Abstract
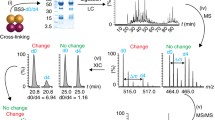
The topology of the GCAP-2 homodimer was investigated by chemical cross-linking and high resolution mass spectrometry. Complementary conducted size-exclusion chromatography and analytical ultracentrifugation studies indicated that GCAP-2 forms a homodimer both in the absence and in the presence of Ca2+. In-depth MS and MS/MS analysis of the cross-linked products was aided by 15 N-labeled GCAP-2. The use of isotope-labeled protein delivered reliable structural information on the GCAP-2 homodimer, enabling an unambiguous discrimination between cross-links within one monomer (intramolecular) or between two subunits (intermolecular). The limited number of cross-links obtained in the Ca2+-bound state allowed us to deduce a defined homodimeric GCAP-2 structure by a docking and molecular dynamics approach. In the Ca2+-free state, GCAP-2 is more flexible as indicated by the higher number of cross-links. We consider stable isotope-labeling to be indispensable for deriving reliable structural information from chemical cross-linking data of multi-subunit protein assemblies.

ᇵ






Similar content being viewed by others
References
Young, M.M., Tang, N., Hempel, J.C., Oshiro, C.M., Taylor, E.W., Kuntz, I.D., Gibson, B.W., Dollinger, G.: High throughput protein fold identification by using experimental constraints derived from intramolecular cross-links and mass spectrometry. Proc. Natl. Acad. Sci. U. S. A. 97, 5802–5806 (2000)
Back, J.W., de Jong, L., Muijsers, A.O., de Koster, C.G.: Chemical cross-linking and mass spectrometry for protein structural modeling. J. Mol. Biol. 331, 303–313 (2003)
Fabris, D., Yu, E.T.: Elucidating the higher-order structure of biopolymers by structural probing and mass spectrometry: MS3D. J. Mass Spectrom. 45, 841–860 (2010)
Sinz, A.: Chemical cross-linking and mass spectrometry to map three-dimensional protein structures and protein-protein interactions. Mass Spectrom. Rev. 25, 663–682 (2006)
Sinz, A.: Investigation of protein–protein interactions in living cells by chemical crosslinking and mass spectrometry. Anal. Bioanal. Chem. 397, 3433–3440 (2010)
Lee, Y.J.: Mass spectrometric analysis of cross-linking sites for the structure of proteins and protein complexes. Mol. Biosyst. 4, 816–823 (2008)
Zhang, H.Z., Tang, X.T., Munske, G.R., Tolic, N., Anderson, G.A., Bruce, J.E.: Identification of protein–protein interactions and topologies in living cells with chemical cross-linking and mass spectrometry. Mol. Cell. Proteom. 8, 409–420 (2009)
Rappsilber, J.: The beginning of a beautiful friendship: Cross-linking/mass spectrometry and modeling of proteins and multi-protein complexes. J. Struct. Biol. 173, 530–540 (2011)
Petrotchenko, E.V., Borchers, C.H.: Cross-linking combined with mass spectrometry for structural proteomics. Mass Spectrom. Rev. 29, 862–876 (2010)
Müller, D.R., Schindler, P., Towbin, H., Wirth, U., Voshol, H., Hoving, S., Steinmetz, M.O.: Isotope-tagged cross-linking reagents. A new tool in mass spectrometric protein interaction analysis. Anal. Chem. 73, 1927–1934 (2001)
Schmidt, A., Kalkhof, S., Ihling, C., Cooper, D.M.F., Sinz, A.: Mapping protein interfaces by chemical cross-linking and Fourier transform ion cyclotron resonance mass spectrometry: application to a calmodulin/adenylyl cyclase 8 peptide complex. Eur. J. Mass Spectrom. 11, 525–534 (2005)
Merkley, E.D., Baker, E.S., Crowell, K.L., Orton, D.J., Taverner, T., Ansong, C., Ibrahim, Y.M., Burnet, M.C., Cort, J.R., Anderson, G.A., Smith, R.D., Adkins, J.N.: Mixed-isotope labeling with LC-IMS-MS for characterization of protein–protein interactions by chemical cross-linking. J. Am. Soc. Mass Spectrom. 24, 444–449 (2013)
