Abstract
Background and Objective
The clearance, by renal elimination or hepatic metabolism, is one of the most important pharmacokinetic parameters of a drug. It allows the half-life, bioavailability, and drug–drug interactions to be predicted, and it can also affect the dose regimen of a drug. Predicting the clearance pathways of new chemical candidates during drug development is vital in order to minimize the risks of possible side effects and drug interactions. Many in vivo methods have been established to predict drug clearance in humans, and these mainly rely on data from in vivo studies in preclinical species—mainly rats, dogs, and monkeys. They are also time consuming and expensive. The aim of this study was to find the relationship between structural parameters of drugs and their clearance pathways.
Methods
The clearance pathway of each drug was obtained from the literature. Various structural descriptors [Abraham solvation parameters, topological polar surface area, numbers of hydrogen-bond donors and acceptors, number of rotatable bonds, molecular weight, logarithm of the partition coefficient (logP), and logarithm of the distribution coefficient at pH 7.4 (logD7.4)] were applied to develop a mechanistic model for predicting clearance pathways.
Results
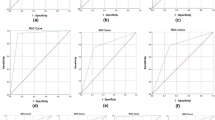
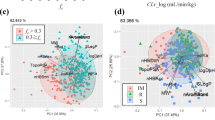
The results of this study indicate that compounds with logD7.4 > 1 or with zero or one hydrogen-bond donor undergo hepatic metabolism, whereas the clearance pathway for chemicals with logD7.4 < − 2 is renal elimination. Furthermore, models established using logistic regression based on five structural parameters for compounds with – 2 < logD7.4 < 1 could be used in a clearance pathway prediction tool. The overall prediction accuracies of the first and second models were 84.8% and 84.4%, respectively.
Conclusion
The developed model can be used to find the clearance pathways of new drug candidates with acceptable accuracy. The main descriptors that are used to evaluate this parameter are the hydrophobicity and the number of hydrogen-bonding functional groups of the compound.



Similar content being viewed by others
References
Kunze A, Huwyler J, Poller B, Gutmann H, Camenisch G. In vitro-in vivo extrapolation method to predict human renal clearance of drugs. J Pharm Sci. 2014;103(3):994–1001. https://doi.org/10.1002/jps.23851.
Rayasilp K, Wonganan P, Chariyavilaskul P, Prompila N, Sukkummee V, Wittayalertpanya S. Effect of pomelo juice on the pharmacokinetics of simvastatin, CYP3A2 activity and Mdr1a, Mdr1b and Slc21a5 expressions in rats. Pharm Sci. 2019;25(4):303–10. https://doi.org/10.15171/PS.2019.45.
Wu CY, Benet LZ. Predicting drug disposition via application of BCS: transport/absorption/ elimination interplay and development of a biopharmaceutics drug disposition classification system. Pharm Res. 2005;22(1):11–23. https://doi.org/10.1007/s11095-004-9004-4.
Ito S, Ando H, Ose A, Kitamura Y, Ando T, Kusuhara H, et al. Relationship between the urinary excretion mechanisms of drugs and their physicochemical properties. J Pharm Sci. 2013;102(9):3294–301. https://doi.org/10.1002/jps.23599.
Berellini G, Waters NJ, Lombardo F. In silico prediction of total human plasma clearance. J Chem Inf Model. 2012;52(8):2069–78. https://doi.org/10.1021/ci300155y.
Wakayama N, Toshimoto K, Maeda K, Hotta S, Ishida T, Akiyama Y, et al. In silico prediction of major clearance pathways of drugs among 9 routes with two-step support vector machines. Pharm Res. 2018. https://doi.org/10.1007/s11095-018-2479-1.
Lombardo F, Obach RS, Varma MV, Stringer R, Berellini G. Clearance mechanism assignment and total clearance prediction in human based upon in silico models. J Med Chem. 2014;57(10):4397–405. https://doi.org/10.1021/jm500436v.
ACD/Labs. ACD/iLab. 2021. Available from https://ilab.acdlabs.com. Accessed 30 July 2021.
Ulrich N, Endo S, Brown TN, Watanabe N, Bronner G, Abraham MH, Goss K-U. UFZ-LSER database v 3.2.1 [Internet]. Leipzig, Germany: Helmholtz Centre for Environmental Research-UFZ; 2017. Available from http://www.ufz.de/lserd. Accessed 30 July 2021.
Hajian-Tilaki K. Receiver operating characteristic (ROC) curve analysis for medical diagnostic test evaluation. Casp J Int Med. 2013;4(2):627–35.
