Abstract
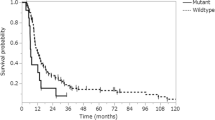
The prognosis of high-grade glioma patients is poor, and the tumors are characterized by resistance to therapy. The aims of this study were to analyze the prognostic value of the expression of the protein tyrosine phosphatase non-receptor type 6 (PTPN6, also referred to as SHP1) in high-grade glioma patients, the epigenetic regulation of the expression of PTPN6, and the role of its expression in chemotherapy resistance in glioma-derived cells. PTPN6 expression was analyzed with immunohistochemistry in 89 high-grade glioma patients. Correlation between PTPN6 expression and overall survival was analyzed with Kaplan-Meier univariate analysis and Cox regression multivariate analysis. Differences in drug sensitivity to a panel of 16 chemotherapeutic drugs between PTPN6-overexpressing clones and control clones were analyzed in vitro with the fluorometric microculture cytotoxicity assay. Cell cycle analysis was done with Krishan staining and flow cytometry. Apoptosis was analyzed with a cell death detection ELISA kit as well as cleaved caspase-3 and caspase-9 Western blotting. Autophagy was analyzed with LC3B Western blotting. Methylation of the PTPN6 promoter was analyzed with bisulfite pyrosequencing, and demethylation of PTPN6 was done with decitabine treatment. The PTPN6 expression correlated in univariate analysis to poor survival for anaplastic glioma patients (p = 0.026). In glioma-derived cell lines, overexpression of PTPN6 caused increase resistance (p < 0.05) to the chemotherapeutic drugs bortezomib, cisplatin, and melphalan. PTPN6 expression did not affect bortezomib-induced cell cycle arrest, apoptosis, or autophagy. Low PTPN6 promoter methylation correlated to protein expression, and the protein expression was increased upon demethylation in glioma-derived cells. PTPN6 expression may be a factor contributing to poor survival for anaplastic glioma patients, and in glioma-derived cells, its expression is epigenetically regulated and influences the response to chemotherapy.





Similar content being viewed by others
References
Louis DN. Molecular pathology of malignant gliomas. Ann Rev Pat. 2006;1:97–117. doi:10.1146/annurev.pathol.1.110304.100043.
Stupp R, Mason WP, van den Bent MJ, Weller M, Fisher B, Taphoorn MJ, Belanger K, Brandes AA, Marosi C, Bogdahn U, Curschmann J, Janzer RC, Ludwin SK, Gorlia T, Allgeier A, Lacombe D, Cairncross JG, Eisenhauer E, Mirimanoff RO. European Organisation for R, Treatment of Cancer Brain T, Radiotherapy G, National Cancer Institute of Canada Clinical Trials G Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med. 2005;352(10):987–996. doi:10.1056/NEJMoa043330.
Stupp R, Tonn JC, Brada M, Pentheroudakis G, Group EGW. High-grade malignant glioma: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol. 2010;21 Suppl 5:v190–3. doi:10.1093/annonc/mdq187.
Wu C, Sun M, Liu L, Zhou GW. The function of the protein tyrosine phosphatase SHP-1 in cancer. Gene. 2003;306:1–12.
Insabato L, Amelio I, Quarto M, Zannetti A, Tolino F, de Mauro G, et al. Elevated expression of the tyrosine phosphatase SHP-1 defines a subset of high-grade breast tumors. Oncology. 2009;77(6):378–84. doi:10.1159/000276765.
Xu SB, Liu XH, Li BH, Zhang Y, Yuan J, Yuan Q, et al. DNA methylation regulates constitutive expression of Stat6 regulatory genes SOCS-1 and SHP-1 in colon cancer cells. J Cancer Res Clin Oncol. 2009;135(12):1791–8. doi:10.1007/s00432-009-0627-z.
Samanta AK, Chakraborty SN, Wang Y, Kantarjian H, Sun X, Hood J, et al. Jak2 inhibition deactivates Lyn kinase through the SET-PP2A-SHP1 pathway, causing apoptosis in drug-resistant cells from chronic myelogenous leukemia patients. Oncogene. 2009;28(14):1669–81. doi:10.1038/onc.2009.7.
Rajendran P, Li F, Manu KA, Shanmugam MK, Loo SY, Kumar AP, et al. Gamma-tocotrienol is a novel inhibitor of constitutive and inducible STAT3 signalling pathway in human hepatocellular carcinoma: potential role as an antiproliferative, pro-apoptotic and chemosensitizing agent. Br J Pharmacol. 2011;163(2):283–98. doi:10.1111/j.1476-5381.2010.01187.x.