Taverner, T., Hall, N.E., O’Hair, R.A.J., Simpson, R.J.: Characterization of an antagonist interleukin-6 dimer by stable isotope labeling, cross-linking, and mass spectrometry. J. Biol. Chem. 277, 46487–46492 (2002)
Olshevskaya, E.V., Hughes, R.E., Hurley, J.B., Dizhoor, A.M.: Calcium binding, but not a calcium-myristoyl switch, controls the ability of guanylyl cyclase-activating protein GCAP-2 to regulate photoreceptor guanylyl cyclase. J. Biol. Chem. 272, 14327–14333 (1997)
Dizhoor, A.M., Olshevskaya, E.V., Henzel, W.J., Wong, S.C., Stults, J.T., Ankoudinova, I., Hurley, J.B.: Cloning, sequencing, and expression of a 24-kDa Ca2+-binding protein activating photo receptor guanylyl cyclase. J. Biol. Chem. 270, 25200–25206 (1995)
Laura, R.P., Dizhoor, A.M., Hurley, J.B.: The membrane guanylyl cyclase, retinal guanylyl cyclase-1, is activated through its intracellular domain. J. Biol. Chem. 271, 11646–11651 (1996)
Sharma, R.K.: Membrane guanylate cyclase is a beautiful signal transduction machine: overview. Mol. Cell. Biochem. 334, 3–36 (2010)
Wilson, E.M., Chinkers, M.: Identification of sequences mediating guanylyl cyclase dimerization. Biochemistry 34, 4696–4701 (1995)
Yang, R.B., Garbers, D.L.: Two eye guanylyl cyclases are expressed in the same photoreceptor cells and form homomers in preference to heteromers. J. Biol. Chem. 272, 13738–13742 (1997)
Olshevskaya, E.V., Ermilov, A.N., Dizhoor, A.M.: Dimerization of guanylyl cyclase-activating protein and a mechanism of photoreceptor guanylyl cyclase activation. J. Biol. Chem. 274, 25583–25587 (1999)
Ames, J.B., Dizhoor, A.M., Ikura, M., Palczewski, K., Stryer, L.: Three-dimensional structure of guanylyl cyclase activating protein-2, a calcium-sensitive modulator of photoreceptor guanylyl cyclases. J. Biol. Chem. 274, 19329–19337 (1999)
Schröder, T., Lilie, H., Lange, C.: The myristoylation of guanylate cyclase-activating protein-2 causes an increase in thermodynamic stability in the presence but not in the absence of Ca2+. Protein Sci. 20, 1155–1165 (2011)
Candiano, G., Bruschi, M., Musante, L., Santucci, L., Ghiggeri, G.M., Carnemolla, B., Orecchia, P., Zardi, L., Righetti, P.G.: Blue silver: a very sensitive colloidal coomassie G-250 staining for proteome analysis. Electrophoresis 25, 1327–1333 (2004)
Jensen, O.N., Shevchenko, A., Mann, M.: Protein analysis by mass spectrometry. In: Creighton, T.E. (ed.) In Protein Structure—A Practical Approach, 2nd ed., pp. 29–57. Oxford University Press, Oxford (1997)
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J.V., Mann, M.: In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2006)
Götze, M., Pettelkau, J., Schaks, S., Bosse, K., Ihling, C., Krauth, F., Fritzsche, R., Kühn, U., Sinz, A.: StavroX—a software for analyzing crosslinked products in protein interaction studies. J. Am. Soc. Mass. Spectrom. 23, 76–87 (2012)
Kalkhof, S., Sinz, A.: Chances and pitfalls of chemical cross-linking with amine-reactive N-hydroxysuccinimide esters. Anal. Bioanal. Chem. 392, 305–312 (2008)
Luther, M.A., Cai, G.Z., Lee, J.C.: Thermodynamics of dimer and tetramer formations in rabbit muscle phosphofructokinase. Biochemistry 25, 7931–7937 (1986)
Berjanskii, M., Zhou, J.J., Liang, Y.J., Lin, G.H., Wishart, D.S.: Resolution-by-proxy: a simple measure for assessing and comparing the overall quality of NMR protein structures. J. Biomol. NMR 53, 167–180 (2012)
Molecular Operating Environment (MOE), Chemical Computing Group Inc.: Montreal, Canada (2010)
Macindoe, G., Mavridis, L., Venkatraman, V., Devignes, M.D., Ritchie, D.W.: HexServer: an FFT-based protein docking server powered by graphics processors. Nucleic Acids Res. 38, W445–W449 (2010)
