Abstract
The expression of nine functional candidates for QT abdominal fat weight and relative abdominal fat content was investigated by real-time polymerase chain reaction (PCR) in the liver, adipose tissue, colon, muscle, pituitary gland and brain of broilers. The high mobility group AT-hook 1 (HMG1A) gene was up-regulated in liver with a ratio of means of 2.90 (P ≤ 0.01) in the «fatty» group (relative abdominal fat content 3.5 ± 0.18%, abdominal fat weight 35.4 ± 6.09 g) relative to the «lean» group (relative abdominal fat content 1.9 ± 0.56%, abdominal fat weight 19.2 ± 5.06 g). Expression of this gene was highly correlated with the relative abdominal fat content (0.70, P ≤ 0.01) and abdominal fat weight (0.70, P ≤ 0.01). The peroxisome proliferator-activated receptor gamma (PPARG) gene was also up-regulated in the liver with a ratio of means of 3.34 (P ≤ 0.01) in the «fatty» group relative to the «lean» group. Correlation of its expression was significant with both the relative abdominal fat content (0.55, P ≤ 0.05) and the abdominal fat weight (0.57, P ≤ 0.01). These data suggest that the HMG1A and PPARG genes were candidate genes for abdominal fat deposition in chickens. Searching of rSNPs in regulatory regions of the HMG1A and PPARG genes could provide a tool for gene-assisted selection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Biological mechanism of the deposition of abdominal fat seems to be important for both the understanding of obesity in humans and increasing productivity in farm animals. Reduction of abdominal fat deposition could allow increasing feed efficiency and carcass value significantly. Based on previously performed research in the Department of Molecular Cytogenetics at the Institute of Genetics and Animal Breeding and the Laboratory of Molecular Genome Organization, Institute of Farm Animal Genetics and Breeding, we selected nine most probable functional candidate genes (FABP1, FABP2, FABP3, HMGA1, MC4R, POMC, PPARG, PPARGC1A and PTPN1) for expression profiling in the adipose tissue, brain, colon, liver, muscle and pituitary gland of broilers.
Fatty acid-binding proteins (FABPs) belong to the family of small cytoplasmic proteins. FABP family members are thought to play roles in fatty acid uptake, transport and metabolism. The FABP genes expression level was significantly increased in obese rats compared with controls (López et al. 2003). The HMGA1 gene encodes high mobility group AT-hook 1 protein. This non-histone protein was involved in a number of cellular processes, such as the integration of retroviruses into chromosomes, metastatic progression of malignant cells and the regulation of inducible gene transcription. The MC4R gene encodes melanocortin 4 receptor. HMGA1 and MC4R were significantly associated with a fat deposition measurement in pigs (Kim et al. 2004). The mutations of the MC4R gene cause several obesity forms in humans (Tan et al. 2009; Calton et al. 2009; Wangensteen et al. 2009) and could be associated with feed intake, fatness and growth in pigs (Meidtner et al. 2006; Bruun et al. 2006). Pro-opiomelanocortin (POMC) plays a key role in the regulation of body weight. The POMC gene mutations were reported to be associated with human obesity (Dubern et al. 2008). The PPARG gene encodes the peroxisome proliferator-activated receptor gamma. It was implicated in the pathology of obesity, diabetes, atherosclerosis and cancer (Qi et al. 2000). PPARGC1A encodes peroxisome proliferator-activated receptor gamma coactivator 1 alpha. This gene plays a significant role in the development of obesity in humans (Okauchi et al. 2008; Lu et al. 2007; Franks et al. 2007) and affects back fat in pigs (Stachowiak et al. 2007). The PTPN1 gene encodes tyrosine phosphatase, non-receptor type 1, which was also reported to dephosphorylate epidermal growth factor receptor kinase. SNPs in this gene are associated with human obesity (Kipfer-Coudreau et al. 2004; Ukkola et al. 2005).
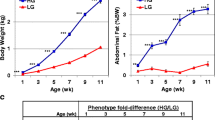
Thirty-five-day-old broilers from cross Iza 15, kindly provided by Dr. O.I. Stanishevskaya (Department of Poultry Science, Institute of Animal Genetics and Breeding, Russian Academy of Agricultural Science), were investigated. Ten birds each from the «fatty» group (relative abdominal fat content 3.5 ± 0.18%, abdominal fat weight 35.4 ± 6.09 g) and the «lean» group (relative abdominal fat content 1.9 ± 0.56%, abdominal fat weight 19.2 ± 5.06 g) were used for expression quantification.
