Abstract
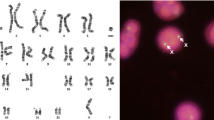
Here a new fluorescence in situ hybridization (FISH-) based probe set is presented and its possible applications are highlighted in 34 exemplary clinical cases. The so-called pericentric-ladder-FISH (PCL-FISH) probe set enables a characterization of chromosomal breakpoints especially in small supernumerary marker chromosomes (sSMC), but can also be applied successfully in large inborn or acquired derivative chromosomes. PCL-FISH was established as 24 different chromosome-specific probe sets and can be used in two- up multicolor-FISH approaches. PCL-FISH enables the determination of a chromosomal breakpoint with a resolution between 1 and ∼10 megabasepairs and is based on locus-specific bacterial artificial chromosome (BAC) probes. Results obtained on 29 sSMC cases and five larger derivative chromosomes are presented and discussed. To confirm the reliability of PCL-FISH, eight of the 29 sSMC cases were studied by array-comparative genomic hybridization (aCGH); the used sSMC-specific DNA was obtained by glass-needle based microdissection and DOP-PCR-amplification. Overall, PCL-FISH leads to a better resolution than most FISH-banding approaches and is a good tool to narrow down chromosomal breakpoints.


Similar content being viewed by others
References
Iourov IY, Vorsanova SG, Yurov YB (2008) Chromosomal mosaicism goes global. Mol Cytogenet 1:26
Lengauer C, Green ED, Cremer T (1992) Fluorescence in situ hybridization of YAC clones after Alu-PCR amplification. Genomics 13:826–828
Liehr T (2012a) Basics and literature on multicolor fluorescence in situ hybridization application. http://www.fish.uniklinikum-jena.de/mFISH.html. [accessed 09/02/2012]
Liehr T (2012b) Small supernumerary marker chromosomes. http://www.fish.uniklinikum-jena.de/sSMC.html. [accessed 09/02/2012]
Liehr T, Heller A, Starke H, Claussen U (2002a) FISH banding methods: applications in research and diagnostics. Exp Rev Mol Diagn 2:217–225
Liehr T, Heller A, Starke H, Rubtsov N, Trifonov V, Mrasek K, Weise A, Kuechler A, Claussen U (2002b) Microdissection based high resolution multicolor banding for all 24 human chromosomes. Int J Mol Med 9:335–339
Liehr T, Claussen U, Starke H (2004) Small supernumerary marker chromosomes (sSMC) in humans. Cytogenet Genome Res 107:55–67
Liehr T, Starke H, Heller A, Kosyakova N, Mrasek K, Gross M, Karst C, Steinhaeuser U, Hunstig F, Fickelscher I, Kuechler A, Trifonov V, Romanenko SA, Weise A (2006a) Multicolor fluorescence in situ hybridization (FISH) applied to FISH-banding. Cytogenet Genome Res 114:240–244
Liehr T, Mrasek K, Weise A, Dufke A, Rodríguez L, Martínez Guardia N, Sanchís A, Vermeesch JR, Ramel C, Polityko A, Haas OA, Anderson J, Claussen U, von Eggeling F, Starke H (2006b) Small supernumerary marker chromosomes–progress towards a genotype-phenotype correlation. Cytogenet Genome Res 112:23–34
Liehr T, Karamysheva T, Merkas M, Brecevic L, Hamid AB, Ewers E, Mrasek K, Kosyakova N, Weise A (2010) Somatic mosaicism in cases with small supernumerary marker chromosomes. Curr Genomics 11:432–439
Liehr T, Bartels I, Zoll B, Ewers E, Mrasek K, Kosyakova N, Merkas M, Hamid AB, von Eggeling F, Posorski N, Weise A (2011) Is there a yet unreported unbalanced chromosomal abnormality without phenotypic consequences in proximal 4p? Cytogenet Genome Res 132:121–123
Manolakos E, Vetro A, Kefalas K, Rapti S-M, Louizou E, Garas A, Kitsos G, Vasileiadis L, Tsoplou P, Eleftheriades M, Peitsidis P, Orru S, Liehr T, Petersen MB, Thomaidis L (2010) The use of array-CGH in a cohort of Greek children with developmental delay. Mol Cytogenet 3:22
