, K. D. Smith, W. L. Wang, A. J. Lazar, D. Lev, The expression of c-met pathway components in unclassified pleomorphic sarcoma/malignant fibrous histiocytoma (UPS/MFH): a tissue microarray study. Histopathology 59, 556–561 (2011)
L. Trusolino, A. Bertotti, P. M. Comoglio, MET signalling: principles and functions in development, organ regeneration and cancer. Nat. Rev. Mol. Cell Biol. 11, 834–848 (2010)
M. Miettinen, J. Lasota, KIT (CD117): a review on expression in normal and neoplastic tissues, and mutations and their clinicopathologic correlation. Appl. Immunohistochem. Mol. Morphol. 13, 205–220 (2005)
L. K. Ashman, R. Griffith, Therapeutic targeting of c-KIT in cancer. Expert Opin. Investig. Drugs 22, 103–115 (2013)
J. Noujaim, D. Gonzalez, K. Thway, R. L. Jones, I. Judson, P.(L576P) -KIT mutation in GIST: favourable prognosis and sensitive to imatinib? Cancer Biol. Ther. 17, 543–555 (2016)
E. D. Fleuren, M. H. Roeffen, W. P. Leenders, U. E. Flucke, M. Vlenterie, H. W. Schreuder, O. C. Boerman, W. T. van der Graaf, Y. M. Versleijen-Jonkers, Expression and clinical relevance of MET and ALK in Ewing sarcomas. Int. J. Cancer 133, 427–436 (2013)
M. Trovato, M. L. Torre, M. Ragonese, A. Simone, R. Scarfì, V. Barresi, G. Giuffrè, S. Benvenga, F. F. Angileri, G. Tuccari, F. Trimarchi, R. M. Ruggeri, S. Cannavò, HGF/c-met system targeting PI3K/AKT, STAT3/phosphorylated-STAT3 pathways in pituitary adenomas: an immunohistochemical characterization in view of targeted therapies. Endocrine 44, 735–743 (2013)
Y. Samuels, Z. Wang, A. Bardelli, N. Silliman, J. Ptak, S. Szabo, H. Yan, A. Gazdar, S. M. Powell, G. J. Riggins, J. K. Willson, S. Markowitz, K. W. Kinzler, B. Vogelstein, V. E. Velculescu, High frequency of mutations of the PIK3CA gene in human cancers. Science 304, 554 (2004)
Acknowledgments
The authors wish to thank Dr. Alba Balladelli for editing the paper and Ms. Cristina Ghinelli for her graphic work. This work was supported by 5‰ donation (Italy). Chiara Novello was supported by a FIRC Triennal fellowship “Mario e Valeria Rindi” (research project n°13748).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors have read and approved the contents of this manuscript and declare that the work is original and has not been submitted or published elsewhere. The authors also state that there is no potential conflict of interest.
Ethical standards
The research protocol was approved by ethic committees of the Rizzoli Institute where the tumor samples were collected. All patients provided informed consent.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Pazzaglia, L., Novello, C., Conti, A. et al. miR-152 down-regulation is associated with MET up-regulation in leiomyosarcoma and undifferentiated pleomorphic sarcoma. Cell Oncol. 40, 77–88 (2017). https://doi.org/10.1007/s13402-016-0306-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-016-0306-4