Abstract
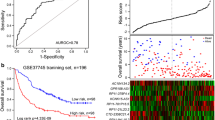
Lung adenocarcinoma (LAD) and lung squamous cell cancer (LSCC) are two major histological types of non-small cell lung cancer. LSCC differs greatly from LAD in many aspects. Accumulating evidence has shown that long noncoding RNA (lncRNA) plays an important role in the process of carcinogenesis and tumor progression. Expression of lncRNA is highly tissue-specific and could be biomarkers for cancer diagnosis and prognosis. Here, we identified differentially expressed lncRNA between LSCC and LAD by data mining of Affymetrix HG-U133 Plus 2.0 microarray. A set of 1646 differentially expressed lncRNA transcripts were identified. Among these lncRNAs, a novel lncRNA, LINC01133, showed the largest fold change among large intergenic noncoding RNAs. Quantitative real-time polymerase chain reaction (PCR) assay confirmed that LINC01133 was upregulated in LSCC (increasing fold 6.4, P < 0.01) but not in the LAD samples. LSCC patients with higher expression level of LINC01133 had shorter survival time (hazard ratio = 2.383; 95 % confidence interval 1.023–5.547, P = 0.044). Wound-healing and transwell assays demonstrated that silence of LINC01133 by small interfering RNA (siRNA) inhibited invasion ability of LSCC cell line. Thus, a set of lncRNA was differentially expressed between LAD and LSCC and could serve as potential biomarkers.



Similar content being viewed by others
References
Siegel R, Naishadham D, Jemal A. Cancer statistics, 2012. CA Cancer J Clin. 2012;62(1):10–29.
Parkin DM, Pisani P, Ferlay J. Estimates of the worldwide incidence of 25 major cancers in 1990. Int J Cancer. 1999;80(6):827–41.
Jemal A, Siegel R, Xu J, Ward E. Cancer statistics, 2010. CA Cancer J Clin. 2010;60(5):277–300.
Yildiz O, Buyuktas D, Ekiz E, Selcukbiricik F, Papila I, Papila C. Facial nerve palsy: an unusual presenting feature of small cell lung cancer. Case Rep Oncol. 2011;4(1):35–8.
Sun S, Schiller JH, Gazdar AF. Lung cancer in never smokers—a different disease. Nat Rev Cancer. 2007;7(10):778–90.
Ponting CP, Oliver PL, Reik W. Evolution and functions of long noncoding RNAs. Cell. 2009;136(4):629–41.
Esteller M. Non-coding RNAs in human disease. Nat Rev Genet. 2011;12(12):861–74.
Consortium EP, Birney E, Stamatoyannopoulos JA, Dutta A, Guigo R, Gingeras TR, et al. Identification and analysis of functional elements in 1 % of the human genome by the ENCODE pilot project. Nature. 2007;447(7146):799–816.
Iyer MK, Niknafs YS, Malik R, Singhal U, Sahu A, Hosono Y, et al. The landscape of long noncoding RNAs in the human transcriptome. Nat Genet. 2015;47(3):199–208.
Prensner JR, Zhao S, Erho N, Schipper M, Iyer MK, Dhanasekaran SM, et al. RNA biomarkers associated with metastatic progression in prostate cancer: a multi-institutional high-throughput analysis of SChLAP1. Lancet Oncol. 2014;15(13):1469–80.
Garzon R, Volinia S, Papaioannou D, Nicolet D, Kohlschmidt J, Yan PS, et al. Expression and prognostic impact of lncRNAs in acute myeloid leukemia. Proc Natl Acad Sci U S A. 2014;111(52):18679–84.
Tsai MC, Spitale RC, Chang HY. Long intergenic noncoding RNAs: new links in cancer progression. Cancer Res. 2011;71(1):3–7.
Qiu MT, Hu JW, Yin R, Xu L. Long noncoding RNA: an emerging paradigm of cancer research. Tumour Biol. 2013;34(2):613–20.
Sanchez Y, Segura V, Marin-Bejar O, Athie A, Marchese FP, Gonzalez J, et al. Genome-wide analysis of the human p53 transcriptional network unveils a lncRNA tumour suppressor signature. Nat Commun. 2014;5:5812.
Ding X, Zhu L, Ji T, Zhang X, Wang F, Gan S, et al. Long intergenic non-coding RNAs (lincRNAs) identified by RNA-seq in breast cancer. PLoS One. 2014;9(8):e103270.
