Abstract
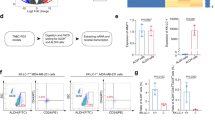
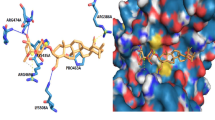
Targeting breast cancer stem cells (BCSCs) offers a promising strategy for breast cancer treatment. We examined the plant alkaloid ellipticine for its efficacy to inhibit the expression of aldehyde dehydrogenase 1 class A1 (ALDH1A1)-positive BCSCs by in vitro and in silico methods. At 3 mM concentration, ellipticine decreased the expression of ALDH1A1-positive BCSCs by 62 % (p = 0.073) in the MCF7 cell line and by 53 % (p = 0.024) in the SUM159 cell line compared to vehicle-treated cultures. Ellipticine significantly reduced the formation of mammospheres, whereas paclitaxel enhanced mammosphere formation in both the treated cell lines. Interestingly, when treated with a combination of ellipticine and paclitaxel, the percentage of ALDH1A1-positive BCSCs dropped by several fold in vitro. A homology model of Homo sapiens ALDH1A1 was built using the crystal structure of NAD-bound sheep liver class I aldehyde dehydrogenase [PDB ID: 1BXS] as a template. Molecular simulation and docking studies revealed that the amino acids Asn-117 and Asn-121, Glu-249, Cys-302, and Gln-350, present in the active site of human ALDH1A1, played a vital role in interacting with the drug. The present study suggests that ellipticine reduces the proliferation and self-renewal ability of ALDH1A1-positive BCSCs and can be used in combination with a cytotoxic drug like paclitaxel for potential targeting of BCSCs.






Similar content being viewed by others
References
National Cancer Registry Programme. http://ncrpindia.org/PBCR_2006_2008/Chapter_2.pdf
Liu S, Dontu G, Wicha MS. Mammary stem cells, self-renewal pathways, and carcinogenesis. Breast Cancer Res. 2005;7:86–95.
Korkaya H, Paulson A, Charafe-Jauffret E, Ginestier C, Brown M, Dutcher J, et al. Regulation of mammary stem/progenitor cells by PTEN/Akt/beta-catenin signaling. PLoS Biol. 2009;7:e1000121.
Dick JE. Breast cancer stem cells revealed. Proc Natl Acad Sci U S A. 2003;100:3547–9.
Al-Hajj M, Becker MW, Wicha M, Weissman I, Clarke MF. Therapeutic implications of cancer stem cells. Curr Opin Genet Dev. 2004;14:43–7.
Dontu G, Al-Hajj M, Abdallah WM, Clarke MF, Wicha MS. Stem cells in normal breast development and breast cancer. Cell Prolif. 2003;36 Suppl 1:59–72.
Pardal R, Clarke MF, Morrison SJ. Applying the principles of stem-cell biology to cancer. Nat Rev Cancer. 2003;3:895–902.
Smalley M, Ashworth A. Stem cells and breast cancer: a field in transit. Nat Rev Cancer. 2003;3:832–44.
Al-Hajj M, Wicha MS, Benito-Hernandez A, Morrison SJ, Clarke MF. Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci U S A. 2003;100:3983–8.
Croker AK, Goodale D, Chu J, Postenka C, Hedley BD, Hess DA, et al. High aldehyde dehydrogenase and expression of cancer stem cell markers selects for breast cancer cells with enhanced malignant and metastatic ability. J Cell Mol Med. 2009;13:2236–52.
Ucar D, Cogle CR, Zucali JR, Ostmark B, Scott EW, Zori R, et al. Aldehyde dehydrogenase activity as a functional marker for lung cancer. Chem Biol Interact. 2009;178:48–55.
Corti S, Locatelli F, Papadimitriou D, Donadoni C, Salani S, Del Bo R, et al. Identification of a primitive brain-derived neural stem cell population based on aldehyde dehydrogenase activity. Stem Cells. 2006;24:975–85.
Vasiliou V, Nebert DW. Analysis and update of the human aldehyde dehydrogenase (ALDH) gene family. Hum Genomics. 2005;2:138–43.
Armstrong L, Stojkovic M, Dimmick I, Ahmad S, Stojkovic P, Hole N, et al. Phenotypic characterization of murine primitive hematopoietic progenitor cells isolated on basis of aldehyde dehydrogenase activity. Stem Cells. 2004;22:1142–51.