The mRNA samples were isolated from frozen tissues using the Aurum total RNA Fatty and Fibrous Kit (Bio-Rad, USA).
Gene-specific primers were designed using database information (http://www.ncbi.nlm.nih.gov and http://www.ensembl.org) and the computer software PRIMER _3 (http://frodo.wi.mit.edu/primer3) (Table 1). GAPDH was used as a reporter gene in our experiment. The polymerase chain reaction (PCR) reaction mix was prepared following standard protocols using the iScript One-Step RT-PCR Kit (Bio-Rad, USA). The 2ΔΔCt method (Livak and Schmittgen 2001) for the calculation of the relative ratio was used. Differences between the mean of candidate-expressed sequence tags quantities in tissues of «fatty» and «lean» groups were tested for by a two-tailed t-test. Pearson’s technique was used for correlation calculation.
No significant difference was detected in expression of the genes FABP1 and PPARGC1A in liver, FABP2 and PTPN1 in colon, MC4R and PTPN1 in brain, FABP3 in skeletal muscle, PPARG in adipose tissue and POMC in pituitary gland in broilers with different relative abdominal fat content and abdominal fat weight. Correlation of their expression with the investigated traits was also not significant (Table 2). No significant difference in expression of the FABP family of genes was also shown by Wang et al. 2009.
The HMGA1 gene was up-regulated in liver with a ratio of means of 2.90 (P ≤ 0.01) in the «fatty» group relative to the «lean» group (Table 2). Expression of this gene was highly correlated with the relative abdominal fat content (0.70, P ≤ 0.01) and the abdominal fat weight (0.70, P ≤ 0.01) (Table 2). Significant influence of HMGA1 and MC4R in growth and fat deposition in duroc pigs has been shown (Kim et al. 2006); however, Stachowiak et al. 2006 did not find an MC4R effect on the production traits of polish large white and polish landrace pig breeds.
The PPARG gene was also up-regulated in liver with a ratio of means of 3.34 (P ≤ 0.01) in the «fatty» group relative to the «lean» group. Correlation of its expression was significant with both the relative abdominal fat content (0.55, P ≤ 0.05) and the abdominal fat weight (0.57, P ≤ 0.01) (Table 2), but this gene did not show a significant difference in expression in adipose tissue. The PPARG gene encodes the peroxisome proliferator-activated receptor gamma, which participates in adipocytes differentiation. Hyperexpression of this gene correlates with obesity in humans (Hindle et al. 2009). Three SNPs in the ESR1 and PPARG genes were shown to be genetically linked with obesity in Han Chinese (Chen et al. 2009). Association of the PPARG expression with fat deposition in broilers was also reported by Sato et al. (2004). This gene was found among the differentially expressed proteins in adipose tissue of divergently selected broilers (Wang et al. 2009).
These data suggest that the HMG1A and PPARG genes were candidate genes for abdominal fat deposition in chickens. It is interesting that both candidate genes (HMGA1 and PPARG) are responsible for adipocytes proliferation and, until 4–5 weeks of age, increase in the number of fat cells (hyperplasia) dominates in abdominal fat deposition in chickens (Leenstra 1986; Mourot and Hermier 2001).