Esmaeili A, Salehi M, Makhdomi N, Ardakani YH, Rajabi M, Namazi S. Evaluation of the association between trough and area under the curve to minimum inhibitory concentration ratio (AUC24/MIC) of vancomycin in infected patients with methicillin resistant Staphylococcus aureus (MRSA). Pharm Sci. 2021;27(2):201–8. https://doi.org/10.34172/PS.2020.70.
Chen J, Yang H, Zhu L, Wu Z, Li W, Tang Y, et al. In silico prediction of human renal clearance of compounds using quantitative structure–pharmacokinetic relationship models. Chem Res Toxicol. 2020;33(2):640–50. https://doi.org/10.1021/acs.chemrestox.9b00447.
Ghotbi G, Hamzeh-Mivehroud M, Taghvimi A, Davaran S, Dastmalchi S. Investigation of experimental and in silico physicochemical properties of thiazole-pyridinium anti-acetylcholinesterase derivatives with potential anti-Alzheimer’s activity. Pharm Sci. 2021;27(3):366–77. https://doi.org/10.34172/ps.2020.81.
Pieńko T, Grudzień M, Taciak PP, Mazurek AP. Cytisine basicity, solvation, log P, and log D theoretical determination as tool for bioavailability prediction. J Mol Graph Model. 2016;63:15–21. https://doi.org/10.1016/j.jmgm.2015.11.003.
Kah M, Brown CD. Log D: Lipophilicity for ionisable compounds. Chemosphere. 2008;72(10):1401–8. https://doi.org/10.1016/j.chemosphere.2008.04.074.
Beheshti S, Shayanfar A. Prediction of the oral bioavailability correlation between humans and preclinical animals. Eur J Drug Metab Pharmacokinet. 2020;45(6):771–83. https://doi.org/10.1007/s13318-020-00636-2.
Golfar Y, Shayanfar A. Prediction of biopharmaceutical drug disposition classification system (BDDCS) by structural parameters. J Pharm Pharm Sci. 2019;22(1):247–69. https://doi.org/10.18433/jpps30271.
Shayanfar S, Shayanfar A. Predicting protein binding of drugs using Abraham parameters: Effect of ionization. J Mazandaran Univ Med Sci. 2019;29(174):96–105.
Kamble S, Loadman P, Abraham MH, Liu X. Structural properties governing drug–plasma protein binding determined by high-performance liquid chromatography method. J Pharm Biomed Anal. 2018;149:16–21. https://doi.org/10.1016/j.jpba.2017.10.022.
Feng B, LaPerle JL, Chang G, Varma MV. Renal clearance in drug discovery and development: molecular descriptors, drug transporters and disease state. Expert Opin Drug Metab Toxicol. 2010;6(8):939–52. https://doi.org/10.1517/17425255.2010.482930.
Dearden JC, Cronin MTD, Kaiser KLE. How not to develop a quantitative structure–activity or structure–property relationship (QSAR/QSPR). SAR QSAR Environ Res. 2009;20(3–4):241–66. https://doi.org/10.1080/10629360902949567.
Guha R. On the interpretation and interpretability of quantitative structure–activity relationship models. J Comp-Aided Mol Des. 2008;22(12):857–71. https://doi.org/10.1007/s10822-008-9240-5.
Hosey CM, Chan R, Benet LZ. BDDCS predictions, self-correcting aspects of BDDCS assignments, BDDCS assignment corrections, and classification for more than 175 additional drugs. AAPS J. 2016;18(1):251–60. https://doi.org/10.1208/s12248-015-9845-2.
Palmer DS, Mitchell JBO. Is experimental data quality the limiting factor in predicting the aqueous solubility of druglike molecules? Mol Pharm. 2014;11(8):2962–72. https://doi.org/10.1021/mp500103r.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
The research reported in this publication was supported by the Elite Researcher Grant Committee under award number 4000425 from the National Institute for Medical Research Development (NIMAD), Tehran, Iran.
Conflict of interest
Navid Kaboudi and Ali Shayanfar have no conflict of interest.
Ethical approval
The study was approved by the research ethics committees of the National Institute for Medical Research Development, Tehran, Iran (IR.NIMAD.REC.1400.037).
Consent to participate
Not applicable.
Consent to publish
Not applicable.
Availability of data and material
All data are available as a supplementary file (Table S1) on the journal’s website along with the published article.
Code availability
Not applicable.
Author contributions
Navid Kaboudi: data collection, data analysis and interpretation, drafting the article; Ali Shayanfar: design of the work, supervision of the project, critical revision of the article. All authors read and approved the final manuscript.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kaboudi, N., Shayanfar, A. Predicting the Drug Clearance Pathway with Structural Descriptors. Eur J Drug Metab Pharmacokinet 47, 363–369 (2022). https://doi.org/10.1007/s13318-021-00748-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13318-021-00748-3