Hess DA, Meyerrose TE, Wirthlin L, Craft TP, Herrbrich PE, Creer MH, et al. Functional characterization of highly purified human hematopoietic repopulating cells isolated according to aldehyde dehydrogenase activity. Blood. 2004;104:1648–55.
Hess DA, Wirthlin L, Craft TP, Herrbrich PE, Hohm SA, Lahey R, et al. Selection based on CD133 and high aldehyde dehydrogenase activity isolates long-term reconstituting human hematopoietic stem cells. Blood. 2006;107:2162–9.
Matsui W, Huff CA, Wang Q, Malehorn MT, Barber J, Tanhehco Y, et al. Characterization of clonogenic multiple myeloma cells. Blood. 2004;103:2332–6.
Goodwin S, Smith AF. Alkaloids of Ochrosia elliptica Labill. J Am Chem Soc. 1959;81:1903–8.
Stiborova M. The anticancer drug ellipticine forms covalent DNA adducts, mediated by human cytochromes P450, through metabolism to 13-hydroxyellipticine and ellipticine N2-oxide. Cancer Res. 2004;64:8374–80.
Stiborova M, Rupertova M, Schmeiser HH, Frei E. Molecular mechanisms of antineoplastic action of an anticancer drug ellipticine. Biomed Pap Med Fac Univ Palacky Olomouc Czech Repub. 2006;150:13–23.
Kuo PL, Hsu YL, Kuo YC, Chang CH, Lin CC. The anti-proliferative inhibition of ellipticine in human breast mda-mb-231 cancer cells is through cell cycle arrest and apoptosis induction. Anticancer Drugs. 2005;16:789–95.
Poljakova J, Eckschlager T, Hrabeta J, Hrebackova J, Smutny S, Frei E, et al. The mechanism of cytotoxicity and DNA adduct formation by the anticancer drug ellipticine in human neuroblastoma cells. Biochem Pharmacol. 2009;77:1466–79.
Martinkova E, Dontenwill M, Frei E, Stiborova M. Cytotoxicity of and DNA adduct formation by ellipticine in human U87MG glioblastoma cancer cells. Neuro Endocrinol Lett. 2009;30 Suppl 1:60–6.
Auclair C. Multimodal action of antitumor agents on DNA: the ellipticine series. Arch Biochem Biophys. 1987;259:1–14.
Huang C, Zhang XM, Tavaluc RT, Hart LS, Dicker DT, Wang W, et al. The combination of 5-fluorouracil plus p53 pathway restoration is associated with depletion of p53-deficient or mutant p53-expressing putative colon cancer stem cells. Cancer Biol Ther. 2009;8:2186–93.
Dontu G, Abdallah WM, Foley JM, Jackson KW, Clarke MF, Kawamura MJ, et al. In vitro propagation and transcriptional profiling of human mammary stem/progenitor cells. Genes Dev. 2003;17:1253–70.
Ginestier C, Hur MH, Charafe-Jauffret E, Monville F, Dutcher J, Brown M, et al. ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell. 2007;1:555–67.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–10.
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–402.
Chenna R. Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Res. 2003;31:3497–500.
Sali A, Blundell TL. Comparative protein modelling by satisfaction of spatial restraints. J Mol Biol. 1993;234:779–815.
Guex N, Peitsch MC. SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis. 1997;18:2714–23.
Laskowski RA, MacArthur MW, Moss DS, Thornton JM. PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr. 1993;26:283–91.
Vriend G. WHAT IF: a molecular modeling and drug design program. J Mol Graph. 1990;8:52–6. 29.
Sippl MJ. Recognition of errors in three-dimensional structures of proteins. Proteins. 1993;17:355–62.
Lüthy R, Bowie JU, Eisenberg D. Assessment of protein models with three-dimensional profiles. Nature. 1992;356:83–5.
Bowie JU, Lüthy R, Eisenberg D. A method to identify protein sequences that fold into a known three-dimensional structure. Science. 1991;253:164–70.
Laskowski RA, Chistyakov VV, Thornton JM. Pdbsum more: new summaries and analyses of the known 3D structures of proteins and nucleic acids. Nucleic Acids Res. 2005;33:D266–268.
Lindal E, Hess B. Gromacs 3.0: a package for molecular simulation and trajectory analysis. J Mol Model. 2001;7:306–17.
Berendsen H. Interaction models for water in relation to protein hydration. In: Pullman B, editor. Intermolecular forces. Dordrecht: Reidel; 1981. p. 331–42.
