Abstract
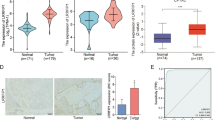
Osteosarcoma (OS) is the most common primary malignant bone tumor in children and adolescents. To identify new biomarkers for early diagnosis of OS and novel therapeutic candidates, we carried out a plasma membrane proteomic study based on two-dimensional electrophoresis (2DE). The OS cell line MG-63 and the human osteoblastic cell line hFOB1.19 were adopted as the comparison model. We extracted plasma membrane by aqueous two-phase partition extraction. The proteins were separated through 2DE. We analyzed the differentially expressed proteins by Imagemaster software and then identified them by liquid chromatography–tandem mass spectrometry, and the location and function of differential proteins were searched through the Gene Ontology database. In total, 220 protein spots were separated by 2DE. Seven proteins with more than 2.0-folds of difference were successfully identified from 13 gel spots, with 6 up-regulated and 1 down-regulated. Gene Ontology analysis of the differentially expressed proteins indicated that these proteins were involved in seven kinds of functions including binding, structural, cell motility, receptor activity, electron carrier activity, NADH dehydrogenase (ubiquinone) activity, and transcription repressor activity. The up-regulation of NDRG1 was verified in osteosarcoma through Western blotting and by immunohistochemistry in paraffin-embedded tissues. The plasma membrane proteins identified in this study may provide new insights into osteosarcoma cancer biology and potential diagnostic and therapeutic biomarkers.





Similar content being viewed by others
Abbreviations
- OS:
-
Osteosarcoma
- 2DE:
-
Two-dimensional electrophoresis
- LC–MS:
-
Liquid chromatography–mass spectrometry
- MS/MS:
-
Tandem MS
- HCT:
-
High capacity trap
- HE:
-
Hematoxylin and eosin
References
Mankin HJ, Hornicek FJ, Rosenberg AE, Harmon DC, Gebhardt MC. Survival data for 648 patients with osteosarcoma treated at one institution. Clin Orthop Relat Res. 2004;(429):286–91.
Leth-Larsen R, Lund RR, Ditzel HJ. Plasma membrane proteomics and its application in clinical cancer biomarker discovery. Mol Cell Proteomics. 2010;9:1369–82.
Sprenger RR, Jensen ON. Proteomics and the dynamic plasma membrane: quo vadis? Proteomics. 2010;10:3997–4011.
Bhattacharyya S, Byrum S, Siegel ER, Suva LJ. Proteomic analysis of bone cancer: a review of current and future developments. Expert Rev Proteomics. 2007;4:371–8.
Byrum S, Montgomery CO, Nicholas RW, Suva LJ. The promise of bone cancer proteomics. Ann N Y Acad Sci. 2010;1192:222–9.
Folio C, Mora MI, Zalacain M, Corrales FJ, Segura V, Sierrasesumaga L, et al. Proteomic analysis of chemonaive pediatric osteosarcomas and corresponding normal bone reveals multiple altered molecular targets. J Proteome Res. 2009;8:3882–8.
Guo QC, Shen JN, Jin S, Wang J, Huang G, Zhang LJ, et al. Comparative proteomic analysis of human osteosarcoma and sv40-immortalized normal osteoblastic cell lines. Acta Pharmacol Sin. 2007;28:850–8.
Zhang Z, Zhang L, Hua Y, Jia X, Li J, Hu S, et al. Comparative proteomic analysis of plasma membrane proteins between human osteosarcoma and normal osteoblastic cell lines. BMC Cancer. 2010;10:206.
Cao R, Li X, Liu Z, Peng X, Hu W, Wang X, et al. Integration of a two-phase partition method into proteomics research on rat liver plasma membrane proteins. J Proteome Res. 2006;5:634–42.
Zhang L, Jia X, Zhang X, Sun J, Peng X, Qi T, et al. Proteomic analysis of PBMCs: characterization of potential HIV-associated proteins. Proteome Sci. 2010;8:12.
Zhang L, Jia X, Zhang X, Cao J, Yang P, Qiu C, et al. Alpha-1 antitrypsin variants in plasma from HIV-infected patients revealed by proteomic and glycoproteomic analysis. Electrophoresis. 2010;31:3437–45.
Zhang L, Xie J, Wang X, Liu X, Tang X, Cao R, et al. Proteomic analysis of mouse liver plasma membrane: use of differential extraction to enrich hydrophobic membrane proteins. Proteomics. 2005;5:4510–24.
de Alava E. Molecular pathology in sarcomas. Clin Transl Oncol. 2007;9:130–44.
Dass CR, Ek ET, Choong PF. PEDF as an emerging therapeutic candidate for osteosarcoma. Curr Cancer Drug Targets. 2008;8:683–90.