Ren S, Peng Z, Mao JH, Yu Y, Yin C, Gao X, et al. RNA-seq analysis of prostate cancer in the Chinese population identifies recurrent gene fusions, cancer-associated long noncoding RNAs and aberrant alternative splicings. Cell Res. 2012;22(5):806–21.
Wang P, Lu S, Mao H, Bai Y, Ma T, Cheng Z, et al. Identification of biomarkers for the detection of early stage lung adenocarcinoma by microarray profiling of long noncoding RNAs. Lung Cancer. 2015.
Xue Y, Ma G, Zhang Z, Hua Q, Chu H, Tong N, et al. A novel antisense long noncoding RNA regulates the expression of MDC1 in bladder cancer. Oncotarget. 2015;6(1):484–93.
Yang F, Huo XS, Yuan SX, Zhang L, Zhou WP, Wang F, et al. Repression of the long noncoding RNA-LET by histone deacetylase 3 contributes to hypoxia-mediated metastasis. Mol Cell. 2013;49(6):1083–96.
Kuner R, Muley T, Meister M, Ruschhaupt M, Buness A, Xu EC, et al. Global gene expression analysis reveals specific patterns of cell junctions in non-small cell lung cancer subtypes. Lung Cancer. 2009;63(1):32–8.
Bonnefont J, Laforge T, Plastre O, Beck B, Sorce S, Dehay C, et al. Primate-specific RFPL1 gene controls cell-cycle progression through cyclin B1/Cdc2 degradation. Cell Death Differ. 2011;18(2):293–303.
Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics. 2003;4(2):249–64.
Smyth GK. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol. 2004;3:Article3.
Yang J, Lin J, Liu T, Chen T, Pan S, Huang W, et al. Analysis of lncRNA expression profiles in non-small cell lung cancers (NSCLC) and their clinical subtypes. Lung Cancer. 2014;85(2):110–5.
Zhang X, Sun S, Pu JK, Tsang AC, Lee D, Man VO, et al. Long non-coding RNA expression profiles predict clinical phenotypes in glioma. Neurobiol Dis. 2012;48(1):1–8.
Derrien T, Johnson R, Bussotti G, Tanzer A, Djebali S, Tilgner H, et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 2012;22(9):1775–89.
Prensner JR, Iyer MK, Balbin OA, Dhanasekaran SM, Cao Q, Brenner JC, et al. Transcriptome sequencing across a prostate cancer cohort identifies PCAT-1, an unannotated lincRNA implicated in disease progression. Nat Biotechnol. 2011;29(8):742–9.
Xu G, Chen J, Pan Q, Huang K, Pan J, Zhang W, et al. Long noncoding RNA expression profiles of lung adenocarcinoma ascertained by microarray analysis. PLoS One. 2014;9(8):e104044.
Giordano TJ, Kuick R, Else T, Gauger PG, Vinco M, Bauersfeld J, et al. Molecular classification and prognostication of adrenocortical tumors by transcriptome profiling. Clin Cancer Res. 2009;15(2):668–76.
Gibb EA, Vucic EA, Enfield KS, Stewart GL, Lonergan KM, Kennett JY, et al. Human cancer long non-coding RNA transcriptomes. PLoS One. 2011;6(10):e25915.
Kretz M, Siprashvili Z, Chu C, Webster DE, Zehnder A, Qu K, et al. Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature. 2013;493(7431):231–5.
Wu W, Bhagat TD, Yang X, Song JH, Cheng Y, Agarwal R, et al. Hypomethylation of noncoding DNA regions and overexpression of the long noncoding RNA, AFAP1-AS1, in Barrett’s esophagus and esophageal adenocarcinoma. Gastroenterology. 2013;144(5):956–66 e4.
Acknowledgments
This work has no funding source.
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding author
Additional information
Jing Zhang and Ning Zhu contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Table 1
Differentially expressed lncRNAs between LSCC and LAD (XLSX 187 kb)
Rights and permissions
About this article
Cite this article
Zhang, J., Zhu, N. & Chen, X. A novel long noncoding RNA LINC01133 is upregulated in lung squamous cell cancer and predicts survival. Tumor Biol. 36, 7465–7471 (2015). https://doi.org/10.1007/s13277-015-3460-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-3460-9