Esteller M. The necessity of a human epigenome project. Carcinogenesis. 2006;27(6):1121–5. doi:10.1093/carcin/bgl033.
Kawakami K, Matsunoki A, Kaneko M, Saito K, Watanabe G, Minamoto T. Long interspersed nuclear element-1 hypomethylation is a potential biomarker for the prediction of response to oral fluoropyrimidines in microsatellite stable and CpG island methylator phenotype-negative colorectal cancer. Cancer Sci. 2011;102(1):166–74. doi:10.1111/j.1349-7006.2010.01776.x.
Dejeux E, Ronneberg JA, Solvang H, Bukholm I, Geisler S, Aas T, et al. DNA methylation profiling in doxorubicin treated primary locally advanced breast tumours identifies novel genes associated with survival and treatment response. Mol Cancer. 2010;9:68. doi:10.1186/1476-4598-9-68.
Hegi ME, Diserens AC, Godard S, Dietrich PY, Regli L, Ostermann S, et al. Clinical trial substantiates the predictive value of O-6-methylguanine-DNA methyltransferase promoter methylation in glioblastoma patients treated with temozolomide. Clin Cancer Res. 2004;10(6):1871–4.
Rivera AL, Pelloski CE, Gilbert MR, Colman H, De La Cruz C, Sulman EP, et al. MGMT promoter methylation is predictive of response to radiotherapy and prognostic in the absence of adjuvant alkylating chemotherapy for glioblastoma. Neuro Oncol. 2010;12(2):116–21. doi:10.1093/neuonc/nop020.
Zhang Q, Wang HY, Marzec M, Raghunath PN, Nagasawa T, Wasik MA. STAT3- and DNA methyltransferase 1-mediated epigenetic silencing of SHP-1 tyrosine phosphatase tumor suppressor gene in malignant T lymphocytes. Proc Natl Acad Sci U S A. 2005;102(19):6948–53. doi:10.1073/pnas.0501959102.
Koyama M, Oka T, Ouchida M, Nakatani Y, Nishiuchi R, Yoshino T, et al. Activated proliferation of B-cell lymphomas/leukemias with the SHP1 gene silencing by aberrant CpG methylation. Lab Investig. 2003;83(12):1849–58.
Kampf C, Olsson I, Ryberg U, Sjostedt E, Ponten F. Production of tissue microarrays, immunohistochemistry staining and digitalization within the human protein atlas. Journal of Visualized Experiments. 2012(63). doi:10.3791/3620.
Popova S, Bergqvist M, Dimberg A, Edqvist P-H, Ekman S, Hesselager G, et al. Molecular classification of gliomas of various WHO grades applying immunohistochemistry. Histopathology. 2013. doi:10.1111/his.12252.
Wickstrom M, Johnsen JI, Ponthan F, Segerstrom L, Sveinbjornsson B, Lindskog M, et al. The novel melphalan prodrug J1 inhibits neuroblastoma growth in vitro and in vivo. Mol Cancer Ther. 2007;6(9):2409–17. doi:10.1158/1535-7163.MCT-07-0156.
Krishan A. Rapid flow cytofluorometric analysis of mammalian cell cycle by propidium iodide staining. J Cell Biol. 1975;66(1):188–93.
Ruchusatsawat K, Wongpiyabovorn J, Shuangshoti S, Hirankarn N, Mutirangura A. SHP-1 promoter 2 methylation in normal epithelial tissues and demethylation in psoriasis. J Mol Med. 2006;84(2):175–82. doi:10.1007/s00109-005-0020-6.
Lennartsson J, Wardega P, Engstrom U, Hellman U, Heldin CH. Alix facilitates the interaction between c-Cbl and platelet-derived growth factor beta-receptor and thereby modulates receptor down-regulation. J Biol Chem. 2006;281(51):39152–8. doi:10.1074/jbc.M608489200.
Uhlen M, Oksvold P, Fagerberg L, Lundberg E, Jonasson K, Forsberg M, et al. Towards a knowledge-based human protein atlas. Nat Biotechnol. 2010;28(12):1248–50. doi:10.1038/nbt1210-1248.
Chanida V, Poonchavist C, Virote S, Apiwat M. The role of SHP-1 promoter 2 hypermethylation detection of lymph node micrometastasis in resectable stage I non-small cell lung cancer as a prognostic marker of disease recurrence. Int J Clin Oncol. 2013. doi:10.1007/s10147-013-0590-1.