References
Bruun CS, Jørgensen CB, Nielsen VH et al (2006) Evaluation of the porcine melanocortin 4 receptor (MC4R) gene as a positional candidate for a fatness QTL in a cross between Landrace and Hampshire. Anim Genet 37:359–362
Calton MA, Ersoy BA, Zhang S et al (2009) Association of functionally significant Melanocortin-4 but not Melanocortin-3 receptor mutations with severe adult obesity in a large North American case–control study. Hum Mol Genet 18:1140–1147
Chen HH, Lee WJ, Fann CS et al (2009) Severe obesity is associated with novel single nucleotide polymorphisms of the ESR1 and PPARgamma locus in Han Chinese. Am J Clin Nutr 90:255–262
Dubern B, Lubrano-Berthelier C, Mencarelli M et al (2008) Mutational analysis of the pro-opiomelanocortin gene in French obese children led to the identification of a novel deleterious heterozygous mutation located in the alpha-melanocyte stimulating hormone domain. Pediatr Res 63:211–216
Franks PW, Ekelund U, Brage S et al (2007) PPARGC1A coding variation may initiate impaired NEFA clearance during glucose challenge. Diabetologia 50:569–573
Hindle AK, Koury J, McCaffrey T et al (2009) Dysregulation of gene expression within the peroxisome proliferator activated receptor pathway in morbidly obese patients. Surg Endosc 23:1292–1297
Kim KS, Thomsen H, Bastiaansen J et al (2004) Investigation of obesity candidate genes on porcine fat deposition quantitative trait loci regions. Obes Res 12:1981–1994
Kim KS, Lee JJ, Shin HY et al (2006) Association of melanocortin 4 receptor (MC4R) and high mobility group AT-hook 1 (HMGA1) polymorphisms with pig growth and fat deposition traits. Anim Genet 37:419–421
Kipfer-Coudreau S, Eberlé D, Sahbatou M et al (2004) Single nucleotide polymorphisms of protein tyrosine phosphatase 1B gene are associated with obesity in morbidly obese French subjects. Diabetologia 47:1278–1284
Leenstra FR (1986) Effect of age, sex, genotype and environment on fat deposition in broiler chickens—a review. Worlds Poult Sci J 42:12–25
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2ΔΔCt method. Methods 25:402–408
López IP, Marti A, Milagro FI et al (2003) DNA microarray analysis of genes differentially expressed in diet-induced (cafeteria) obese rats. Obes Res 11:188–194
Lu WS, Yan XD, Liu HY et al (2007) The cSNPs analysis in whole extron-wide of PGC-1alpha gene in Chinese population and the domain MEF2C bioinformatics study. Zhonghua Yi Xue Yi Chuan Xue Za Zhi 24:409–416
Meidtner K, Wermter AK, Hinney A et al (2006) Association of the melanocortin 4 receptor with feed intake and daily gain in F2 Mangalitsa × Piétrain pigs. Anim Genet 37:245–247
Mourot J, Hermier D (2001) Lipids in monogastric animal meat. Reprod Nutr Dev 41:109–118
Okauchi Y, Iwahashi H, Okita K et al (2008) PGC-1alpha Gly482Ser polymorphism is associated with the plasma adiponectin level in type 2 diabetic men. Endocr J 55:991–997
Qi C, Zhu Y, Reddy JK (2000) Peroxisome proliferator-activated receptors, coactivators, and downstream targets. Cell Biochem Biophys 32:187–204
Sato K, Fukao K, Seki Y et al (2004) Expression of the chicken peroxisome proliferator-activated receptor-gamma gene is influenced by aging, nutrition, and agonist administration. Poult Sci 83:1342–1347
Stachowiak M, Szydlowski M, Obarzanek-Fojt M et al (2006) An effect of a missense mutation in the porcine melanocortin-4 receptor (MC4R) gene on production traits in polish pig breeds is doubtful. Anim Genet 37(1):55–57
Stachowiak M, Szydlowski M, Cieslak J et al (2007) SNPs in the porcine PPARGC1a gene: interbreed differences and their phenotypic effects. Cell Mol Biol Lett 12:231–239
Tan K, Pogozheva ID, Yeo GS et al (2009) Functional characterization and structural modeling of obesity associated mutations in the melanocortin 4 receptor. Endocrinology 150:114–125
Ukkola O, Rankinen T, Lakka T et al (2005) Protein tyrosine phosphatase 1B variant associated with fat distribution and insulin metabolism. Obes Res 13:829–834
Wang D, Wang N, Li N et al (2009) Identification of differentially expressed proteins in adipose tissue of divergently selected broilers. Poult Sci 88:2285–2292
Wangensteen T, Kolsgaard ML, Mattingsdal M et al (2009) Mutations in the melanocortin 4 receptor (MC4R) gene in obese patients in Norway. Exp Clin Endocrinol Diabetes 117:266–273
Acknowledgements
This work was supported by MSHE grant 311 32 34 36.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Larkina, T.A., Sazanova, A.L., Fomichev, K.A. et al. HMG1A and PPARG are differently expressed in the liver of fat and lean broilers. J Appl Genetics 52, 225–228 (2011). https://doi.org/10.1007/s13353-010-0023-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-010-0023-z