Abstract
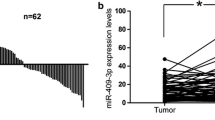
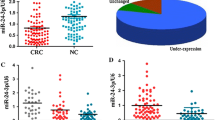
Aberrant expression of miR-720 had been reported in several cancers. However, the expression level and prognostic value of miR-720 in colorectal cancer (CRC) had not been addressed. In our study, we detected the expression level of miR-720 in 96 CRC tissues to evaluate its clinicopathological characteristics in colorectal cancer. Kaplan–Meier survival curve was performed to evaluate the prognostic role of miR-720 in patients with CRC. Furthermore, in vitro, we transfected the miR-720 mimics or inhibitors into the corresponding CRC cell lines and evaluated the effects on the abilities of cell growth, colony formation, migration, wound healing, and invasion in CRC cells. Our data showed that miR-720 level was significantly upregulated in CRC tissues than that in corresponding normal-appearing tissues (NATs) (p < 0.05), and high miR-720 correlated with the tumor size (p = 0.014), tumor–node–metastasis (TNM) stage (p = 0.040), lymphatic metastasis (p = 0.008), and distant metastasis (p = 0.016), which led to a poorer 5-year overall survival rate in CRC patients (p < 0.05). Our experiments in vitro also confirmed that miR-720 could promote the cell growth (p < 0.05), abilities of colony formation (p < 0.05), wound healing (p < 0.05), migration (p < 0.05), and invasion of CRC cells (p < 0.05). We identified StarD13 gene as a putative target of miR-720 in colorectal cancer by bioinformatics analysis, and subsequent dual luciferase activity and Western blot assay further certified that miR-720 might specifically target the StarD13 3′-untranslated region (UTR) at the 795 region (p < 0.05). miR-720 might act as a promoting factor in the development of CRC and could be a prognostic indicator in the prognosis of CRC. Downregulation of miR-720 might be considered to be a potentially important molecular treatment strategy for early stage CRC patients.




Similar content being viewed by others
References
Siegel R, Ward E, Brawley O, Jemal A. Cancer statistics, 2011: the impact of eliminating socioeconomic and racial disparities on premature cancer deaths. CA Cancer J Clin. 2011;61(4):212–36.
Karsa LV, Lignini TA, Patnick J, Lambert R, Sauvaget C. The dimensions of the CRC problem. Best Pract Res Clin Gastroenterol. 2010;24(4):381–96.
Peters U, Hutter CM, Hsu L, et al. Meta-analysis of new genome-wide association studies of colorectal cancer risk. Hum Genet. 2012;131(2):217–34.
Chan AT, Giovannucci EL. Primary prevention of colorectal cancer. Gastroenterology. 2010;138(6):2029–43. e2010.
Binefa G, Rodriguez-Moranta F, Teule A, Medina-Hayas M. Colorectal cancer: from prevention to personalized medicine. World J Gastroenterol. 2014;20(22):6786–808.
Lai EC. Micro RNAs are complementary to 3′ UTR sequence motifs that mediate negative post-transcriptional regulation. Nat Genet. 2002;30(4):363–4.
Iorio MV, Croce CM. Causes and consequences of microRNA dysregulation. Cancer J. 2012;18(3):215–22.
Kozomara A, Griffiths-Jones S. miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 2014;42(Database issue):D68–73.
Griffiths-Jones S, Grocock RJ, van Dongen S, Bateman A, Enright AJ. miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res. 2006;34(Database issue):D140–4.
Lewis BP, Burge CB, Bartel DP. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell. 2005;120(1):15–20.
Kunej T, Godnic I, Ferdin J, Horvat S, Dovc P, Calin GA. Epigenetic regulation of microRNAs in cancer: an integrated review of literature. Mutat Res. 2011;717(1–2):77–84.
Winter J, Jung S, Keller S, Gregory RI, Diederichs S. Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat Cell Biol. 2009;11(3):228–34.
Pizzini S, Bisognin A, Mandruzzato S, et al. Impact of microRNAs on regulatory networks and pathways in human colorectal carcinogenesis and development of metastasis. BMC Genomics. 2013;14:589.
Zhang L, Pickard K, Jenei V, et al. miR-153 supports colorectal cancer progression via pleiotropic effects that enhance invasion and chemotherapeutic resistance. Cancer Res. 2013;73(21):6435–47.