Manvelyan M, Schreyer I, Höls-Herpertz I, Köhler S, Niemann R, Hehr U, Belitz B, Bartels I, Götz J, Huhle D, Kossakiewicz M, Tittelbach H, Neubauer S, Polityko A, Mazauric ML, Wegner R, Stumm M, Küpferling P, Süss F, Kunze H, Weise A, Liehr T, Mrasek K (2007) Forty-eight new cases with infertility due to balanced chromosomal rearrangements: detailed molecular cytogenetic analysis of the 90 involved breakpoints. Int J Mol Med 19:855–864
Nietzel A, Rocchi M, Starke H, Heller A, Fiedler W, Wlodarska I, Loncarevic IF, Beensen V, Claussen U, Liehr T (2001) A new multicolor-FISH approach for the characterization of marker chromosomes: centromere-specific multicolor-FISH (cenM-FISH). Hum Genet 108:199–204
Schröck E, du Manoir S, Veldman T, Schoell B, Wienberg J, Ferguson-Smith MA, Ning Y, Ledbetter DH, Bar-Am I, Soenksen D, Garini Y, Ried T (1996) Multicolor spectral karyotyping of human chromosomes. Science 273:494–497
Speicher MR, Gwyn Ballard S, Ward DC (1996) Karyotyping human chromosomes by combinatorial multi-fluor FISH. Nat Genet 12:368–375
Telenius H, Carter NP, Bebb CE, Nordenskjöld M, Ponder BA, Tunnacliffe A (1992) Degenerate oligonucleotide-primed PCR: general amplification of target DNA by a single degenerate primer. Genomics 13:718–725
Tsuchiya KD, Opheim KE, Hannibal MC, Hing AV, Glass IA, Raff ML, Norwood T, Torchia BA (2008) Unexpected structural complexity of supernumerary marker chromosomes characterized by microarray comparative genomic hybridization. Mol Cytogenet 1:7
van der Veken LT, Dieleman MMJ, Douben H, van de Brug JC, van de Graaf R, Hoogeboom AJM, Poddighe PJ, de Klein A (2010) Low grade mosaic for a complex supernumerary ring chromosome 18 in an adult patient with multiple congenital anomalies. Mol Cytogenet 3:13
Weimer J, Heidemann S, von Kaisenberg CS, Grote W, Arnold N, Bens S, Caliebe A (2011) Isolated trisomy 7q21.2-31.31 resulting from a complex familial rearrangement involving chromosomes 7, 9 and 10. Mol Cytogenet 4:28
Weise A, Starke H, Heller A, Tönnies H, Volleth M, Stumm M, Senger G, Nietzel A, Claussen U, Liehr T (2002) Chromosome 2 aberrations in clinical cases characterised by high resolution multicolour banding and region specific FISH probes. J Med Genet 39:434–439
Weise A, Mrasek K, Fickelscher I, Claussen U, Cheung SW, Cai WW, Liehr T, Kosyakova N (2008) Molecular definition of high-resolution multicolor banding probes: first within the human DNA sequence anchored FISH banding probe set. J Histochem Cytochem 56:487–493
Acknowledgments
The clinical cases were kindly provided by the following colleagues: Australia: J Anderson, Brisbane; Belgium: Dr. J. Vermeesch, Leuven; France: Dr. C. Yardin, Montpellier; Germany: Dr. I. Bartels, Göttingen; Dr. B. Belitz, Berlin; Dr. U. Beudt, Frankfurt; Dr. H.-M. Burow, Oberkirch; Dr. A. Dufke, Tübingen; Dr. G. Gillessen-Kasebach, Lübeck; Dr. D. Huhle, Leipzig; Dr. A. Kuechler, Essen; Dr. T. Martin, Homburg; Dr. A. Meiner, Halle; Dr. D. Mitter, Leipzig; Dr. S. Morlot, Hannover; Dr. A. Ovens-Raeder, München, Dr. G. Schwan, Dortmund; Dr. S. Singer, Tübingen; Dr. S. Spranger, Bremen; Portugal: Dr. J. Melo, Coimbra; Serbia: Dr. G. Josik, Vinca; Turkey: Dr. B. Seher, Ankara; UK: Dr. K. Ren Ong, Birmingham.
Supported in parts by Deutsche Forschungsgemeinschaft (DFG LI 820/22-1), Else Kröner-Fresenius-Stiftung (2011_A42), the Deutscher Akademischer Austauschdienst (DAAD), the Monika-Kutzner-Stiftung and the Stefan-Morsch-Stiftung.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hamid, A.B., Kreskowski, K., Weise, A. et al. How to narrow down chromosomal breakpoints in small and large derivative chromosomes – a new probe set. J Appl Genetics 53, 259–269 (2012). https://doi.org/10.1007/s13353-012-0098-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-012-0098-9