Zhang Y, Zhao D, Zhao H, Wu X, Zhao W, Wang Y, et al. Hypermethylation of SHP-1 promoter in patient with high-risk myelodysplastic syndrome and it predicts poor prognosis. Med Oncol. 2012;29(4):2359–63. doi:10.1007/s12032-012-0163-6.
Keilhack H, Muller M, Bohmer SA, Frank C, Weidner KM, Birchmeier W, et al. Negative regulation of Ros receptor tyrosine kinase signaling. An epithelial function of the SH2 domain protein tyrosine phosphatase SHP-1. J Cell Biol. 2001;152(2):325–34.
Rodriguez-Ubreva FJ, Cariaga-Martinez AE, Cortes MA, Romero-De Pablos M, Ropero S, Lopez-Ruiz P, et al. Knockdown of protein tyrosine phosphatase SHP-1 inhibits G1/S progression in prostate cancer cells through the regulation of components of the cell-cycle machinery. Oncogene. 2010;29(3):345–55. doi:10.1038/onc.2009.329.
Montano X. Repression of SHP-1 expression by p53 leads to trkA tyrosine phosphorylation and suppression of breast cancer cell proliferation. Oncogene. 2009;28(43):3787–800. doi:10.1038/onc.2009.143.
Youssef G, Gillett C, Agbaje O, Crompton T, Montano X. Phosphorylation of NTRK1 at Y674/Y675 induced by TP53-dependent repression of PTPN6 expression: a potential novel prognostic marker for breast cancer. Mod Pathol. 2013. doi:10.1038/modpathol.2013.129.
Mena-Duran AV, Togo SH, Bazhenova L, Cervera J, Bethel K, Senent ML, et al. SHP1 expression in bone marrow biopsies of myelodysplastic syndrome patients: a new prognostic factor. Br J Haematol. 2005;129(6):791–4. doi:10.1111/j.1365-2141.2005.05516.x.
Wang N, Li Z, Ding R, Frank GD, Senbonmatsu T, Landon EJ, et al. Antagonism or synergism. Role of tyrosine phosphatases SHP-1 and SHP-2 in growth factor signaling. J Biol Chem. 2006;281(31):21878–83.
Forget G, Gregory DJ, Whitcombe LA, Olivier M. Role of host protein tyrosine phosphatase SHP-1 in Leishmania donovani-induced inhibition of nitric oxide production. Infect Immun. 2006;74(11):6272–9. doi:10.1128/IAI.00853-05.
Maiti AK. Genetic determinants of oxidative stress-mediated sensitization of drug-resistant cancer cells. Int J Cancer. 2012;130(1):1–9. doi:10.1002/ijc.26306.
Berg KL, Siminovitch KA, Stanley ER. SHP-1 regulation of p62(DOK) tyrosine phosphorylation in macrophages. J Biol Chem. 1999;274(50):35855–65.
Acknowledgments
The authors would like to thank Prof. Frank Böhmer, Jena University, for kindly providing the pRK5PTPN6 vector; Dr. H Hedman, Umea University, the UCSF Tissue Bank, and Dr. JS Guillamo for providing us with the glioma cell lines used in the experiments; and the Science for Life Laboratory BioVis Technology Platform in Uppsala for supporting the flow cytometry analyses. We would also like to express our gratitude for the financial support from the Cancer Foundation at Gavle Hospital, the Research Fund at the Department of Oncology, Uppsala University Hospital, the Swedish Cancer Society, the Swedish Research Council, and the Knut and Alice Wallenberg Foundation.
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Online resource 1
Mean dose response curves from FMCA analysis for MGMT and PTPN6 overexpressing clones and control clones. PTPN6 clones (n = 5) were less sensitive than control clones (n = 5) to the drugs bortezomib (A), cisplatin (B) and melphalan (C). (JPEG 41 kb)
Online resource 2
Mean cell cycle distribution of three PTPN6 and three control clones with or without bortezomib treatment. Bortezomib increased the percentage of cells in G0/G1 phase (p < 0.05) and decreased the percentage of cells in S and G2/M phases (p < 0.05) after 72 h in both PTPN6 overexpressing clones and control clones. (JPEG 22 kb)
Rights and permissions
About this article
Cite this article
Sooman, L., Ekman, S., Tsakonas, G. et al. PTPN6 expression is epigenetically regulated and influences survival and response to chemotherapy in high-grade gliomas. Tumor Biol. 35, 4479–4488 (2014). https://doi.org/10.1007/s13277-013-1590-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-013-1590-5