Abstract
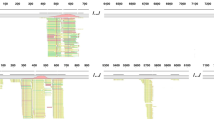
We screened for viral DNA in cerebrospinal fluid samples using next-generation sequencing (NGS) technology to diagnose CNS viral infections. We collected CSF samples from four cases with clinically suspected viral meningoencephalitis. DNA extracted from the samples was analyzed with NGS, and the results were further validated using PCR. Herpes simplex virus 1 (HSV-1) was detected in the CSF of two patients, HSV-2 and human herpes virus type 3 (HHV-3, VZV) in the CSF of two other patients separately. The number of unique reads of the identified viral genes ranged from 144 to 44205 (93.51 to 99.57 %). The coverage of identified viral genes ranged from 12 to 98 % with a depth value of 1.1 to 35, respectively. The results were further confirmed using PCR in three cases. The clinical presentation and outcomes of these four cases were consistent with the diagnostic results of NGS. NGS of CSF samples can be used as a diagnostic assay for CNS viral infection. Its further application for “pan-viral” or even “pan-microbial” screening of CSF might influence the diagnosis of CNS infectious diseases.



Similar content being viewed by others
Abbreviations
- CSF:
-
Cerebrospinal fluid
- NGS:
-
Next-generation sequencing
- EEG:
-
Electroencephalogram
- HSV:
-
Human herpes virus
- VZV:
-
Varicella zoster virus
References
Fournier PE, Dubourg G, Raoult D (2014) Clinical detection and characterization of bacterial pathogens in the genomics era. Genome Med 6(11):114. doi:10.1186/s13073-014-0114-2. eCollection 2014
Glaser CA, Honarmand S, Anderson LJ, Schnurr DP, Forghani B, Cossen CK, Schuster FL, Christie LJ, Tureen JH (2006) Beyond viruses: clinical profiles and etiologies associated with encephalitis. Clin Infect Dis 43:1565–1577
Jeon YJ, Zhou Y, Li Y, et al. (2014) The feasibility study of non-invasive fetal trisomy 18 and 21 detection with semiconductor sequencing platform. PLoSOne.2014;20;9(10):e110240. doi: 10.1371/journal.pone.0110240. eCollection 2014
Li H, Durbin R (2010) Fast and accurate short read alignment with Burrows Wheeler transform. Bioinformatics 26(5):589–595
Naccache SN, Federman S, Veeraraghavan N, Zaharia M, Lee D, Samayoa E, Bouquet J, Greninger AL, Luk KC, Enge B, Wadford DA, Messenger SL, Genrich GL, Pellegrino K, Grard G, Leroy E, Schneider BS, Fair JN, Martínez MA, Isa P, Crump JA, DeRisi JL, Sittler T, Hackett J Jr, Miller S, Chiu CY (2014) A cloud-compatible bioinformatics pipeline for ultrarapid pathogen identification from next-generation sequencing of clinical samples. Genome Res 24:1180–1192
Padmanabhan R, Mishra AK, Raoult D, Fournier PE (2013) Genomics and metagenomics in medical microbiology. J Microbiol Methods 95(3):415–424
Sherry NL, Porter JL, Seemann T, Watkins A, Stinear TP, Howden BP (2013) Outbreak investigation using high-throughput genome sequencing within a diagnostic microbiology laboratory. J Clin Microbiol 51(5):1396–1401
Wilson MR, Naccache SN, Samayoa E, Biagtan M, Bashir H, Yu G, Salamat SM, Somasekar S, Federman S, Miller S, Sokolic R, Garabedian E, Candotti F, Buckley RH, Reed KD, Meyer TL, Seroogy CM, Galloway R, Henderson SL, Gern JE, DeRisi JL, Chiu CY (2014) Actionable diagnosis of neuroleptospirosis by next-generation sequencing. N Engl J Med 370:2408–2417
Authors’ contributions
Hongzhi Guan, MD, is the principal investigator who drafted the original manuscript. Ao Shen, PhD, participated in laboratory analysis of CSF and drafted parts of the original manuscript. Xia Lv, MD, was involved in case and sample collection as well as analysis or interpretation of data. Xunzhe Yang, MD, was involved in case and sample collection, analysis, or interpretation of the data. Haitao Ren, BA, Yanhuan Zhao, BA, Yinxin Zhang, PhD, and Yanping Gong, PhD, participated in the laboratory analysis of CSF. Peixiang Ni, PhD, analyzed or interpreted the data. Honglong Wu, PhD, Yicheng Zhu, MD, and Liying Cui, MD, conceptualized the study and revised the manuscript.
Conflict of interest
We declare that we have no financial and personal relationships with other people or organizations that may inappropriately influence our work. There is no professional or other personal interest of any nature or kind in any product, service, and/or company that could be construed as influencing the position presented in or the review of the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Hongzhi Guan and Ao Shen contributed equally to this work.
Rights and permissions
About this article
Cite this article
Guan, H., Shen, A., Lv, X. et al. Detection of virus in CSF from the cases with meningoencephalitis by next-generation sequencing. J. Neurovirol. 22, 240–245 (2016). https://doi.org/10.1007/s13365-015-0390-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13365-015-0390-7