Nikitovic D, Berdiaki K, Chalkiadaki G, Karamanos N, Tzanakakis G. The role of SLRP-proteoglycans in osteosarcoma pathogenesis. Connect Tissue Res. 2008;49:235–8.
Zhang L, Peng X, Zhang Z, Feng Y, Jia X, Shi Y, et al. Subcellular proteome analysis unraveled annexin A2 related to immune liver fibrosis. J Cell Biochem. 2010;110:219–28.
Zhang L, Jia X, Feng Y, Peng X, Zhang Z, Zhou W, et al. Plasma membrane proteome analysis of the early effect of alcohol on liver: implications for alcoholic liver disease. Acta Biochim Biophys Sin (Shanghai). 2011;43:19–29.
Lubke T, Lobel P, Sleat DE. Proteomics of the lysosome. Biochim Biophys Acta. 2009;1793:625–35.
Jung EU, Yoon JH, Lee YJ, Lee JH, Kim BH, Yu SJ, et al. Hypoxia and retinoic acid-inducible NDRG1 expression is responsible for doxorubicin and retinoic acid resistance in hepatocellular carcinoma cells. Cancer Lett. 2010;298:9–15.
Sibold S, Roh V, Keogh A, Studer P, Tiffon C, Angst E, et al. Hypoxia increases cytoplasmic expression of NDRG1, but is insufficient for its membrane localization in human hepatocellular carcinoma. FEBS Lett. 2007;581:989–94.
Angst E, Sibold S, Tiffon C, Weimann R, Gloor B, Candinas D, et al. Cellular differentiation determines the expression of the hypoxia-inducible protein NDRG1 in pancreatic cancer. Br J Cancer. 2006;95:307–13.
Cangul H. Hypoxia upregulates the expression of the NDRG1 gene leading to its overexpression in various human cancers. BMC Genet. 2004;5:27.
Gerhard R, Nonogaki S, Fregnani JH, Soares FA, Nagai MA. Ndrg1 protein overexpression in malignant thyroid neoplasms. Clinics (Sao Paulo). 2010;65:757–62.
Matsugaki T, Zenmyo M, Hiraoka K, Fukushima N, Shoda T, Komiya S, et al. N-myc downstream-regulated gene 1/Cap43 expression promotes cell differentiation of human osteosarcoma cells. Oncol Rep. 2010;24:721–5.
Wu M, Bai X, Xu G, Wei J, Zhu T, Zhang Y, et al. Proteome analysis of human androgen-independent prostate cancer cell lines: variable metastatic potentials correlated with vimentin expression. Proteomics. 2007;7:1973–83.
Hayashi E, Kuramitsu Y, Okada F, Fujimoto M, Zhang X, Kobayashi M, et al. Proteomic profiling for cancer progression: differential display analysis for the expression of intracellular proteins between regressive and progressive cancer cell lines. Proteomics. 2005;5:1024–32.
He X, Cao X. Identification of alternatively spliced GRIM-19 mRNA in kidney cancer tissues. J Hum Genet. 2010;55:507–11.
Li Y, Liang Q, Wen YQ, Chen LL, Wang LT, Liu YL, et al. Comparative proteomics analysis of human osteosarcomas and benign tumor of bone. Cancer Genet Cytogenet. 2010;198:97–106.
Shimizu T, Chiqira M, Nagase M, Watanabe H, Udagawa E. HLA phenotypes in patients who have osteosarcoma. J Bone Jt Surg Am. 1990;72:68–70.
Acknowledgments
We thank the grants from the National 973 Project of China (2011CB910700), Shanghai Natural Science Foundation (09ZR1426300), and Shanghai Hospital Clinical Research Platform for Clinical Research Source (No. SHDC12007203). We thank Kirill Gorshkov for carefully editing the whole manuscript.
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding authors
Additional information
Yingqi Hua, Xiaofang Jia, Mengxiong Sun, and Longpo Zheng contributed equally to this work.
Electronic supplementary materials
Below is the link to the electronic supplementary material.
Figure S1
The SDS-PAGE gels of 50 ug MG and Normal PM proteins to make sure equal protein load for western blotting analysis. (JPEG 15 kb)
Table S1
Table S1 Lists of non-redundant peptides identified in each protein spot. (XLS 24 kb)
Table S2
Information of proteins involved in protein-protein interaction network. (XLS 113 kb)
Rights and permissions
About this article
Cite this article
Hua, Y., Jia, X., Sun, M. et al. Plasma membrane proteomic analysis of human osteosarcoma and osteoblastic cells: revealing NDRG1 as a marker for osteosarcoma. Tumor Biol. 32, 1013–1021 (2011). https://doi.org/10.1007/s13277-011-0203-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-011-0203-4