Abstract
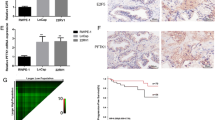
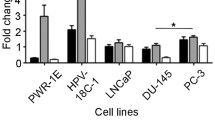
The long-lasting inadequacy of existing treatments for prostate cancer has led to increasing efforts for developing novel therapies for this disease. MicroRNAs (miRNAs) are believed to have considerable therapeutic potential due to their role in regulating gene expression and cellular pathways. Identifying miRNAs that efficiently target genes and pathways is a key step in using these molecules for therapeutic purposes. Moreover, computational methods have been devised to help identify candidate miRNAs for each gene/pathway. MAPK and JAK/STAT pathways are known to have essential roles in cell proliferation and neoplastic transformation in different cancers including prostate cancer. Herein, we tried to identify miRNAs that target these pathways in the context of prostate cancer as therapeutic molecules. Genes involved in these pathways were analyzed with various algorithms to identify potentially targeting miRNAs. miR-23a and miR-23b were then selected as the best potential candidates that target a higher number of genes in these pathways with greater predictive scores. We then analyzed the expression of candidate miRNAs in LNCAP and PC3 cell lines as well as prostate cancer clinical samples. miR-23a and miR-23b showed a significant downregulation in cell line and tissue samples, a finding which is consistent with overactivation of these pathways in prostate cancer. In addition, we overexpressed miR-23a and miR-23b in LNCAP and PC3 cell lines, and these two miRNAs decreased IL-6R expression which has a critical role in these pathways. These results suggest the probability of utilizing miR-23a and miR-23b as therapeutic targets for the treatment of prostate cancer.





Similar content being viewed by others
References
Ferlay J SI, Ervik M, Dikshit R, Eser S, Mathers C, Rebelo M, Parkin DM, Forman D, Bray, F. Globocan 2012 v1.0, cancer incidence and mortality worldwide: IARC CancerBase no. 11; in Cancer IAfRo (ed), 2013.
Center MM, Jemal A, Lortet-Tieulent J, Ward E, Ferlay J, Brawley O, et al. International variation in prostate cancer incidence and mortality rates. Eur Urol. 2012;61:1079–92.
Lassi K, Dawson NA. Drug development for metastatic castration-resistant prostate cancer: current status and future perspectives. Future Oncol. 2011;7:551–8.
Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136:215–33.
Huntzinger E, Izaurralde E. Gene silencing by microRNAs: contributions of translational repression and mRNA decay. Nat Rev Genet. 2011;12:99–110.
Thorsen SB, Obad S, Jensen NF, Stenvang J, Kauppinen S. The therapeutic potential of microRNAs in cancer. Cancer J. 2012;18:275–84.
Gebert LF, Rebhan MA, Crivelli SE, Denzler R, Stoffel M, Hall J. Miravirsen (spc3649) can inhibit the biogenesis of mir-122. Nucleic Acids Res. 2014;42:609–21.
Bader AG, Brown D, Stoudemire J, Lammers P. Developing therapeutic microRNAs for cancer. Gene Ther. 2011;18:1121–6.
Bader AG. Mir-34—a microRNA replacement therapy is headed to the clinic. Front Genet. 2012;3:120.
Heyn H, Schreek S, Buurman R, Focken T, Schlegelberger B, Beger C. MicroRNA mir-548d is a superior regulator in pancreatic cancer. Pancreas. 2012;41:218–21.
Ritchie W, Rasko JE, Flamant S. MicroRNA target prediction and validation. Adv Exp Med Biol. 2013;774:39–53.
Vergoulis T, Vlachos IS, Alexiou P, Georgakilas G, Maragkakis M, Reczko M, et al. Tarbase 6.0: capturing the exponential growth of miRNA targets with experimental support. Nucleic Acids Res. 2012;40:D222–9.
Tan Gana NH, Victoriano AF, Okamoto T. Evaluation of online miRNA resources for biomedical applications. Genes Cells: Devoted Mol Cell Mech. 2012;17:11–27.
da Silva HB, Amaral EP, Nolasco EL, de Victo NC, Atique R, Jank CC, et al. Dissecting major signaling pathways throughout the development of prostate cancer. Prostate Cancer. 2013;2013:920612.
Wagner EF, Nebreda AR. Signal integration by JNK and p38 MAPK pathways in cancer development. Nat Rev Cancer. 2009;9:537–49.
Bradham C, McClay DR. P38 MAPK in development and cancer. Cell Cycle. 2006;5:824–8.
Kinkade CW, Castillo-Martin M, Puzio-Kuter A, Yan J, Foster TH, Gao H, et al. Targeting AKT/mTOR and ERK MAPK signaling inhibits hormone-refractory prostate cancer in a preclinical mouse model. J Clin Invest. 2008;118:3051–64.
Gioeli D, Mandell JW, Petroni GR, Frierson Jr HF, Weber MJ. Activation of mitogen-activated protein kinase associated with prostate cancer progression. Cancer Res. 1999;59:279–84.
Harrison DA. The jak/stat pathway. Cold Spring Harbor perspectives in biology 2012;4
Kwon EM, Holt SK, Fu R, Kolb S, Williams G, Stanford JL, et al. Androgen metabolism and JAK/STAT pathway genes and prostate cancer risk. Cancer Epidemiol. 2012;36:347–53.
Liu X, He Z, Li CH, Huang G, Ding C, Liu H. Correlation analysis of JAK-STAT pathway components on prognosis of patients with prostate cancer. Pathol Oncol Res: POR. 2012;18:17–23.