Hofsli E, Sjursen W, Prestvik WS, et al. Identification of serum microRNA profiles in colon cancer. Br J Cancer. 2013;108(8):1712–9.
Peng XH, Huang HR, Lu J, et al. MiR-124 suppresses tumor growth and metastasis by targeting Foxq1 in nasopharyngeal carcinoma. Mol Cancer. 2014;13(1):186.
Duan JH, Fang L. MicroRNA-92 promotes gastric cancer cell proliferation and invasion through targeting FXR. Tumour Biol 2014.
Muller V, Gade S, Steinbach B, et al. Changes in serum levels of miR-21, miR-210, and miR-373 in HER2-positive breast cancer patients undergoing neoadjuvant therapy: a translational research project within the Geparquinto trial. Breast Cancer Res Treat. 2014;147(1):61–8.
Li LZ, Zhang CZ, Liu LL, et al. miR-720 inhibits tumor invasion and migration in breast cancer by targeting TWIST1. Carcinogenesis. 2014;35(2):469–78.
Jones CI, Zabolotskaya MV, King AJ, et al. Identification of circulating microRNAs as diagnostic biomarkers for use in multiple myeloma. Br J Cancer. 2012;107(12):1987–96.
Shinozuka E, Miyashita M, Mizuguchi Y, et al. SnoN/SKIL modulates proliferation through control of hsa-miR-720 transcription in esophageal cancer cells. Biochem Biophys Res Commun. 2013;430(1):101–6.
Sand M, Skrygan M, Sand D, et al. Comparative microarray analysis of microRNA expression profiles in primary cutaneous malignant melanoma, cutaneous malignant melanoma metastases, and benign melanocytic nevi. Cell Tissue Res. 2013;351(1):85–98.
Ragusa M, Statello L, Maugeri M, et al. Specific alterations of the microRNA transcriptome and global network structure in colorectal cancer after treatment with MAPK/ERK inhibitors. J Mol Med (Berl). 2012;90(12):1421–38.
Lerebours F, Cizeron-Clairac G, Susini A, et al. miRNA expression profiling of inflammatory breast cancer identifies a 5-miRNA signature predictive of breast tumor aggressiveness. Int J Cancer. 2013;133(7):1614–23.
Gupta GP, Massague J. Cancer metastasis: building a framework. Cell. 2006;127(4):679–95.
Eccles SA, Welch DR. Metastasis: recent discoveries and novel treatment strategies. Lancet. 2007;369(9574):1742–57.
Ching YP, Wong CM, Chan SF, et al. Deleted in liver cancer (DLC) 2 encodes a RhoGAP protein with growth suppressor function and is underexpressed in hepatocellular carcinoma. J Biol Chem. 2003;278(12):10824–30.
El-Sitt S, Khalil BD, Hanna S, El-Sabban M, Fakhreddine N, El-Sibai M. DLC2/StarD13 plays a role of a tumor suppressor in astrocytoma. Oncol Rep. 2012;28(2):511–8.
Hanna S, Khalil B, Nasrallah A, et al. StarD13 is a tumor suppressor in breast cancer that regulates cell motility and invasion. Int J Oncol. 2014;44(5):1499–511.
Tang F, Zhang R, He Y, Zou M, Guo L, Xi T. MicroRNA-125b induces metastasis by targeting STARD13 in MCF-7 and MDA-MB-231 breast cancer cells. PLoS One. 2012;7(5):e35435.
Nasrallah A, Saykali B, Al Dimassi S, Khoury N, Hanna S, El-Sibai M. Effect of StarD13 on colorectal cancer proliferation, motility and invasion. Oncol Rep. 2014;31(1):505–15.
Acknowledgments
This study was supported by a grant from the National Youthful Science Foundation of China (Nos. 81101858 and 81302147); the National Science Foundation of Jiangsu Province, China (No. BK20130270); and the Science and Education Youth Health Foundation, Suzhou, China (No. KJXW2012005).
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding authors
Additional information
Xu Wang, Yuting Kuang, and Xiaochun Shen contributed equally to this work.
Rights and permissions
About this article
Cite this article
Wang, X., Kuang, Y., Shen, X. et al. Evaluation of miR-720 prognostic significance in patients with colorectal cancer. Tumor Biol. 36, 719–727 (2015). https://doi.org/10.1007/s13277-014-2697-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-014-2697-z