DeLano WL. The PyMOL molecular graphics system. SanCarlos: DeLano Scientific. http://wwwpymolorg (2006).
Fosse P, Rene B, Charra M, Paoletti C, Saucier JM. Stimulation of topoisomerase II-mediated DNA cleavage by ellipticine derivatives: structure–activity relationship. Mol Pharmacol. 1992;42:590–5.
Li X, Lewis MT, Huang J, Gutierrez C, Osborne CK, Wu MF, et al. Intrinsic resistance of tumorigenic breast cancer cells to chemotherapy. J Natl Cancer Inst. 2008;100:672–9.
Yu F, Yao H, Zhu P, Zhang X, Pan Q, Gong C, et al. let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell. 2007;131:1109–23.
Charafe-Jauffret E, Monville F, Ginestier C, Dontu G, Birnbaum D, Wicha MS. Cancer stem cells in breast: current opinion and future challenges. Pathobiology. 2008;75:75–84.
Ohashi M, Sugikawa E, Nakanishi N. Inhibition of p53 protein phosphorylation by 9-hydroxyellipticine: a possible anticancer mechanism. Jpn J Cancer Res. 1995;86:819–27.
Stiborova M, Bieler CA, Wiessler M, Frei E. The anticancer agent ellipticine on activation by cytochrome P450 forms covalent DNA adducts. Biochem Pharmacol. 2001;62:1675–84.
Poljakova J, Frei E, Gomez JE, Aimova D, Eckschlager T, Hrabeta J, et al. DNA adduct formation by the anticancer drug ellipticine in human leukemia HL-60 and CCRF-CEM cells. Cancer Lett. 2007;252:270–9.
Tian E, Landowski TH, Stephens OW, Yaccoby S, Barlogie B, Shaughnessy Jr JD. Ellipticine derivative NSC 338258 represents a potential new antineoplastic agent for the treatment of multiple myeloma. Mol Cancer Ther. 2008;7:500–9.
Fang K, Chen SP, Lin CW, Cheng WC, Huang HT. Ellipticine-induced apoptosis depends on Akt translocation and signaling in lung epithelial cancer cells. Lung Cancer. 2009;63:227–34.
Juret P, Couette JE, Delozier T, Le Talaer JY. [Hydroxy-9-methyl-2-ellipticinium (NSC 264-137) for osseous metastases from breast cancer. A 4 year experience (author’s transl)]. Bull Cancer. 1981;68:224–31.
Peng Y, Li C, Chen L, Sebti S, Chen J. Rescue of mutant p53 transcription function by ellipticine. Oncogene. 2003;22:4478–87.
Wang X, Wu X, Wang C, Zhang W, Ouyang Y, Yu Y, He Z. Transcriptional suppression of breast cancer resistance protein (BCRP) by wild-type p53 through the NF-kappaB pathway in MCF-7 cells. FEBS Lett. 2010;584:3392–7.
Godar S, Ince TA, Bell GW, Feldser D, Donaher JL, Bergh J, et al. Growth-inhibitory and tumor-suppressive functions of p53 depend on its repression of CD44 expression. Cell. 2008;134:62–73.
Phillips TM, McBride WH, Pajonk F. The response of CD24(−/low)/CD44+ breast cancer-initiating cells to radiation. J Natl Cancer Inst. 2006;98:1777–85.
Moreb JS, Baker HV, Chang L-J, Amaya M, Lopez MC, Ostmark B, et al. ALDH isozymes downregulation affects cell growth, cell motility and gene expression in lung cancer cells. Mol Cancer. 2008;7:87–87.
Acknowledgments
At the Tumor Biology Laboratory, National Institute of Pathology, New Delhi, India, we wish to thank Dr. Sourav Verma and Ms. Anvesha Malik for their help in the flow cytometry experiments. We also thank the BD Biosciences Flow Cytometry Core Group for the excellent support in cell sorting. SLP and PSC want to thank the Indian Council of Medical Research, New Delhi, Government of India for the financial support and RSC gratefully acknowledges the University Grant Commission, New Delhi, India for his fellowship.
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pandrangi, S.L., Chikati, R., Chauhan, P.S. et al. Effects of ellipticine on ALDH1A1-expressing breast cancer stem cells—an in vitro and in silico study. Tumor Biol. 35, 723–737 (2014). https://doi.org/10.1007/s13277-013-1099-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-013-1099-y