Aalinkeel R, Hu Z, Nair BB, Sykes DE, Reynolds JL, Mahajan SD, et al. Genomic analysis highlights the role of the JAK-STAT signaling in the anti-proliferative effects of dietary flavonoid-‘ashwagandha’ in prostate cancer cells. Evid Based Complement Alternat Med: eCAM. 2010;7:177–87.
Chiromatzo AO, Oliveira TY, Pereira G, Costa AY, Montesco CA, Gras DE, et al. miRNApath: a database of miRNAs, target genes and metabolic pathways. Genet Mol Res: GMR. 2007;6:859–65.
Vlachos IS, Kostoulas N, Vergoulis T, Georgakilas G, Reczko M, Maragkakis M, et al. DIANA miRPath v.2.0: investigating the combinatorial effect of microRNAs in pathways. Nucleic Acids Res. 2012;40:W498–504.
Ritchie W, Flamant S, Rasko JE. mimiRNA: a microRNA expression profiler and classification resource designed to identify functional correlations between microRNAs and their targets. Bioinformatics. 2010;26:223–7.
Mohammadi-Yeganeh S, Paryan M, Mirab Samiee S, Soleimani M, Arefian E, Azadmanesh K, et al. Development of a robust, low cost stem-loop real-time quantification PCR technique for miRNA expression analysis. Mol Biol Rep. 2013;40:3665–74.
Aghaee-Bakhtiari SH, Arefian E, Soleimani M, Noorbakhsh F, Samiee SM, Fard-Esfahani P, Mahdian R. Reproducible and reliable real-time pcr assay to measure mature form of mir-141. Appl Immunohistochem Mol Morphol 2015;Epub Ahead of Print
Bader AG, Brown D, Winkler M. The promise of microRNA replacement therapy. Cancer Res. 2010;70:7027–30.
Liu C, Kelnar K, Liu B, Chen X, Calhoun-Davis T, Li H, et al. The microRNA mir-34a inhibits prostate cancer stem cells and metastasis by directly repressing CD44. Nat Med. 2011;17:211–5.
Pulukuri SM, Gondi CS, Lakka SS, Jutla A, Estes N, Gujrati M, et al. RNA interference-directed knockdown of urokinase plasminogen activator and urokinase plasminogen activator receptor inhibits prostate cancer cell invasion, survival, and tumorigenicity in vivo. J Biol Chem. 2005;280:36529–40.
Walter BA, Valera VA, Pinto PA, Merino MJ. Comprehensive microRNA profiling of prostate cancer. J Cancer. 2013;4:350–7.
Coppola V, De Maria R, Bonci D. Micrornas and prostate cancer. Endocrine-related cancer 2010;17:F1-17
Majid S, Dar AA, Saini S, Arora S, Shahryari V, Zaman MS, et al. miR-23b represses proto-oncogene Src kinase and functions as methylation-silenced tumor suppressor with diagnostic and prognostic significance in prostate cancer. Cancer Res. 2012;72:6435–46.
Sun T, Yang M, Chen S, Balk S, Pomerantz M, Hsieh CL, et al. The altered expression of MiR-221/-222 and MiR-23b/-27b is associated with the development of human castration resistant prostate cancer. Prostate. 2012;72:1093–103.
Smith PC, Hobisch A, Lin DL, Culig Z, Keller ET. Interleukin-6 and prostate cancer progression. Cytokine Growth Factor Rev. 2001;12:33–40.
Culig Z, Steiner H, Bartsch G, Hobisch A. Interleukin-6 regulation of prostate cancer cell growth. J Cell Biochem. 2005;95:497–505.
Sakai I, Miyake H, Terakawa T, Fujisawa M. Inhibition of tumor growth and sensitization to chemotherapy by RNA interference targeting interleukin-6 in the androgen-independent human prostate cancer PC3 model. Cancer Sci. 2011;102:769–75.
Zhu LH, Liu T, Tang H, Tian RQ, Su C, Liu M, et al. MicroRNA-23a promotes the growth of gastric adenocarcinoma cell line MGC803 and downregulates interleukin-6 receptor. FEBS J. 2010;277:3726–34.
Acknowledgments
The authors appreciate the Pasteur Institute of Iran, Tehran, Iran for providing technical support. The authors are thankful to Vahid Haghpanah and Mohsen Khorashadizadeh for reading the manuscript and critical discussions. The authors are grateful to Zahra Masoumi and Fatemeh Kohram for editing the manuscript.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Funding
This study was funded by the Stem Cell Technology Research Center, Tehran, Iran.
Authors’ contributions
Seyed Hamid Aghaee-Bakhtiari carried out the practical tests and drafted the manuscript. Ehsan Arefian carried out the study design and drafting. Mahmood Naderi participated in the design of the study and drafting. Farshid Noorbakhsh participated in study design, data analysis, and draft preparation. Vahideh Nodouzi helped in clinical sample gathering and assisted in data analysis. Mojgan Asgari aided sample gathering and analyzed the stage of tumor samples. Pezhman Fard-Esfahani participated in data analysis and assisted in drafting the manuscript. Reza Mahdian helped in study design and aided in manuscript editing. Masoud Soleimani assisted in study design, data analysis, and draft preparation. All authors read and approved the final manuscript.
Conflicts of interest
None.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Aghaee-Bakhtiari, S.H., Arefian, E., Naderi, M. et al. MAPK and JAK/STAT pathways targeted by miR-23a and miR-23b in prostate cancer: computational and in vitro approaches. Tumor Biol. 36, 4203–4212 (2015). https://doi.org/10.1007/s13277-015-3057-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-015-3057-3