Comeau, S.R., Gatchell, D.W., Vajda, S., Camacho, C.J.: ClusPro: an automated docking and discrimination method for the prediction of protein complexes. Bioinformatics 20, 45–50 (2004)
Tovchigrechko, A., Vakser, I.A.: GRAMM-X public web server for protein–protein docking. Nucleic Acids Res. 34, W310–W314 (2006)
Pierce, B.G., Hourai, Y., Weng, Z.P.: Accelerating protein docking in ZDOCK using an advanced 3D convolution library. Plos One 6, (9): e24657 (2011). doi:10.1371/journal.pone.0024657
Torchala, M., Moal, I.H., Chaleil, R.A.G., Fernandez-Recio, J., Bates, P.A.: SwarmDock: a server for flexible protein-protein docking. Bioinformatics 29, 807–809 (2013)
Schneidman-Duhovny, D., Inbar, Y., Nussinov, R., Wolfson, H.J.: PatchDock and SymmDock: servers for rigid and symmetric docking. Nucleic Acids Res. 33, W363–W367 (2005)
De Vries, S.J., van Dijk, M., Bonvin, A.: The HADDOCK web server for data-driven biomolecular docking. Nat. Protoc. 5, 883–897 (2010)
Case, D.A., Darden, T.A., Cheatham III, T.E., Simmerling, C.L., Wang, J., Duke, R.E., Luo, R., Walker, R.C., Zhang, W., Merz, K.M., Roberts, B., Hayik, S., Roitberg, A., Seabra, G., Swails, J., Goetz, A.W., Kolossváry, I., Wong, K.F., Paesani, F., Vanicek, J., Wolf, R.M., Liu, J., Wu, X., Brozell, S.R., Steinbrecher, T., Gohlke, H., Cai, Q., Ye, X., Wang, J., Hsieh, M.J., Cui, G., Roe, D.R., Mathews, D.H., Seetin, M.G., Salomon-Ferrer, R., Sagui, C., Babin, V., Luchko, T., Gusarov, S., Kovalenko, A., Kollman, P.A.: AMBER 12. University of California, San Francisco (2012)
Mädler, S., Bich, C., Touboul, D., Zenobi, R.: Chemical cross-linking with NHS esters: a systematic study on amino acid reactivities. J. Mass Spectrom. 44, 694–706 (2009)
Pettelkau, J., Schröder, T., Ihling, C.H., Olausson, B.E.S., Kölbel, K., Lange, C., Sinz, A.: Structural insights into retinal guanylylcyclase-GCAP-2 interaction determined by cross-linking and mass spectrometry. Biochemistry 51, 4932–4949 (2012)
Chen, Z.A., Jawhari, A., Fischer, L., Buchen, C., Tahir, S., Kamenski, T., Rasmussen, M., Lariviere, L., Bukowski-Wills, J.C., Nilges, M., Cramer, P., Rappsilber, J.: Architecture of the RNA polymerase II-TFIIF complex revealed by cross-linking and mass spectrometry. EMBO J. 29, 717–726 (2010)
Green, N.S., Reisler, E., Houk, K.N.: Quantitative evaluation of the lengths of homobifunctional protein cross-linking reagents used as molecular rulers. Protein Sci. 10, 1293–1304 (2001)
Schilling, B., Row, R.H., Gibson, B.W., Guo, X., Young, M.M.: MS2Assign, automated assignment and nomenclature of tandem mass spectra of chemically crosslinked peptides. J. Am. Soc. Mass. Spectrom. 14, 834–850 (2003)
Santos, L.F.A., Iglesias, A.H., Gozzo, F.C.: Fragmentation features of intermolecular cross-linked peptides using N-hydroxy-succinimide esters by MALDI- and ESI-MS/MS for use in structural proteomics. J. Mass Spectrom. 46, 742–750 (2011)
Kalkhof, S., Haehn, S., Paulsson, M., Smyth, N., Meiler, J., Sinz, A.: Computational modeling of laminin N-terminal domains using sparse distance constraints from disulfide bonds and chemical cross-linking. Proteins 78, 3409–3427 (2010)
Acknowledgments
This work was funded by the Deutsche Forschungsgemeinschaft (DFG, project Si 867/13-1) and the region of Sachsen-Anhalt.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 3510 kb)
Rights and permissions
About this article
Cite this article
Pettelkau, J., Thondorf, I., Theisgen, S. et al. Structural Analysis of Guanylyl Cyclase-Activating Protein-2 (GCAP-2) Homodimer by Stable Isotope-Labeling, Chemical Cross-Linking, and Mass Spectrometry. J. Am. Soc. Mass Spectrom. 24, 1969–1979 (2013). https://doi.org/10.1007/s13361-013-0734-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-013-0734-6